Abstract
Broad-scale mass spectrometric analyses of glycopeptides are constrained by the considerable complexity inherent to glycoproteomics, and techniques are still being actively developed to address the associated analytical difficulties. Here we apply Orbitrap mass analysis and higher-energy C-trap dissociation (HCD) to facilitate detailed insights into the compositions and heterogeneity of complex mixtures of low abundance glycopeptides. By generating diagnostic oxonium product ions at mass measurement errors of <5 ppm, highly selective glycopeptide precursor ion detections are made at sub-fmol limits of detection: analyses of proteolytic digests of a hen egg glycoprotein mixture detect 88 previously uncharacterized glycopeptides from 666 precursor ions selected for MS/MS, with only one false positive due to co-fragmentation of a non-glycosylated peptide with a glycopeptide. We also demonstrate that by (1) identifying multiple series of glycoforms using high mass accuracy single stage MS spectra, and (2) performing product ion scans at optimized HCD collision energies, the identification of peptide + N-acetylhexosamine (HexNAc) ions (Y1 ions) can be readily achieved at <5 ppm mass measurement errors. These data allow base peptide sequences and glycan compositional information to be attained with high confidence, even for glycopeptides that produce weak precursor ion signals and/or low quality MS/MS spectra. The glycopeptides characterized from low fmol abundances using these methods allow two previously unreported glycosylation sites on the Gallus gallus protein ovoglycoprotein (amino acids 82 and 90) to be confirmed; considerable glycan heterogeneities at amino acid 90 of ovoglycoprotein, and amino acids 34 and 77 of Gallus gallus ovomucoid are also revealed.
Similar content being viewed by others
1 Introduction
Over half of all proteins are thought to undergo post-translational modification (PTM) by carbohydrates [1]. Such modifications have been implicated in numerous intracellular, cell–cell and cell–matrix recognition events [2], which has underscored the recognition of protein glycosylation as a critical area of research [2–4]. Mass spectrometry (MS) has emerged as the analytical discipline of choice within this field of study [5–12], and developments in MS instrumentation and techniques have significantly enhanced our ability to profile protein glycosylations. Efforts at characterizing these PTMs, however, remain constrained by the considerable heterogeneity of oligosaccharide structure. As such, the full potential held by MS to facilitate broad-scale, systemic analyses of protein glycosylations remains unfulfilled, and methodologies are still being actively developed to address the analytical difficulties associated with this area of research.
Existing methodologies commonly treat the peptide portion of glycoproteins and the attached glycan moieties as independent entities. To this end, complete or partial deglycosylation of peptides prior to MS analysis is generally undertaken. Though these protocols allow for the large-scale identification of glycoproteins and their glycosylation sites [13–17], the biological conclusions that can be drawn from these techniques are severely limited by their inability to determine site-specific oligosaccharide structures.
An alternate strategy is to characterize intact glycopeptides following glycoprotein proteolysis [5, 6]. It has been well established that peptide sequences, glycan structures and glycan attachment sites can be comprehensively elucidated from intact glycopeptides using various modes of fragmentation in tandem MS (MS/MS) [5]. The use of pronase to achieve nonspecific enzymatic proteolysis can also contribute towards an exposition of oligosaccharide structural and attachment site heterogeneity [18–24]. To characterize intact glycopeptides on a proteome-wide scale, the steps associated with such analyses must ultimately be addressed using automated, high throughput data analysis workflows such as Peptoonist [25], GlycoMiner [26] and the automated workflows of Joenväärä et al. [27, 28] (summarized in Scheme 1). Analyses of this breadth dictate that these workflows must be capable of being applied to complex mixtures featuring glycopeptides of low abundance. As outlined below, the promise held by such automated workflows, however, remains constrained by existing methods of data collection.
Step 1 of such analyses, the detection of glycopeptides within the mixture subjected to MS, is critical to the analysis of glycoprotein digests produced via high specificity enzymes. To this end, marker ion observation [29–33] has emerged as the primary method of choice, and glycan derived oxonium ions produced via collision-induced dissociation (CID) in MS/MS have proven to be particularly robust markers [29–31]. For this purpose the oxonium ions of hexose (Hex) at mass-to-charge ratio (m/z) 163.0601, N-acetylhexosamine (HexNAc) at m/z 204.0867, Hex1HexNAc1 at m/z 366.1395, and N-acetylneuraminic acid (NeuNAc) at m/z 292.1027 are frequently employed. A major drawback of this technique is that detection of these ions can be rendered impossible when mass analyzers that exhibit low mass cutoffs following CID are utilized. Another drawback relates to interference from non-glycosylated peptide fragmentation [31, 34]. Though high mass accuracy marker ion detection in quadrupole time-of-flight (Q-TOF) precursor ion scanning experiments has been shown to minimize this issue [34], the long scan times associated with this method severely limit its applicability. Alternatively, source CID followed by selected ion monitoring (SIM) [35] or MS/MS in pseudo-MS3 experiments [36] allow for high levels of sensitivity in oxonium marker ion detection. The nonselective nature of the source CID, however, precludes the assignment of these oxonium ions to specific precursor ions. As such, sensitive and selective methods for detecting oxonium marker ions in broad-scale glycopeptide analyses are still lacking.
Steps 2a and 2b of such analyses involve, respectively, base peptide and oligosaccharide characterization. In undertaking these steps, one approach—utilized by the GlycoMod tool [37] —involves matching theoretical glycopeptide masses, generated for base peptides associated with proteins identified in the sample, against those obtained experimentally. The simplicity of this technique is offset by its potential for high false discovery rates, even when high mass accuracy data is utilized [38]; MS/MS product ion scans are therefore often required to achieve adequate confirmation of glycopeptide matches. During step 2a this can be achieved using specific instrumentation [32, 33, 39, 40] or fragmentation methods [5, 41–46]; of particular note is the use of the often intensive peptide + HexNAc (Y1) ion in CID derived MS/MS spectra [47], which can support the identification of base peptide masses [48]. During step 2b, the glycosidic cleavages of sugar moieties, which dominate CID derived MS/MS spectra of glycopeptides, can aid in the assignment of glycan compositions [49, 50]. Automated glycopeptide characterization workflows typically use one or a combination of the abovementioned techniques to establish either the glycan portion or the peptide portion; mass differences between the identified portion and the observed glycopeptide mass are then used to complete the glycopeptide assignment. (Alternatively, the Peptoonist software uses MS/MS to identify both the peptide and glycan portions of a limited number of glycopeptides; the mass differences between these fully characterized glycopeptides and other glycopeptide candidates can be ascertained from MS-1 data, allowing glycoforms to be identified.) As such, glycopeptide false discovery rates are not only dictated by the accuracy of these methods, but are also constrained by a need to obtain high mass accuracy single-stage MS (MS-1) data.
