Abstract
Background:
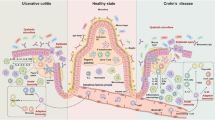
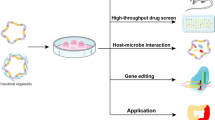
Intestinal organoids have evolved as potential molecular tools that could be used to study host-microbiome interactions, nutrient uptake, and drug screening. Gut epithelial barrier functions play a crucial role in health and diseases, especially in autoimmune diseases, such as inflammatory bowel diseases (IBDs), because they disrupt the epithelial mucosa and impair barrier function.
Methods:
In this study, we generated an in vitro IBD model based on dextran sodium sulfate (DSS) and intestinal organoids that could potentially be used to assess barrier integrity. Intestinal organoids were long-term cultivated and characterized with several specific markers, and the key functionality of paracellular permeability was determined using FITC-dextran 4 kDa. Intestinal organoids that had been treated with 2 µM DSS for 3 h were developed and the intestinal epithelial barrier function was sequentially evaluated.
Results:
The results indicated that the paracellular permeability represented epithelial characteristics and their barrier function had declined when they were exposed to FITC-dextran 4 kDa after DSS treatment. In addition, we analyzed the endogenous mRNA expression of pro-inflammatory cytokines and their downstream effector genes. The results demonstrated that the inflammatory cytokines genes significantly increased in inflamed organoids compared to the control, leading to epithelial barrier damage and dysfunction.
Conclusion:
The collective results showed that in vitro 3D organoids mimic in vivo tissue topology and functionality with minor limitations, and hence are helpful for testing disease models.



Similar content being viewed by others
References
Chelakkot C, Ghim J, Ryu SH. Mechanisms regulating intestinal barrier integrity and its pathological implications. Exp Mol Med. 2018;50:103.
Deuring JJ, de Haar C, Kuipers EJ, Peppelenbosch MP, van der Woude CJ. The cell biology of the intestinal epithelium and its relation to inflammatory bowel disease. Int J Biochem Cell Biol. 2013;45:798–806.
Kraiczy J, Zilbauer M. Intestinal epithelial organoids as tools to study epigenetics in gut health and disease. Stem Cells Int. 2019;2019:7242415.
Lancaster MA, Knoblich JA. Organogenesis in a dish: modeling development and disease using organoid technologies. Science. 2014;345:1247125.
Sato T, Vries RG, Snippert HJ, van de Wetering M, Barker N, Stange DE, et al. Single Lgr5 stem cells build crypt-villus structures in vitro without a mesenchymal niche. Nature. 2009;459:262–5.
Sato T, Stange DE, Ferrante M, Vries RG, Van Es JH, Van den Brink S, et al. Long-term expansion of epithelial organoids from human colon, adenoma, adenocarcinoma, and Barrett’s epithelium. Gastroenterology. 2011;141:1762–72.
McCracken KW, Catá EM, Crawford CM, Sinagoga KL, Schumacher M, Rockich BE, et al. Modelling human development and disease in pluripotent stem-cell-derived gastric organoids. Nature. 2014;516:400–4.
Jee JH, Lee DH, Ko J, Hahn S, Jeong SY, Kim HK, et al. Development of collagen-based 3D matrix for gastrointestinal tract-derived organoid culture. Stem Cells Int. 2019;2019:8472712.
Ko JK, Auyeung KK. Inflammatory bowel disease: etiology, pathogenesis and current therapy. Curr Pharm Des. 2014;20:1082–96.
Verstockt B, Smith KG, Lee JC. Genome-wide association studies in Crohn’s disease: past, present and future. Clin Transl Immunology. 2018;7:e1001.
Zhang YZ, Li YY. Inflammatory bowel disease: pathogenesis. World J Gastroenterol. 2014;20:91–9.
Westbrook AM, Szakmary A, Schiestl RH. Mechanisms of intestinal inflammation and development of associated cancers: lessons learned from mouse models. Mutat Res. 2010;705:40–59.
Hibiya S, Tsuchiya K, Hayashi R, Fukushima K, Horita N, Watanabe S, et al. Long-term inflammation transforms intestinal epithelial cells of colonic organoids. J Crohns Colitis. 2017;11:621–30.
DeVoss J, Diehl L. Murine models of inflammatory bowel disease (IBD): challenges of modeling human disease. Toxicol Pathol. 2014;42:99–110.
