Abstract
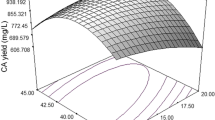
Nitrilases are enzymes with varied applications preferably used in preparation of industrially important compounds as well as in bioremediation processes. The production of enzymes is influenced by medium compositions as some of the constituents of medium directly affect the enzyme synthesis. In the present study, the components of the medium are optimized by classical and response surface methodology to maximize the production of intracellular nitrilase from Streptomyces sp. MTCC 7546. Glucose was observed as the most suitable carbon source for production of nitrilase by ‘one factor at a time’. The optimal concentration of glucose (5 g/L) was set as zero coded level for further optimization study. A central composite design was employed to study the effect of interaction among glucose, yeast extract and mineral base and evaluated for maximum nitrilase yield. The nitrilase production of 11.54 U/mg was observed at 24 h in medium containing 6.5 g/L glucose, 1.0 g/L yeast extract and 220 mL/L mineral base with 10 mM of benzonitrile as inducer. A twofold increment from 5.95 U/mg in the enzyme yield was observed after the optimization study of response surface methodology.




Similar content being viewed by others
References
Gupta V, Gaind S, Verma PK, Sood N, Srivastava AK (2010) Purification and characterization of intracellular nitrilases from Rhodococcus sp.—potential role of periplasmic nitrilase. Afr J Microbiol Res 4:1148–1153
Ghanayem BI, Nyska A, Haseman JK, Bucher JR (2002) Acrylonitrile is a multisite carcinogen in male and female B6C3F1 mice. Toxicol Sci 68:59–68
Singh R, Sharma R, Tewari N, Geetanjali Rawat DS (2006) Nitrilase and its application as a ‘green’ catalyst. Chem Biodivers 3:1279–1287
Nigam VK, Khandelwal AK, Gothwal RK, Mohan MK, Choudhury B, Vidyarthi AS, Ghosh P (2009) Nitrilase catalyzed conversion of acrylonitrile by free and immobilized cells of Streptomyces sp. J Biosci 34:21–26
Mathew CD, NagasawaT Kobayashi M, Yamada H (1998) Nitrilase-catalyzed production of Nicotinnic acid from 3-cyanopyridine in Rhodococcus rhodochrous J1. Appl Environ Microbiol 54:1030–1032
Nagasawa T, Nakamura T, Yamada H (1990) Production acrylic acid and Methacrylic acid using Rhodococcus rhodochrous J1 nitrilase. Appl Microbiol Biotechnol 34:322–324
Nigam VK, Agarwal A, Sharma M, Ghosh P, Choudhury B (2009) Bioconversion of 3-cyanopyridine to nicotinic acid by a thermostable nitrilase. Res J Biotech 4:32–36
Agarwal A, Nigam VK (2014) Nitrilase mediated conversion of indole-3-acetonitrile to indole-3-acetic acid. Biocatal Agric Biotechnol 3:351–357
Shen M, Zheng Y, Shen YC (2009) Biosynthesis of p-methoxyphenylacetic acid from p-methoxyphenylacetonitrile by immobilized Bacillus subtilis ZJB-063. Process Biochem 43:978–983
Vega-Hernάndez MC, Leόn-Barrios M, Pèrez-Galdona R (2002) Indole-3-acetic acid production from indole-3-acetonitrile in Bradyrhizobium. Soil Biol Biochem 34:665–668
Wang YJ, Zheng YG, Xue JP, Shen YC (2006) Microbial transformation of indole-3-acetonitrile to indole-3-acetamide by Nocardia sp. 108. Process Biochem 41:1746–1750
Kobayashi M, Suzuki T, Fujita T, Masuda M, Shimizu S (1995) Occurrence of enzymes involved in biosynthesis of indole-3-acetic acid from indole-3-acetonitrile in plant associated bacteria, Agrobacterium and Rhizobium. Proc Natl Acad Sci USA 92:714–718
Kobayashsi M, Izui H, Nagasawa T, Yamada H (1993) Nitrilase in biosynthesis of the plant hormone indole-3-acetic acid from indole-3-acetonitrile: cloning of the Alcaligenes gene and site directed mutagenesis of cysteine residues. Proc Natl Acad Sci USA 90:247–251
Kim YB, Yang SJ, Lee JW, Cha J, Hong SY, Chung SH, Lee ST, Rhim SL (1996) Cloning and expression of an amylase gene from Streptomyces albus KSM-35 in Escherichia coli. Food Biotechnol 5:249–253