In attending to the abovementioned constraints, Orbitrap mass analysis shows great promise. The capabilities of these mass analyzers in proteomics experiments are now strongly established [51–53], and Orbitrap derived high mass accuracy MS-1 spectra of intact glycopeptides have recently been implemented in the elucidation of glycan compositions [54]. Notably, following the recent implementation of beam-type CID in LTQ Orbitrap hybrid mass spectrometers, which has been termed higher-energy C-trap dissociation (HCD) [55], Orbitraps can now be used to produce, over full mass ranges, triple quadrupole-like product ion scans of high mass accuracy. Segu and Mechref have examined the applicability of this technique to glycopeptide analysis in relation to the observation of Y1 ions in model glycoproteins [56]. Such high mass accuracy MS-1 and MS/MS spectra, which are generated concomitantly during Orbitrap mass analysis and HCD in data dependent acquisition (DDA)/MS, should be expected to play a significant role in addressing the data collection requirements for high throughput glycopeptide characterization workflows. The present contribution demonstrates the utility of Orbitrap mass analysis and HCD in DDA/MS towards this purpose, particularly in relation to the analysis of complex peptide mixtures featuring low abundance glycopeptides. Tryptic digests of a well-characterized model glycoprotein, ribonuclease B (RNase B), are firstly used to obtain the low abundance glycopeptide derived data required to establish an appropriate analytical workflow using these experimental methods. Guided by the outcomes of the RNase B analyses, proteolytic digests of a mixture containing previously uncharacterized low abundance hen egg glycoproteins are then analyzed within the framework of the steps associated with high throughput glycopeptide characterizations. These analyses establish that, for each glycopeptide characterization step, the described methods of data collection are capable of facilitating strong outcomes when applied to complex peptide mixtures featuring low abundance glycopeptides; namely: highly sensitive and selective glycopeptide detections during step 1, and high confidence base peptide and glycan compositional characterizations during steps 2a and 2b.
2 Experimental
2.1 Materials
RNase B from bovine pancreas (≥80%) and hen egg white powder (≥98% ovalbumin) were obtained from Sigma-Aldrich (St Louis, MO, USA). Sequencing grade trypsin and Glu-C were obtained from Roche (Penzberg, Germany). All other reagents, chemicals, and solvents were analytical or HPLC grade.
2.2 Proteolytic Digestions
Disulfide bond reduction of RNase B (5 mg/mL in 6 M urea, 30 mM NH4HCO3, and 5 mM dithiothreitol (DTT)) was performed for 20 min at 37 °C. Samples were diluted 40-fold and trypsin was added at an enzyme to protein ratio of 1:15. Digestion proceeded for 20 h at 37 °C. Following digestion, samples were stored at −20 °C prior to analysis.
Disulfide bond reduction of hen egg white powder (2.5 mg/mL in 2 M urea, 12 mM NH4HCO3, and 1.2 mM DTT) was performed for 20 min at 37 °C. Samples were either diluted 2.5-fold and trypsin added at an enzyme to protein ratio of 1:35, or diluted 2-fold and Glu-C added at an enzyme to protein ratio of 1:25. Trypsin digests proceeded for 23 h at 37 °C, and Glu-C digests proceeded for 15 h at 37 °C. Following digestion, samples were stored at −20 °C prior to analysis.
2.3 Mass Spectrometry
RNase B and hen egg white powder digest solutions were diluted 40-fold and 100-fold, respectively, in 0.1% formic acid prior to injection for DDA/MS experiments. Injection volumes between 0.5 and 1.5 μL were used (corresponding to a maximum of 100 to 300 fmol of proteolytic peptides, or a maximum of ~2 to 6 fmol non-ovalbumin derived glycopeptides in the hen egg white powder digests). Proteolytic peptides were separated by nano-liquid chromatography (LC) using an UltiMate 3000 HPLC and autosampler system (Dionex, Amsterdam, The Netherlands). Samples were concentrated and desalted onto a CapTrap peptide precolumn (500 μm × 2 mm, Michrom Bioresources, Auburn, CA, USA) with H2O:CH3CN (98:2, 0.1% trifluoroacetic acid) at 10 μL/min. The use of 0.1% trifluoroacetic acid improves the retention of hydrophilic peptides; some RNase B derived glycopeptides, which are otherwise difficult to retain on the employed CapTrap peptide precolumn, were thus capable of being be analyzed. After a 4 min wash, the precolumn was switched (Valco 10 port valve; Dionex) into line with a fritless nanocolumn (75 μm i.d. × 10 cm) containing C18 RP packing material (5 μ Magic; Michrom Bioresources) manufactured according to Gatlin et al. [57]. Peptides were eluted using a linear gradient of H2O:CH3CN (98:2, 0.1% formic acid) to H2O:CH3CN (55:45, 0.1% formic acid) at 250 nL/min over 30 min. High voltage (2000 V) was applied to a low volume tee (Upchurch Scientific), and the column tip was positioned ~5 mm from the heated capillary (T = 275 °C) of an LTQ Orbitrap Velos (Thermo Electron, Bremen, Germany) hybrid linear ion trap and Orbitrap mass spectrometer. Positive ions were generated by electrospray, and the LTQ Orbitrap Velos operated in DDA mode.
Survey scans m/z 350–2000 were acquired in the Orbitrap (resolution = 30 000 at m/z 400, with an initial accumulation target value of 1,000,000 ions in the linear ion trap; lock mass applied to polycyclodimethylsiloxane background ions of exact m/z 445.1200 and 429.0887). Up to the 10 most abundant ions (>5000 counts) with charge states of > +2 were sequentially isolated and fragmented via HCD within the HCD collision cell with an activation q = 0.25, an activation time of 40 ms, normalized collision energies of 27%, 35%, and 42% and at a target value of 50,000 ions; fragment ions were mass analyzed in the Orbitrap without the use of a lock mass; m/z selected for MS/MS were dynamically excluded for 45 s. Peak lists were generated using Mascot Daemon/extract_msn (ver. 2.3; Matrix Science, London, England) using the default parameters. Similar DDA/MS experiments were conducted using collision induced dissociation (CID) with an activation q = 0.25, an activation time of 30 ms, normalized collision energy of 30%, and at a target value of 10,000 ions; fragment ions were mass analyzed in the linear ion trap. Each nano-LC DDA/MS experiment was replicated three times.
Additional experiments were conducted by direct infusion of undiluted proteolytic digest mixtures using nanospray emitters manufactured in-house. Source conditions were identical to those applied to nano-LC experiments with the exception of the applied voltage (900 V). To optimize HCD parameters for DDA/MS, selected precursor ions were isolated and fragmented using HCD with the activation q, activation time and normalized collision energies systematically altered within the ranges of 0.3–0.75, 0.01–100 ms, and 15%–70%, respectively. Experiments were also conducted upon selected precursor or fragment ions within the linear ion trap in MS/MS or MS3 experiments using CID with an activation q = 0.25, an activation time of 30 ms, normalized collision energies ranging from 25% to 35% and at target values of 10,000 ions.
2.4 Database Searches
DDA/MS derived peak lists were submitted to the database search program Mascot (ver. 2.3; Matrix Science, London, UK). The following search parameters were employed: precursor and MS/MS tolerances were ±4 ppm and ±0.4 Da, respectively; acrylamide (C), carbamidomethyl (C), and oxidation (M) were specified as variable modifications; enzyme specificity was either trypsin with one missed cleavage allowed, or V8-DE with two missed cleavages allowed; and the NCBInr database was searched. Protein identifications were deemed to be high confidence if they produced Mascot scores greater than the significance threshold of P = 0.05.
3 Results and Discussion
3.1 Ribonuclease B Analysis
The following RNase B analyses serve to illustrate the nature of the data capable of being produced from low abundance glycopeptides using HCD. These data, presented in relation to the steps associated with glycopeptide characterizations, are used to guide the development of an analytical workflow suitable for the characterization of complex mixtures containing previously uncharacterized low abundance glycopeptides. Such a workflow is applied and evaluated in the subsequent hen egg glycoprotein mixture analysis.
