Abstract
Purpose
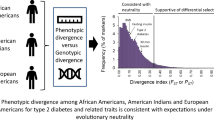
Type 2 diabetes mellitus (T2DM) is highly heritable and exhibits significant variability in prevalence between different populations. Prevalence of T2DM is higher in Asian and African relative to European populations. During evolution, traditional feast-famine cycles likely led to significant natural selection impacting metabolic genes. Human adaptation to environmental changes (food supply, lifestyle, climate, and geography) likely influenced differential selection of T2DM-associated genes. Together, insulin receptor substrate-1 and -2 (IRS1 and IRS2) genes encode the major ligands of insulin and IGF1 receptors. Irs2-deficient mice exhibit a T2DM phenotype with severe insulin resistance, and a common IRS2 polymorphism is associated with T2DM. Therefore, the present study sought evidence of natural selection at IRS2 loci.
Methods
Data were sourced from the HapMap and 1000 Genomes projects, comprising four different populations with distinct ancestries: European, Yoruba, Han Chinese, and Japanese. A three-step method was applied to detect IRS2 locus selection. The long-range haplotype (LRH) test detected unusual extended haplotypes, the integrated haplotype score (iHS) detected selection, and Wright’s F-statistics (particularly Wright’s fixation index: FST) were calculated as a measure of population differentiation.
Results
The African population exhibited highly significant LRH findings (percentile >99.9, p = 0.005–0.0009), while both the European and African populations exhibited extreme positive iHS test scores ([iHS] >2.5).
Conclusion
These findings indicate that genetic selection has occurred at the IRS2 locus, warranting further research into the adaptive evolution of metabolic disorder-associated genes.
Similar content being viewed by others
References
Diamond J. Evolution, consequences and future of plant and animal domestication. Nature. 2002;418:700–7.
Sabeti PC, Reich DE, Higgins JM, Levine HZ, Richter DJ, Schaffner SF, et al. Detecting recent positive selection in the human genome from haplotype structure. Nature. 2002;419:832–7.
Yoshiuchi I. Evidence of selection at insulin receptor substrate-1 gene loci. Acta Diabetol. 2013;50:775–9. https://doi.org/10.1007/s00592-012-0414-1.
Yoshiuchi I. Two SNPs associated with type 2 diabetes and obesity at melanocortin-4 receptor gene loci exhibited high Fst values and natural selection. J Diabetes Metab. 2013;S11:005.
NEEL JV. Diabetes mellitus: a "thrifty" genotype rendered detrimental by "progress"? Am J Hum Genet. 1962;14:353–62.
Almgren P, Lehtovirta M, Isomaa B, Sarelin L, Taskinen MR, Lyssenko V, et al. Heritability and familiarity of type 2 diabetes and related quantitative traits in the Botnia study. Diabetologia. 2011;54:2811–9.
Manolio TA, Collins FS, Cox NJ, Goldstein DB, Hindorff LA, Hunter DJ, et al. Finding the missing heritability of complex diseases. Nature. 2009;461:747–53.
Diamond J. The double puzzle of diabetes. Nature. 2003;423:599–602.
Withers DJ, Gutierrez JS, Towery H, Burks DJ, Ren JM, Previs S, et al. Disruption of IRS-2 causes type 2 diabetes in mice. Nature. 1998;391:900–4.
Mammarella S, Romano F, Di Valerio A Creati B, Esposito DL, Palmirotta R, et al. Interaction between the G1057D variant of IRS-2 and overweight in the pathogenesis of type 2 diabetes. Hum Mol Genet. 2000;9:2517–21.
Voight BF, Kudaravalli S, Wen X, Pritchard JK. A map of recent positive selection in the human genome. PLoS Biol. 2006;4:446–58.
Novembre J, Di Rienzo A. Spatial patterns of variation due to natural selection in humans. Nat Rev Genet. 2009;10:745–55.
The International HapMap Consortium. A haplotype map of the human genome. Nature. 2005;437:1299–320.
Pickrell JK, Coop G, Novembre J, Kudaravalli S, Li JZ, Absher D, et al. Signals of recent positive selection in a worldwide sample of human populations. Genome Res. 2009;19:826–37.
Holsinger KE, Weir BS. Genetics in geographically structured populations: defining, estimating and interpreting FST. Nat Rev Genet. 2009;10(9):639–50.
The 1000 Genomes Project Consortium. A global reference for human genetic variation. Nature.2015; 526: 68–74. doi:https://doi.org/10.1038/nature15393
Ehrmann DA, Tang X, Yoshiuchi I, Cox NJ, Bell GI. Relationship of insulin receptor substrate-1 and -2 genotypes to phenotypic features of polycystic ovary syndrome. J Clin Endocrinol Metab. 2002;87:4297–300.