Scheiffele F, Fuss IJ. Induction of TNBS colitis in mice. Curr Protoc Immunol. 2002;Chapter 15:Unit 15.19.
Kiesler P, Fuss IJ, Strober W. Experimental models of inflammatory bowel diseases. Cell Mol Gastroenterol Hepatol. 2015;1:154–70.
Bredenoord AL, Clevers H, Knoblich JA. Human tissues in a dish: the research and ethical implications of organoid technology. Science. 2017;355:eaaf9414.
Sato T, Clevers H. Growing self-organizing mini-guts from a single intestinal stem cell: mechanism and applications. Science. 2013;340:1190–4.
Lee BR, Kim H, Park TS, Moon S, Cho S, Park T, et al. A set of stage-specific gene transcripts identified in EK stage X and HH stage 3 chick embryos. BMC Dev Biol. 2007;7:60.
Rallabandi HR, Yang H, Jo YJ, Lee HC, Byun SJ, Lee BR. Identification of female specific genes in the W chromosome that are expressed during gonadal differentiation in the chicken. Korean J Poult Sci. 2019;46:287–96.
Pearce SC, Al-Jawadi A, Kishida K, Yu S, Hu M, Fritzky LF, et al. Marked differences in tight junction composition and macromolecular permeability among different intestinal cell types. BMC Biol. 2018;16:19.
Sala FG, Kunisaki SM, Ochoa ER, Vacanti J, Grikscheit TC. Tissue-engineered small intestine and stomach form from autologous tissue in a preclinical large animal model. J Surg Res. 2009;156:205–12.
Yan KS, Chia LA, Li X, Ootani A, Su J, Lee JY, et al. The intestinal stem cell markers Bmi1 and Lgr5 identify two functionally distinct populations. Proc Natl Acad Sci U S A. 2012;109:466–71.
Pastula A, Middelhoff M, Brandtner A, Tobiasch M, Höhl B, Nuber AH, et al. Three-dimensional gastrointestinal organoid culture in combination with nerves or fibroblasts: a method to characterize the gastrointestinal stem cell niche. Stem Cells Int. 2016;2016:3710836.
Nagatake T, Fujita H, Minato N, Hamazaki Y. Enteroendocrine cells are specifically marked by cell surface expression of claudin-4 in mouse small intestine. PLoS One. 2014;9:e90638.
Altay G, Larrañaga E, Tosi S, Barriga FM, Batlle E, Fernández-Majada V, et al. Self-organized intestinal epithelial monolayers in crypt and villus-like domains show effective barrier function. Sci Rep. 2019;9:10140.
Schmitt M, Schewe M, Sacchetti A, Feijtel D, van de Geer WS, Teeuwssen M, et al. Paneth cells respond to inflammation and contribute to tissue regeneration by acquiring stem-like features through SCF/c-Kit signaling. Cell Rep. 2018;24:2312–28.e7.
Dotti I, Salas A. Potential use of human stem cell-derived intestinal organoids to study inflammatory bowel diseases. Inflamm Bowel Dis. 2018;24:2501–9.
Kitajima S, Takuma S, Morimoto M. Histological analysis of murine colitis induced by dextran sulfate sodium of different molecular weights. Exp Anim. 2000;49:9–15.
Okayasu I, Hatakeyama S, Yamada M, Ohkusa T, Inagaki Y, Nakaya R. A novel method in the induction of reliable experimental acute and chronic ulcerative colitis in mice. Gastroenterology. 1990;98:694–702.
Bardenbacher M, Ruder B, Britzen-Laurent N, Schmid B, Waldner M, Naschberger E, et al. Permeability analyses and three dimensional imaging of interferon gamma-induced barrier disintegration in intestinal organoids. Stem Cell Res. 2019;35:101383.
Michielan A, D’Incà R. Intestinal permeability in inflammatory bowel disease: pathogenesis, clinical evaluation, and therapy of leaky gut. Mediators Inflamm. 2015;2015:628157.
Jaworska K, Konop M, Bielinska K, Hutsch T, Dziekiewicz M, Banaszkiewicz A, et al. Inflammatory bowel disease is associated with increased gut-to-blood penetration of short-chain fatty acids: a new, non-invasive marker of a functional intestinal lesion. Exp Physiol. 2019;104:1226–36.
Dobre M, Milanesi E, Mănuc TE, Arsene DE, Ţieranu CG, Maj C, et al. Differential intestinal mucosa transcriptomic biomarkers for Crohn’s disease and ulcerative colitis. J Immunol Res. 2018;2018:9208274.