Step 1 – Detection of glycopeptide precursor ions
The present experiments suggest that, when generating data via HCD and Orbitrap mass analysis, highly selective glycopeptide detections can be achieved via the observation of oxonium ions. Figure 1a shows an extracted ion chromatogram (XIC) for fragment ions of m/z 204.0836–204.0898 obtained via DDA/MS using HCD during MS/MS. In relation to HexNAc derived oxonium ions, the bounds of this m/z window correspond to mass measurement errors of 15 ppm; this value is beyond the maximum mass measurement error of the employed Orbitrap mass analyzer, and as such, all ions with m/z values possibly corresponding to HexNAc derived oxonium ions are encompassed within this window. As summarized in the pie chart in Figure 1a, of the 107 precursor ions selected for MS/MS in this experiment, only four are associated with this XIC and therefore identified as potential glycopeptides. Each of these precursor ions do indeed correspond to known RNase B derived glycopeptides (as will be elaborated upon in the subsequent sub-section), indicating an excellent level of selectivity in glycopeptide detection.
(a) XIC for fragment ions with m/z corresponding to HexNAc derived oxonium ions generated using HCD and Orbitrap mass analysis during DDA/MS. (b) XIC’s for fragment ions with m/z corresponding to Hex1HexNAc1 derived oxonium ions generated using CID and linear ion trap mass analysis during DDA/MS. In both (a) and (b), signals confirmed as belonging to oxonium ions are highlighted in grey. For each experiment, pie charts illustrate the total number of precursor ions selected for fragmentation, precursors identified as potential glycopeptides via observation of fragment ions with m/z corresponding to glycan derived oxonium ions, and correct glycopeptide detections. All results obtained from a tryptic digest of RNase B
To confirm that HCD does indeed reduce the negative impact of non-glycosylated peptide fragmentation on the selectivity of glycopeptide detections, otherwise identical experiments were conducted using CID and linear ion trap mass analysis during MS/MS. Figure 1b shows the resulting XIC for fragment ions of m/z 365.97–366.31, and Figure S1 shows the resulting XIC for fragment ions of m/z 204.00–204.18. These m/z windows encompass all ions with m/z values possibly corresponding to Hex1HexNAc1 and HexNAc derived oxonium ions, respectively, within the maximum mass measurement errors of the employed linear ion trap mass analyzer. (Maximum mass measurement errors in CID derived MS/MS spectra from DDA/MS experiments were determined to be 450 ppm based on the observed m/z for known oxonium ions.) As summarized in the pie charts in Figure 1b and in Figure S1, 85 precursor ions were selected for MS/MS in this experiment; 63 produced fragment ions within the m/z 365.97–366.31 window and 27 produced fragment ions within the m/z 204.00–204.18 window. Careful inspection of the individual MS/MS spectra reveal that only 6 of the ions associated with the m/z 365.97–366.31 XIC can be attributed to glycopeptides (spectra not shown). None of the precursor ions associated with the m/z 204.00–204.18 XIC can be attributed to glycopeptides; this is unsurprisingly given the low mass cutoff of the linear ion trap. (Though two glycopeptides were detected via CID DDA/MS and not detected via HCD DDA/MS, the associated glycopeptide precursor ions were not selected for fragmentation during HCD DDA/MS. These additional detections can therefore be traced to variability in precursor ion selection during DDA/MS, rather than sensitivity differences stemming from the employed fragmentation techniques.) Though these results are specific to the employed instrument, and similar glycopeptide detection methods would not generally be relied upon in practice, these poor levels of selectivity reinforce the notion that interference from non-glycosylated peptide fragmentation can have an undesirable impact [31, 34]. In practice, such poor selectivity can be overcome through the use of additional glycopeptide detection criteria. As has been summarized in Scheme 1, GlycoMiner [26] and the automated workflows of Joenväärä et al. [27, 28] require the concomitant observation of ions corresponding to peptide + fragmented glycans in MS/MS spectra before a precursor ion is confirmed as a glycopeptide. This added detection criterion, however, reduces sensitivity; glycopeptides that produce relatively few glycan fragment ions will have a decreased likelihood of being detected. The current results suggest that the data generated using HCD and Orbitrap mass analysis should allow excellent levels of selectivity to be achieved without the need for these additional glycopeptide detection criteria.
Steps 2a and 2b – Characterization of the peptide and glycan portions
The experiments described below suggest that, when data is gathered using HCD and Orbitrap mass analysis in DDA/MS, the characterization steps 2a and 2b are most likely to be accurately facilitated through the use of high mass accuracy MS-1 measurements, and the concomitant observation of Y1 ions. These conclusions can be drawn through a consideration of the mass accuracy associated with the MS-1 data and the nature of the fragment ions produced during MS/MS, as discussed below.
To verify that high mass accuracies are attained from the MS-1 data associated with the present experiments, confident glycopeptide characterizations were first established. The glycopeptide characterizations made from the current sample are presented in Table 1, and detailed descriptions of the evidence used to confirm these characterizations are given in the Supplementary Material. This information allows one to conclude that sub ppm mass measurement errors are regularly obtained during MS-1 (root mean square (RMS) error of 0.5 ppm, with no mass measurement errors exceeding 1.2 ppm). These mass measurement errors are consistent with those expected from an LTQ Orbitrap with lock mass applied [58]. For studies of limited scale such as the present RNase B analysis, which is restricted to the characterization of N-linked glycopeptides, this level of mass accuracy is sufficient to allow steps 2a and 2b of glycopeptide characterizations to be undertaken at high confidence without the need for MS/MS verification. These levels of mass accuracy are not, however, sufficient to allow broader-scale MS-1 based characterizations to be free of false positives [38]; for studies of even moderate breadth, MS/MS data must be drawn upon to verify glycopeptide assignments.
In developing appropriate analytical workflows for use with HCD derived MS/MS spectra, the information required for accurate glycopeptide characterizations must be considered. As outlined in the introduction and as summarized in Scheme 1, different types of fragment ions can be used to obtain glycopeptide matches: base peptide fragment ions (b and y ions), oxonium ions, peptide + fragmented glycan ions, and more specifically, Y1 ions. Accurate glycopeptide characterizations can be made from such data if: (A) multiple fragment ions associated with one or more fragment ion type match with those expected from a given glycopeptide, and/or (B) ions highly specific to a given glycopeptide can be identified with confidence.
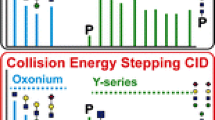
To gain insight into the fragment ions produced via HCD, relatively high abundances of glycopeptides were first isolated and fragmented via direct infusion nanospray. Figure 2a illustrates results obtained from doubly charged ions of glycopeptide III; relative abundances are shown for selected fragment ions over the 15% to 70% range of collision energies. Using collision energies in the range of 25% to 40%, one is most likely to observe peptide + fragmented glycan ions whilst also observing oxonium ions; this is reinforced in part (b) of Figure 2, which shows the total number of peptide + fragmented glycan ions observed over the 15% to 70% range of collision energies. At collision energies above 40%, the likelihood of observing peptide + fragmented glycan ions other than the Y1 ions is diminished. At collision energies below 25%, the likelihood of observing oxonium ions is unacceptably low if these ions are to be used for glycopeptide detection during DDA/MS. These experiments also show that Y1 fragment ions dominate the product ion scans over the 25% to 55% range of collision energies. Notably, b and y ions are not observed over the entire 15% to 70% range of collision energies. Altering the activation q and activation times over the ranges outlined in Section 2 did not significantly affect the relative abundances of the fragment ions of interest. These results are consistent with findings reported by Segu and Mechref, who found that for glycopeptide ions over a range of different charge states, glycan compositions, peptide compositions and peptide lengths, HCD is more conducive to producing product ion spectra dominated by Y1 ions and oxonium ions than CID in a quadrupole ion trap, which generally produces spectra dominated by peptide + fragmented glycan ions [56].