Excoffier L, Laval G, Schneider S. Arlequin (version 3.0): an integrated software package for population genetics data analysis. Evol Bioinformatics Online. 2005;1:47–50.
Southam L, Soranzo N, Montgomery SB, Frayling TM, McCarthy MI, Barroso I, et al. Is the thrifty genotype hypothesis supported by evidence based on confirmed type 2 diabetes- and obesity-susceptibility variants? Diabetologia. 2009;52:1846–51.
Mountain JL, Risch N. Assessing genetic contributions to phenotypic differences among 'racial' and 'ethnic' groups. Nat Genet. 2004;36:S48–53.
Darwin C. The origin of species by means of natural selection. London:John Murray;1859.
Rosenberg NA, Pritchard JK, Weber JL, Cann HM, Kidd KK, Zhivotovsky LA, et al. Genetic structure of human populations. Science. 2002;298:2381–5. https://doi.org/10.1126/science.1078311.
McDougall I, Brown FH, Fleagle JG.Stratigraphic placement and age of modern humans from Kibish, Ethiopia. Nature. 2005;433:733–736. doi: https://doi.org/10.1038/nature03258.
Nielsen R, Akey JM, Jakobsson M, Pritchard JK, Tishkoff S, Willerslev E. Tracing the peopling of the world through genomics. Nature. 2017;541:302–10. https://doi.org/10.1038/nature21347.
Vigilant L, Stoneking M, Harpending H, Hawkes K, Wilson AC. African populations and the evolution of human mitochondrial DNA. Science. 1991;253:1503–7. https://doi.org/10.1126/science.1840702.
Fu Q, Posth C, Hajdinjak M, Petr M, Mallick S, Fernandes D, et al. The genetic history of ice age Europe. Nature. 2016;534:200–5. https://doi.org/10.1038/nature17993.
Allentoft ME, Sikora M, Sjögren KG, Rasmussen S, Rasmussen M, Stenderup J, et al. Population genomics of Bronze Age Eurasia. Nature. 2015;522:167–72. https://doi.org/10.1038/nature14507.
Ayub Q, Moutsianas L, Chen Y, Panoutsopoulou K, Colonna V, Pagani L, et al. Revisiting the thrifty gene hypothesis via 65 loci associated with susceptibility to type 2 diabetes. Am J Hum Genet. 2015;94:176–85. https://doi.org/10.1016/j.ajhg.2013.12.010.
Berg JJ, Coop G. A population genetic signal of polygenic adaptation. PLoS Genet. 2014;10:e1004412. https://doi.org/10.1371/journal.pgen.1004412.
Mathieson I. Human adaptation over the past 40,000 years. Curr Opin Genet. 2020;62:97–104. https://doi.org/10.1016/j.gde.2020.06.003.
Chen R, Corona E, Sikora M, Dudley JT, Morgan AA, Moreno-Estrada A, et al. Type 2 diabetes risk alleles demonstrate extreme directional differentiation among human populations, compared to other diseases. PLoS Genet. 2012;8:e1002621. https://doi.org/10.1371/journal.pgen.1002621.
Ségurel L, Austerlitz F, Toupance B, Gautier M, Kelley JL, Pasquet P, et al. Positive selection of protective variants for type 2 diabetes from the Neolithic onward: a case study in Central Asia. Eur J Hum Genet. 2013;21:1146–51. https://doi.org/10.1038/ejhg.2012.295.
Field Y, Boyle EA, Telis N, Gao Z, Gaulton KJ, Golan D, et al. Detection of human adaptation during the past 2000 years. Science. 2016;354:760–4. https://doi.org/10.1126/science.aag0776.
Acknowledgements
We thank Kyoko Yoshiuchi for secretarial contribution. We would like to thank Editage (www.editage.com) for English language editing. Dr. Issei Yoshiuchi planned this study, examined data, wrote this manuscript, thought about discussion, and edited this manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interests
The author declares no conflict of interest.
Duality of interest
The authors declare that there is no duality of interest associated with this manuscript.
Ethics approval
All procedures followed were in accordance with the ethical standards of the responsible committee on human experimentation and with the Helsinki Declaration of 1975, as revised in 2008.
This article does not contain any studies with human participants performed by any of the authors.
Informed consent
Informed consent was obtained from all individual participants included in the study.
Prior Presentation
We presented parts of this project at the 57th Annual Meeting of Japan Diabetes Association, Osaka, Japan, in May 2014.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
ESM 1
(DOCX 13 kb)
Rights and permissions
About this article
Cite this article
Yoshiuchi, I. Analysis of genetic selection at insulin receptor substrate-2 gene loci. J Diabetes Metab Disord 20, 307–311 (2021). https://doi.org/10.1007/s40200-021-00745-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40200-021-00745-y