Voff JN, Altenbuchner J (1998) Genetic instability of the Streptomyces chromomsome. Mol Microbiol 27:239–246
Banga J, Tripathi CKM (2009) Response surface methodology for optimization of medium components in submerged culture of Aspergillus flavus for enhanced heparinise production. Lett Appl Microbiol 49:204–209
Adinarayana K, Ellaiah P (2002) Response surface optimization of the critical medium components for the production of alkaline protease by a new isolated Bacillus sp. J Pharm Pharmaceut Sci 5:272–278
Dubey S, Singh A, Banerjee UC (2011) Response surface methodology of nitrilase production by recombinant Escherichia coli. Braz J Microbiol 42:1085–1092
Kumari A, Mahapatra P, Banerjee R (2009) Statistical optimization of culture conditions by response surface methodology for synthesis of lipase with Enterobacter aerogens. Braz Arch Biol Technol 52:1349–1356
Shen M, Liu ZQ, Zheng YG, Shen YC (2009) Enhancing endonitrilase production by a newly isolated Arthrobacter nitroguajacolicus ZJUTB06-99 through optimization of culture medium. Biotechnol Bioprocess Eng 14:795–802
Dong HP, Liu ZQ, Zheng YG, Shen YC (2011) Medium optimization for nitrilase production by newly isolated Rhodococcus erythropolis ZJB-0910 using statistical designs. Chem Biochem Eng 25:351–358
Khandelwal AK, Nigam VK, Choudhury B, Medicherla KM, Ghosh P (2007) Optimization of nitrilase production from a new thermophillic isolate. J Chem Technol Biotechnol 82:642–661
Piotrowski M, Schonfelder S, Weiler EW (2001) The Arabidopsis thaliana isogene NIT4 and its orthologs in Tabacco encode β-cyano-l-alanine hydratase/nitrilase. J Biol Chem 276:2616–2621
Miller GL (1959) Use of Dinitrosalicylic acid reagent for determination of reducing sugar. Anal Chem 31:426–428
Standbury PF, Whitaker A, Hall SJ (1998) Princsssiples of fermentation technology. Aditya, New Delhi
Tanyildizi MS, Özer D, Elibol M (2005) Optimization of α-amylase production by Bacillus sp. using response surface methodology. Process Biochem 40:2291–2296
Kashmiri ZN, Mankar SA (2014) Free radicals and oxidative stress in bacteria. Int J Curr Microbiol App Sci 3:34–40
Kamal A, Kumar MS, Kumar CG, Shaik T (2011) Bioconversion of acrylonitrile to acrylic acid by Rhodococcus ruber strain AKSH-84. J Microbiol Biotechnol 21:37–42
Yamamoto K, Komatsu K (1991) Purification and characterization of nitrilase responsible for the hydrolysis from Acinetobacter sp. AK 226. Agric Biol Chem 55:1459–1466
Varshney AK, Mohan MK, Vidyarthi AS, Nigam VK, Ghosh P (2013) Statistical optimization of medium components to increase the manganese peroxidise by Phanerochaete chrysoporium NCIM 1197. Biotechnol Bioprocess Eng 18:1176–1184
Muralidhar RV, Chirumamila RR, Marchant R, Nigam P (2001) A response surface approach for the comparison of lipase production by Candida cylindracea using two different carbon sources. Biochem Eng J 9:17–23
Acknowledgments
Authors are thankful to the University Grants Commission, New Delhi, India for providing Junior Research Fellowship to Astha Agarwal (F.2-29/2011; SA-1) for carrying out this research work. Authors are also thankful to Centre of Excellence (COE), Department of Biotechnology under TEQIP phase II (Ref No: NPIU/TEQIP II/FIN/31/158; Dated 16th April 2013) for laboratory facilities and infrastructure support and lastly, authors are grateful to Dr. Alok Varshney for sincere help during experimental work. Authors also declare that there is no financial or commercial conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Agarwal, A., Nigam, V.K. Enhanced Production of Nitrilase from Streptomyces sp. MTCC 7546 by Response Surface Method. Proc. Natl. Acad. Sci., India, Sect. B Biol. Sci. 87, 603–609 (2017). https://doi.org/10.1007/s40011-015-0638-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40011-015-0638-2