Results of HCD product ion scans of glycopeptide III of Table 1 (doubly charged, m/z 927.4) produced via direct infusion nanospray over the 15% to 70% range of collision energies in terms of: (a) the abundances of various product ions observed (relative to the base peak); and (b) the number of singly charged and doubly charged peptide + fragmented glycan ions observed. The peptide + fragmented glycan ions of (b) do not include Y1 ions, and are plotted for the most abundant ions observed
The abovementioned experiments suggest that the HCD collision energies employed during DDA/MS in the present RNase B analysis—27%, 35%, and 42%—should maximize the potential for detection of fragment ions required for glycopeptide characterizations based on method (A). Detailed assessment of the HCD DDA/MS data, however, indicates that the observation of such fragment ions remains severely limited in the low abundance glycopeptides that were analyzed. Figure 3a labels all b and y ions, intact glycan derived oxonium ions, and peptide + fragmented glycan ions produced from glycopeptides I and III in selected DDA/MS experiments. It can be seen that no b and y ions are detected, and though Y1 ions appear at high relative abundances, limited other peptide + fragmented glycan ions are observed. Though Hex and HexNAc derived oxonium ions are clearly apparent, these ions are only of limited utility in the characterization of glycopeptide structures. As can be seen in Figure 3b, which illustrates the relative abundances of different ion types observed over the entire set of RNase B DDA/MS experiments, these results can be considered to be typical. It is likely that the high production of Y1 ions stems from the propensity for beam-type CID to induce further dissociation of primary fragments; that is, peptide + fragmented glycan primary fragments are likely to undergo further dissociation to form relatively stable Y1 ions. As the observed oxonium ions are internal glycan fragments, it is likely that their high relative abundances can also be largely explained by dissociation of primary fragments. As should be expected, a greater number of peptide + fragmented glycan ions are observed in the direct infusion nanospray experiments, as the integration of multiple scans to produce a mass spectrum increases spectral quality. This optimal level of spectral quality cannot, however, be replicated during DDA/MS when there is a need to generally maximize the number of precursor ions selected for fragmentation. These results therefore indicate that accurate characterizations of low abundance glycopeptides are unlikely to be made in HCD DDA/MS experiments using method (A).
(a) Illustrative HCD derived product ion scans obtained via fragmentation of low abundance tryptic glycopeptides of RNase B (doubly charged ions of glycopeptides I and III of Table 1; HCD collision energy of 42%) and Orbitrap mass analysis during DDA/MS. All b and y ions, intact glycan derived oxonium ions and peptide + fragmented glycan ions observed within the spectra are labeled. (b) Relative abundances of Y1 ions, the most abundant oxonium ions and the most abundant peptide + fragmented glycan ions (excluding Y1 ions) observed in low abundance RNase B derived glycopeptides (all ion counts in MS/MS spectra at normalized levels of <8 × 103); data collected across HCD DDA/MS experiments conducted using normalized collision energies of 27%, 35%, and 42%
The strong likelihood of detecting Y1 ions during HCD product ion scans, as described by Segu and Mechref [56] and as reinforced by the above results, indicates that the possibility remains, however, for accurate glycopeptide characterizations to be made using method (B). Detection of Y1 ions is utilized in step 2a of various analytical workflows [26–28, 48]; their identification may allow base peptide sequences to be ascertained without the need for additional fragment ion matches. The possibility then exists for step 2b to be undertaken via mass differences between base peptides and precursor ion derived monoisotopic masses.
In determining base peptide sequence compositions from the observation of Y1 ions alone, the mass accuracies at which the Y1 ions are observed are of central importance. An inspection of Figure 3a reveals that for the illustrated spectra, NLTK + HexNAc ions are observed at m/z 678.3666 and 678.3693; the respective mass measurement errors of these ions are 0.4 and 3.6 ppm. These values are typical of the Y1 product ions observed from the present sample using HCD DDA/MS in which a RMS error of 3 ppm was obtained, with maximum mass measurement errors of ±5 ppm; these mass measurement errors are consistent with those expected from Orbitrap mass analysis without the application of lock mass [58]. The degree to which this level of mass accuracy can be relied upon to determine base peptide compositions is discussed during evaluation of the workflow applied to the hen egg glycoprotein analysis.
After base peptide compositions have been determined, the characterization of glycan compositions during step 2b is dictated by the mass accuracy of the MS-1 data. For the N-linked glycans under present consideration, it is noteworthy that the <2.5 ppm mass measurement errors in MS-1 are sufficient to produce zero false positives: if a GlycoMod search is conducted upon the experimentally observed glycopeptide masses listed in Table 1 against the known masses of the base peptides using a mass tolerance ±2.5 ppm, only correct compositions are returned. False positives are, however, returned if the GlycoMod search is expanded to consider O-linked glycans; as summarized in Table S1 of the Supplementary Material, false positive glycan compositions are generated for glycopeptides II, V, and VI, 13 false positive glycan compositions in total. These results confirm that accurate mass data alone cannot be relied upon to produce zero false positives during step 2b of the analysis. These results do, however, suggest that the present levels of mass accuracy can result in the generation of relatively small pools of candidate glycan compositions, particularly if only N-linked glycopeptides are being considered. It is feasible that additional information can be used to further refine putative glycan compositional assignments; such information is applied to the workflow used in the hen egg glycoprotein analysis.
3.2 Hen Egg Glycoprotein Mixture Analysis
The RNase B experiments described above indicate that when data is gathered using HCD and Orbitrap mass analysis during DDA/MS, steps 1, 2a, and 2b of glycopeptide characterizations are most likely to be well facilitated through the observation of oxonium marker ions and Y1 ions at high mass accuracy, and the concomitant generation of high mass accuracy MS-1 data. The present hen egg glycoprotein analyses provide a proof of principle that by implementing such data in a glycopeptide characterization workflow, strong outcomes can be achieved in the characterization of complex digest mixtures of previously uncharacterized low abundance glycoproteins. The specific aspects of the employed workflow in relation to steps 2a and 2b are outlined following evaluation of step 1.
Step 1 – Detection of glycopeptide precursor ions
The following experiments confirm that the observation of HCD derived oxonium ions can allow highly selective and sensitive glycopeptide detections to be made from complex peptide mixtures of low abundance peptides. Figure 4 illustrates data obtained from two illustrative HCD DDA/MS experiments conducted upon Glu-C and trypsin hen egg glycoprotein digests. The total number of precursor ions selected for fragmentation and the number of these ions that produce fragments of m/z corresponding to glycan derived oxonium ions (within the m/z windows described in the RNase B analysis) are shown. Of the 241 precursor ions selected for MS/MS in the Glu-C digest, 37 are identified as potential glycopeptides; each of these glycopeptide detections can be confidently confirmed, as described in the following sub-section, and/or via the observation of strong glycan fragment peaks in complementary CID data. In the trypsin digest, of the 425 precursor ions selected for MS/MS, 51 are identified as potential glycopeptides and 50 of these glycopeptide detections can be confidently confirmed. The remaining precursor ion shows strong evidence for the production of doubly charged peptide + fragmented glycan ions at a m/z range beyond the m/z of the doubly charged molecular ion, suggesting that co-fragmentation with a glycopeptide precursor produces the observed oxonium marker ion.