Kobayashi K, Arimura Y, Goto A, Okahara S, Endo T, Shinomura Y, et al. Therapeutic implications of the specific inhibition of causative matrix metalloproteinases in experimental colitis induced by dextran sulphate sodium. J Pathol. 2006;209:376–83.
Kawamoto A, Nagata S, Anzai S, Takahashi J, Kawai M, Hama M, et al. Ubiquitin D is upregulated by synergy of notch signalling and TNF-α in the inflamed intestinal epithelia of IBD patients. J Crohns Colitis. 2019;13:495–509.
Dhillon SS, Mastropaolo LA, Murchie R, Griffiths C, Thöni C, Elkadri A, et al. Higher activity of the inducible nitric oxide synthase contributes to very early onset inflammatory bowel disease. Clin Transl Gastroenterol. 2014;5:e46.
Giles EM, Sanders TJ, McCarthy NE, Lung J, Pathak M, MacDonald TT, et al. Regulation of human intestinal T-cell responses by type 1 interferon-STAT1 signaling is disrupted in inflammatory bowel disease. Mucosal Immunol. 2017;10:184–93.
Armaka M, Apostolaki M, Jacques P, Kontoyiannis DL, Elewaut D, Kollias G. Mesenchymal cell targeting by TNF as a common pathogenic principle in chronic inflammatory joint and intestinal diseases. J Exp Med. 2008;205:331–7.
Sandborn WJ, Hanauer SB. Antitumor necrosis factor therapy for inflammatory bowel disease: a review of agents, pharmacology, clinical results, and safety. Inflamm Bowel Dis. 1999;5:119–33.
Acknowledgement
This work was supported with the National Institute of Animal Sciences (Grant No. PJ01422201), Rural Development Administration (RDA), Korea.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
The authors declare no conflicts of interest.
Ethical statement
The experimental use of mouse was performed after receiving approval of the Institutional Animal Care and Use Committee (IACUC) of National Institute of Animal Science (NIAS-2019-366), Korea.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Figure S1.
Isolation of intestinal crypts. A ~5-week-old ICR mouse was used to extract the intestinal stem cells (left upper panel). A section of the small intestine was obtained from the euthanized mouse and washed thoroughly with PBS under aseptic conditions (left lower panel). Histological H and E staining of vertical and horizontal sections from the small intestine showing the presence of crypts (oval structures at the bottom) and villi (finger like structures). The purple spots show nuclei and the epithelium is stained pink. Scale bar: 100 μm (right upper panel) and 50 μm (right middle and lower panels). (PPT 6462 kb)
Figure S2. In vitro intestinal organoid IBD model generation using DSS. (A) The organoids were treated with the chemical agent DSS at 1 µM, 2 µM, or 5 µM per well (upper panel) to identify and optimize the lethal dose needed to induce inflammation. After incubation for 1 h, inflammation was observed in the 2 µM and 5 µM wells, but not in 1 µM wells. (B) Time course data showing the effects of the 2 µM lethal dose at 30 min, 60 min, and 180 min post-treatment (lower panel). There was minimal damage at 30 min post-treatment, but the organoids were highly inflamed and had begun to disintegrate at 180 min. There was also damage to the basement membrane, magnitude of inflammation was indicated in circles using designated colors yellow (damage) and red (lethal) in the figure. Scale bar: 100 μm (PPT 6462 kb)
Figure S3. Culturing and measuring the growth rate of mouse intestinal organoids in long-term cultures. (A) Long term culturing of intestinal organoids for up to 10 generations (P2-P10). Middle and lower panels show various developmental stages. The organoids can be spheroidal (round shaped), show budding (spheroids with extension), and have mature villus and crypt like structures (branched structures). (B) Growth rate graph showing the number of organoids/well (mean n = 3 wells) growing in a 100 µL matrigel dome on a 24 well plate. Scale bar: 100 μm (PPT 6462 kb)
Rights and permissions
About this article
Cite this article
Rallabandi, H.R., Yang, H., Oh, K.B. et al. Evaluation of Intestinal Epithelial Barrier Function in Inflammatory Bowel Diseases Using Murine Intestinal Organoids. Tissue Eng Regen Med 17, 641–650 (2020). https://doi.org/10.1007/s13770-020-00278-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13770-020-00278-0