Results of illustrative HCD and Orbitrap mass analysis DDA/MS experiments upon Glu-C and trypsin digests of a hen egg glycoprotein mixture in terms of: total precursor ions selected for fragmentation, precursors identified as potential glycopeptides via observation of fragment ions with m/z corresponding to glycan derived oxonium ions, and correct glycopeptide identifications. Illustrated data collected at 35% for the Glu-C digest sample, and 42% for the trypsin digest sample
The present results reveal that in detecting glycopeptides from complex peptide mixtures using HCD in DDA/MS, HexNAc derived oxonium ions are particularly robust markers for N-linked high mannose glycopeptides (the glycan compositions of which are determined in the following sub-section). Figure 5 takes its data from the entire series of HCD DDA/MS experiments conducted upon the hen egg glycoprotein digest mixtures, which consisted of experiments conducted using normalized collision energies of 27%, 35%, and 42%. In the trypsin digest sample, an average of 57% and 6% of the glycopeptides capable of generating Hex1HexNAc1 and Hex derived oxonium ions do indeed produce these respective ions, whereas all detected glycopeptides produce HexNAc derived oxonium ions. In the Glu-C digest sample, though the production of Hex1HexNAc1 and Hex derived oxonium ions show more variation across the different collision energies employed, qualitatively similar results are nonetheless observed: on average, 46% and 5% of detected glycopeptides produce Hex1HexNAc1 and Hex derived oxonium ions respectively, whilst all detected glycopeptides produce HexNAc derived oxonium ions.
Percentages of product ion spectra in which HexNAc, Hex1HexNAc1, and Hex derived oxonium ions are observed from Gallus gallus ovomucoid and ovoglycoprotein derived high mannose glycopeptides produced via Glu-C or trypsin digestions. Data obtained from HCD spectra collected from DDA/MS experiments using normalized collision energies of 27%, 35%, and 42%
The present results also reveal that glycopeptide detections are readily achieved at low fmol levels. DDA/MS experiments conducted upon 100 fmol of sample detect numerous glycopeptides derived from hen egg glycoproteins other than ovalbumin, which makes up >98% of the mixture under study; the maximum concentration of a non-ovalbumin derived glycopeptide is therefore 2 fmol. Given that certain identified glycopeptides are distributed amongst at least 14 glycoforms (as determined in the following sub-section), it can be concluded that sub-fmol limits of glycopeptide detection are achieved.
Steps 2a and 2b – Characterization of the peptide and glycan portions
The data obtained from the present samples confirm that high confidence glycopeptide compositional characterizations can be made during steps 2a and 2b through the concomitant generation of high mass accuracy Y1 ions and MS-1 data; compositions for numerous previously uncharacterized hen egg glycopeptides are thus determined. These glycopeptides, present at low fmol concentrations, confirm the presence of two previously unreported glycosylation sites on Gallus gallus ovoglycoprotein (amino acids 82 and 90), with significant glycan heterogeneity observed at amino acid 90. Considerable glycan heterogeneities at amino acids 34 and 77 of Gallus gallus ovomucoid are also revealed. Tables 2 and 3 list all of the glycopeptides detected in the current experiments and their final characterizations, as determined using the following workflow: (i) MS-1 derived monoisotopic masses of detected glycopeptides are differentiated, thus allowing series of glycoforms to be recognized; (ii) candidate Y1 ions are identified within the different series of glycoforms; (iii) the masses of Y1 ion candidates are matched against the masses of possible base peptides (obtained via conventional sequence database searching); (iv) pools of glycan compositions are obtained by subtracting the masses of matched base peptides from the monoisotopic masses of the glycopeptide precursor ions; and (v) the confidence of glycan compositional matches are refined by comparing them against the compositions of families of glycoforms obtained from step (i). These steps are elaborated upon below.
During step (i), it can be demonstrated that the present levels of mass accuracy in MS-1 allow glycoforms to be confidently ascertained from the pools of glycopeptides detected via oxonium ion observation; that is, mass differences between glycopeptides corresponding to monosaccharide or oligosaccharide masses are identified. Goldberg et al. have previously demonstrated the utility of an analogous approach to identifying glycoforms using Q-TOF generated data [25]. In the present Glu-C digest mixture, the <2.5 ppm mass measurement errors in MS-1 allows five different series of glycoforms to be clearly identified; in the trypsin digest mixture, eight different series of glycoforms are apparent. Of the 83 distinct glycopeptides detected across the two samples, the present levels of mass accuracy allow these glycoform series to be determined without any ambiguity (i.e., no mass coincidences allowing single glycopeptides to be placed in different series of glycoform are observed). Within each individual series of glycoforms the precursor ions elute at distinct but similar retention times, as listed in Tables 2 and 3 and illustrated in the chromatograms provided in Figures S5 to S16. Precursor ions resulting from in-source fragmentation of a common glycopeptide should have identical retention times; these results therefore indicate that the bulk of the observed microheterogeneity is not an artifact of in-source fragmentation.
Having identified series of glycoforms, Y1 ions, which are observed at identical m/z when related glycoforms are subjected to MS/MS, may then be identified in step (ii); this is possible even when glycopeptides produce weak precursor ion signals and/or poor product ion spectra. Figure 6 shows averaged product ion spectra obtained from the fragmentation of related series of glycoforms, providing an illustration of the principle by which Y1 ion identification may be achieved. Tables 4 and 5 list—for illustrative datasets—the final confidently identified Y1 ions, the relative abundances of these ions, and how these relative abundances rank in relation to other product ions. Qualitatively identical outcomes in Y1 ion identification were achieved using datasets produced using collision energies of 27%, 35%, and 42%. Unlike the RNase B derived data, Y1 ions rarely form the most abundant peaks within the spectra produced from the present glycopeptides; in the vast majority of spectra, the most abundant peaks stem from glycan derived oxonium ions (and product ions associated with their loss of H2O). This is consistent across the 27% to 42% range of collision energies employed in the DDA/MS experiments, and across the various charge states observed for the precursor ions (predominantly 2+ and 3+). As suggested by Segu and Mechref’s study [56], this lowered relative abundance of Y1 ions can likely be attributed to the increased sizes and altered compositions of the peptide backbones associated with the present samples [as determined in step (iii)] relative to the RNase B derived NLTK peptide. Despite this, if low m/z product ions (<m/z 500) are dismissed, it can be seen that, when observed, Y1 ions still typically rank amongst the top two peaks in individual spectra. With few exceptions, even when Y1 ions do not rank as the highest abundance peak above m/z 500, the higher-ranking peaks are poorly reproducible and can be dismissed as candidate Y1 ions. Notably, this remains true for spectra in which Y1 ions are not observed at all. The few non-Y1 ion peaks retained from these analyses are artifacts of the HCD process (m/z 1868.5, 1905.7, 1907.7, and 1974.1) and base peptide peaks. The HCD artifacts, which are observed consistently but with relatively poor reproducibility in m/z, may be immediately dismissed as candidate Y1 ions. For each of the glycoform series, this leaves a maximum of two ions retained as Y1 ion candidates; as can be seen in the examples shown in Figure 6, these candidate ions differ in m/z by 203.08—the mass of a HexNAc unit—providing strong evidence to suggest that the lower m/z peak is the base peptide ion, and the higher m/z peak the Y1 ion. As will be discussed below, step (iii) of the current workflow provides further evidence to rule out base peptide ion peaks as Y1 ion candidates, allowing the true Y1 ions to be confirmed.
By obtaining Y1 ion candidates in step (ii), the possibility then exists for base peptide sequences to be ascertained during step (iii). In broad-scale studies of unknown glycoproteins, this step strongly depends on the ability to determine the proteins present in the sample, thus allowing a list of theoretical peptide masses to be generated, and on the mass accuracy of the Y1 ions. Table 6 lists the proteins and number of peptides identified from the present samples using HCD DDA/MS and CID DDA/MS and the Mascot search parameters listed in Section 2. Though both fragmentation methods allow for the confident identification of three Gallus gallus proteins and thus the generation of identical lists of theoretical peptide masses, the use of CID allows a greater number of unique peptides to be identified, which is consistent with previous reports [59]. In broader-scale studies, the use of CID as a complement to HCD data should therefore be considered if glycoprotein identifications, and thus the overall sensitivity of the analysis, are to be maximized. For the three proteins identified from the present study, theoretical peptide masses were generated with no enzyme cleavage specificity, taking into account the possibility for oxidation of peptide backbone M units. Using the resultant theoretical base peptide masses, for each of the Y1 ion candidates identified from step (ii), no more than one associated base peptide featuring possible sites of N- or O-linked glycosylation can be identified within a mass tolerance of 5 ppm. For each glycoform series in which two Y1 candidates are identified, only the higher m/z ions generate matches, confirming these peaks as the Y1 ions. Though this data provides a proof of principle that highly selective base peptide characterizations can be made using the present techniques, the reproducibility of these results will depend upon the breadth of each study to which these techniques are applied. It is envisaged that the present ±5 ppm levels of mass accuracy in MS/MS should, however, readily allow minimal false positive base peptide characterizations for studies featuring over an order of magnitude more proteins than the current investigation [60–62].
The confident assignment of base peptide sequences during step (iii) allows glycan compositional assignments to be inferred from MS-1 data during step (iv). In the present analyses, each glycopeptide with an assigned base peptide sequence returned only one possible glycan compositional match from GlycoMod within a mass tolerance of ±2.5 ppm, suggesting zero false positive characterizations. These outstanding results are unsurprising given that Desaire and Hua have shown that, even without knowledge of the peptide backbone composition, ±2.5 ppm levels of mass accuracy in MS-1 can allow 86% of the glycopeptides in a complex glycoprotein to be unambiguously characterized; for a simple glycoprotein, this value was determined to be 98% [38]. These final glycopeptide compositional characterizations are further supported by their observed reversed-phase LC behavior: the more polar glycopeptides, according to their putative structures, do indeed elute earlier than the less polar glycopeptides (see data listed in Tables 2 and 3, and illustrated in Figures S5 to S16). Of particular note in this regard, several possibly truncated N-linked glycans, which should be relatively non-polar, are identified to have the longest retention times within their respective glycoform series. These data provide particularly strong evidence to suggest that these unusual glycan structures are indeed present in the samples under analysis, and are not formed via in-source fragmentation. It must be noted, however, that no evidence has been presented to suggest that these glycans are biologically relevant, and the possibility of in vitro glycan truncation cannot be ruled out.
For samples in which ambiguous glycan compositional matches are returned during step (iv), step (v) of the present workflow can allow false positives to be presented as low confidences matches. Though no false positives are returned from the present analyses, the principles of this step can be illustrated using the previously described RNase B data (see Table S1 for details). Three RNase B derived glycopeptides (glycopeptides II, V, and VI of Tables 1 and S1), all confirmed to contain a NLTK backbone, produce ambiguous glycan compositional matches in GlycoMod when using a ±2.5 ppm mass tolerance. The monoisotopic mass differences of these three glycopeptides reveal that they differ in composition by single Hex units only. The false positive glycan compositions represent a departure from this observed relationship; each differs from the other glycopeptides in the series by combinations of monosaccharide units other than Hex, allowing these glycopeptides to be deemed low confidence matches.
For the workflow described above, glycopeptides observed to be in large glycoform series should, in general, be more likely to produce confident matches; the possibility of obtaining confident Y1 ion matches during step (ii) is increased, and greater discriminatory power can be attained during step (v). Though high certainty matches are less likely for glycopeptides observed alone, or in small glycoform series, it should, however, be noted that the compositions of such glycopeptides can still be firmly established in specific instances. For instance the matches obtained for glycoforms of the ovoglycoprotein derived base peptide FNVTE in the Glu-C digest sample can be considered high confidence: ±2.5 ppm mass tolerance GlycoMod searches on the associated monoisotopic precursor ion masses return single hits, the base peptide features no missed cleavages, and the glycan compositions are well established in the GlycoSuiteDB database [63, 64]. The glycoforms of the ovoglycoprotein derived base peptide LNETCVVK in the trypsin digest can be considered high certainty for the same reasons. Within any given study, the likelihood of obtaining similar high confidence matches from glycopeptides observed alone or in small glycoform series will depend on the breadth of the analysis.
4 Conclusions
Progress towards broad-scale, systemic analyses of protein glycosylations will likely involve the integration of various complementary analytical approaches. The present findings suggest that in fulfilling the potential for MS to facilitate such large-scale glycopeptide characterizations, the role of Orbitrap mass analysis and HCD in DDA/MS is likely to be significant. The technique offers the significant advantages of allowing glycopeptides to be selectively detected at sub-fmol limits of detection, and for highly confident compositional characterizations to be made for these glycopeptides. Detailed insights into the compositions and heterogeneity of previously uncharacterized glycopeptides present at low fmol concentrations are thus capable of being gained. These excellent outcomes stem from a reduced requirement for high overall spectral quality, as the observation of a severely limited number of fragment ions at high mass accuracy still results in the potential for high confidence characterizations. This represents a significant advance over conventional CID DDA/MS, in which the fragmentation of glycopeptides at low concentrations can be expected to produce poor quality MS/MS spectra, either precluding the detection of these ions as glycopeptides, or making their characterization difficult. Goldberg et al. bypass this problem in their Peptoonist software by expanding the relatively few high confidence glycopeptide characterizations obtained via CID DDA/MS to other glycopeptide candidates observed during MS-1, thus increasing overall sensitivity [25]. It can be expected that the present methods of data collection should allow for even greater overall levels of sensitivity when this technique is employed, as the potential exists for the initial number of high confidence DDA/MS derived glycopeptide characterizations to be increased when using HCD instead of CID.
When used in isolation, data collection via HCD DDA/MS does, however, feature several limitations. Though highly sensitive individual glycopeptide compositional characterizations are capable of being achieved, the overall sensitivity of a given workflow relying upon DDA/MS can remain limited by MS processes; for instance glycopeptides that ionize poorly, such as those containing negatively charged sialic acid moieties, may be unlikely to be selected for fragmentation during DDA/MS. Additionally, though potential glycan attachment sites can be ascertained through the identification of base peptide sequences, the present results suggest that HCD produces few product ions conducive to the specific localization of glycan attachment sites. Similarly, few glycan fragment ion peaks for glycan structural (as opposed to compositional) determination are observed. In accessing such information, alternate modes of activation during MS/MS such as CID or electron transfer dissociation (ETD), which can be applied alongside HCD in individual DDA/MS experiments, should serve as a complement to the present methods of data collection. Despite these limitations, the particularly strong capabilities of Orbitrap mass analysis and HCD in DDA/MS demonstrated for low abundance glycopeptides suggest that when used in collaboration with complementary technologies, this method of data collection holds great promise to help facilitate robust, systematic, broad-scale characterizations of protein glycosylations.
References
Apweiler, R., Hermjakob, H., Sharon, N.: On the frequency of protein glycosylation, as deduced from analysis of the SWISS-PROT database. Biochim. Biophys. Acta 1473, 4–8 (1999)
Ohtsubo, K., Marth, J.D.: Glycosylation in cellular mechanisms of health and disease. Cell 126, 855–867 (2006)
Dwek, R.A.: Glycobiology: Toward understanding the function of sugars. Chem. Rev. 96, 683–720 (1996)
Spiro, R.G.: Protein glycosylation: Nature, distribution, enzymatic formation, and disease implications of glycopeptide bonds. Glycobiology 12, 43r–56r (2002)
Wuhrer, M., Catalina, M.I., Deelder, A.M., Hokke, C.H.: Glycoproteomics based on tandem mass spectrometry of glycopeptides. J. Chromatogr. B 849, 115–128 (2007)
Dalpathado, D.S., Desaire, H.: Glycopeptide analysis by mass spectrometry. Analyst 133, 731–738 (2008)
Dell, A., Morris, H.R.: Glycoprotein structure determination by mass spectrometry. Science 291, 2351–2356 (2001)
North, S.J., Hitchen, P.G., Haslam, S.M., Dell, A.: Mass spectrometry in the analysis of N-linked and O-linked glycans. Curr. Opin. Struct. Biol. 19, 498–506 (2009)
Morelle, W., Canis, K., Chirat, F., Faid, V., Michaelski, J.C.: The use of mass spectrometry for the proteomic analysis of glycosylation. Proteomics 6, 3993–4015 (2006)
Morelle, W., Michaelski, J.C.: Analysis of protein glycosylation by mass spectrometry. Nat Protoc 2, 1585–1602 (2007)
Morris, H.R., Chalabi, S., Panico, M., Sutton-Smith, M., Clark, G.F., Goldberg, D., Dell, A.: Glycoproteomics: past, present and future. Int. J. Mass spectrom. 259, 16–31 (2007)
Temporini, C., Calleri, E., Massolini, G., Caccialanza, G.: Integrated analytical strategies for the study of phosphorylation and glycosylation in proteins. Mass Spectrom. Rev. 27, 207–236 (2008)
Liu, T., Qian, W.J., Gritsenko, M.A., Camp, D.G.I., Monroe, M.E., Moore, R.J., Smith, R.D.: Human plasma N-glycoproteome analysis by immunoaffinity subtraction, hydrazide chemistry, and mass spectrometry. J. Proteome Res. 4, 2070–2080 (2005)
Pan, S., Wang, Y., Quinn, J.F., Peskind, E.R., Waichunas, D., Wimberger, J.T., Jin, J., Li, J.G., Zhu, D., Pan, C., Zhang, J.: Identification of glycoproteins in human cerebrospinal fluid with a complementary proteomic approach. J. Proteome Res. 5, 2769–2779 (2006)
Lewandrowski, U., Moebius, J., Walter, U., Sickmann, A.: Elucidation of N-glycosylation sites on human platelet proteins. Mol. Cell. Proteom. 5, 226–233 (2006)
Ramachandran, P., Boontheung, P., Xie, Y., Sondej, M., Wong, D.T., Loo, J.A.: Identification of N-linked glycoproteins in human saliva by glycoprotein capture and mass spectrometry. J. Proteome Res. 5, 1493–1503 (2006)
Bernhard, O.K., Kapp, E.A., Simpson, R.J.: Enhanced analysis of the mouse plasma proteome using cysteine-containing tryptic glycopeptides. J. Proteome Res. 6, 987–995 (2007)
Juhasz, P., Martin, S.A.: The utility of nonspecific proteases in the characterization of glycoproteins by high-resolution time-of-flight mass spectrometry. Int. J. Mass spectrom. 169, 217–230 (1997)
An, H.J., Peavy, T.R., Hedrick, J.L., Lebrilla, C.B.: Determination of N-glycosylation sites and site heterogeneity in glycoproteins. Anal. Chem. 75, 5628–5637 (2003)
Wuhrer, M., Koeleman, C.A.M., Hokke, C.H., Deelder, A.M.: Protein glycosylation analyzed by normal-phase nano-liquid chromatography-mass spectrometry of glycopeptides. Anal. Chem. 77, 886–894 (2005)
Temporini, C., Perani, E., Calleri, E., Dolcini, L., Lubda, D., Caccialanza, G., Massolini, G.: Pronase-immobilized enzyme reactor: an approach for automation in glycoprotein analysis by LC/LS-ESI/MSn. Anal. Chem. 79, 355–363 (2007)
Clowers, B.H., Dodds, E.D., Seipert, R.R., Lebrilla, C.B.: Site determination of protein glycosylation based on digestion with immobilized nonspecific proteases and Fourier transform ion cyclotron resonance mass spectrometry. J. Proteome Res. 6, 4032–4040 (2007)
Yu, Y.Q., Fournier, J., Gilar, M., Gebler, J.C.: Identification of N-linked glycosylation sites using glycoprotein digestion with pronase prior to MALDI tandem time-of-flight mass spectrometry. Anal. Chem. 79, 1731–1738 (2007)
Dodds, E.D., Seipert, R.R., Clowers, B.H., German, J.B., Lebrilla, C.B.: Analytical performance of immobilized pronase for glycopeptide footprinting and implications for surpassing reductionist glycoproteomics. J. Proteome Res. 8, 502–512 (2009)
Goldberg, D., Bern, M., Parry, S., Sutton-Smith, M., Panico, M.: Automated N-glycopeptide identification using a combination of single- and tandem-MS. J. Proteome Res. 6, 3995–4005 (2007)
Ozohanics, O., Krenyacz, J., Ludanyi, K., Pollreisz, F., Vekey, K., Drahos, L.: GlycoMiner: a new software tool to elucidate glycopeptide composition. Rapid Commun. Mass Spectrom. 22, 3245–3254 (2008)
Joenväärä, S., Ritamo, I., Peltoniemi, H., Renkonen, R.: N-glycoproteomics—an automated workflow approach. Glycobiology 18, 339–349 (2008)
Peltoniemi, H., Joenväärä, S., Renkonen, R.: De novo glycan structure search with the CID MS/MS spectra of native N-glycopeptides. Glycobiology 19, 707–714 (2009)
Carr, S.A., Huddleston, M.J., Bean, M.F.: Selective identification and differentiation of N- and O-linked oligosaccharides in glycoproteins by liquid chromatography-mass spectrometry. Protein Sci. 2, 183–196 (1993)
Conboy, J.J., Henion, J.D.: The determination of glycopeptides by liquid chromatography/mass spectrometry with collision-induced dissociation. J. Am. Soc. Mass. Spectrom. 3, 804–814 (1992)
Huddleston, M.J., Bean, M.F., Carr, S.A.: Collision fragmentation of glycopeptides by electrospray ionization LC/MS and LC/MS/MS: methods for selective detection of glycopeptides in protein digests. Anal. Chem. 65, 877–884 (1993)
Krokhin, O., Ens, W., Standing, K.G., Wilkins, J., Perreault, H.: Site-specific N-glycosylation analysis: Matrix-assisted laser desorption/ionization quadrupole-quadrupole time-of-flight tandem mass spectral signatures for recognition and identification of glycopeptides. Rapid Commun. Mass Spectrom. 18, 2020–2030 (2004)
Bykova, N.V., Rampitsch, C., Krokhin, O., Standing, K.G., Ens, W.: Determination and characterization of site-specific N-glycosylation using MALDI-Qq-TOF tandem mass spectrometry: Case study with a plant protease. Anal. Chem. 78, 1093–1103 (2006)
Jebanathirajah, J., Steen, H., Roepstorff, P.: Using optimized collision energies and high resolution, high accuracy fragment ion selection to improve glycopeptide detection by precursor ion scanning. J. Am. Soc. Mass. Spectrom. 14, 777–784 (2003)
Sullivan, B., Addona, T.A., Carr, S.A.: Selective detection of glycopeptides on ion trap mass spectrometers. Anal. Chem. 76, 3112–3118 (2004)
Peterman, S.M., Mulholland, J.J.: A novel approach for identification and characterization of glycoproteins using a hybrid linear ion trap/FT-ICR mass spectrometer. J. Am. Soc. Mass Spectrom. 17, 168–179 (2006)
Cooper, C.A., Gasteiger, E., Packer, N.H.: GlycoMod—a software tool for determining glycosylation compositions from mass spectrometric data. Proteomics 1, 340–349 (2001)
Desaire, H., Hua, D.: When can glycopeptides be assigned based solely on high-resolution mass spectrometry data? Int. J. Mass spectrom 287, 21–26 (2009)
Demelbauer, U.M., Zehl, M., Plematl, A., Allmaier, G., Rizzi, A.: Determination of glycopeptide structures by multistage mass spectrometry with low-energy collision-induced dissociation: Comparison of electrospray ionization quadrupole ion trap and matrix-assisted laser desorption/ionization quadrupole ion trap reflectron time-of-flight approaches. Rapid Commun. Mass Spectrom. 18, 1575–1582 (2004)
Wuhrer, M., Balog, C.I.A., Catalina, M.I., Jones, F.M., Schramm, G., Haas, H., Doenhoff, M.J., Dunne, D.W., Deelder, A.M., Hokke, C.H.: IPSE/↦-1, a major secretory glycoprotein antigen from schistosome eggs, expresses the Lewis X motif on core-difucosylated N-glycans. FEBS J. 273, 2276–2292 (2006)
Mikesh, L.M., Ueberheide, B., Chi, A., Coon, J.J., Syka, J.E.P., Shabanowitz, J., Hunt, D.F.: The utility of ETD mass spectrometry in proteomic analysis. Biochim. Biophys. Acta 1764, 1811–1822 (2006)
Hogan, J.M., Pitteri, S.J., Chrisman, P.A., McLuckey, S.A.: Complementary structural information from a tryptic N-linked glycopeptide via electron transfer ion/ion reactions and collision-induced dissociation. J. Proteome Res. 4, 628–632 (2005)
Hakansson, K., Cooper, H.J., Emmett, M.R., Costello, C.E., Marshall, A.G., Nilsson, C.L.: Electron capture dissociation and infrared multiphoton dissociation MS/MS of an N-glycosylated tryptic peptic to yield complementary sequence information. Anal. Chem. 73, 4530–4536 (2001)
Hakansson, K., Chalmers, M.J., Quinn, J.F., McFarland, M.A., Hendrickson, C.L., Marshall, A.G.: Combined electron capture and infrared multiphoton dissociation for multistage MS/MS in a Fourier transform ion cyclotron resonance mass spectrometer. Anal. Chem. 75, 3256–3262 (2003)
Alley, W.R., Mechref, Y., Novotny, M.V.: Characterization of glycopeptides by combining collision-induced dissociation and electron-transfer dissociation mass spectrometry data. Rapid Commun. Mass Spectrom. 23, 161–170 (2009)
Perdivara, I., Petrovich, R., Allinquant, B., Deterding, L.J., Tomer, K.B., Pryzbylski, M.: Elucidation of O-glycosylation structures of the β-amyloid precursor protein by liquid chromatography-mass spectrometry using electron transfer dissociation and collision induced dissociation. J. Proteome Res. 8, 631–642 (2009)
Ritchie, M.A., Gill, A.C., Deery, M.J., Lilley, K.: Precursor ion scanning for detection and structural characterization of heterogeneous glycopeptide mixtures. J. Am. Soc. Mass. Spectrom. 13, 1065–1077 (2002)
Irunga, J., Go, E.P., Dalpathado, D.S., Desaire, H.: Simplification of mass spectral analysis of acidic glycopeptides using GlycoPep ID. Anal. Chem. 79, 3065–3074 (2007)
Ren, J.M., Rejtar, T., Li, L., Karger, B.L.: N-glycan structure annotation of glycopeptides using a linearized glycan structure database (GlyDB). J. Proteome Res. 6, 3162–3173 (2007)
Shan, B., Ma, B., Zhang, K., Lajoie, G.: Complexities and algorithms for glycan sequencing using tandem mass spectrometry. J. Bioinf. Comput. Biol. 6, 77–91 (2008)
Han, X., Aslanian, A., Yates, J.R.I.: Mass spectrometry for proteomics. Curr. Opin. Chem. Biol. 12, 483–490 (2008)
Perry, R.H., Cooks, R.G., Noll, R.J.: Orbitrap mass spectrometry: instrumentation, ion motion and applications. Mass Spectrom. Rev. 27, 661–699 (2008)
Yates, J.R.I., Ruse, C.I., Nakorchevsky, A.: Proteomics by mass spectrometry: approaches, advances, and applications. Annu. Rev. Biomed. Eng. 11, 49–79 (2009)
Kuzmanov, U., Jiang, N., Smith, C.R., Soosaipillai, A., Diamandis, E.P.: Differential N-glycosylation of kallikrein 6 derived from ovarian cancer cells or the central nervous system. Mol. Cell. Proteom. 8, 791–798 (2009)
Olsen, J.V., Macek, B., Lange, O., Makarov, A., Horning, S., Mann, M.: Higher-energy C-trap dissociation for peptide modification analysis. Nat. Methods 4, 709–712 (2007)
Segu, Z.M., Mechref, Y.: Characterizing protein glycosylation sites through higher-energy C-trap dissociation. Rapid Commun. Mass Spectrom. 24, 1217–1225 (2010)
Gatlin, C.L., Kleemann, G.R., Hays, L.G., Link, A.J., Yates, J.R.I.: Protein identification at the low femtomole level from silver-stained gels using a new fritless electrospray interface for liquid chromatography microspray and nanospray mass spectrometry. Anal. Biochem. 263, 93–101 (1998)
Olsen, J.V., de Godoy, L.M.F., Li, G., Macek, B., Mortensen, P., Pesch, R., Makarov, A.A., Lange, O., Horning, S., Mann, M.: Parts per million mass accuracy on an Orbitrap mass spectrometer via lock mass injection into a C-trap. Mol. Cell. Proteom. 4, 2010–2021 (2005)
Scherl, A., Shaffer, S.A., Taylor, G.K., Hernandez, P., Appel, R.D., Binz, P.A., Goodlett, D.R.: On the benefits of acquiring peptide fragment ions at high measured mass accuracy. J. Am. Soc. Mass Spectrom. 19, 891–901 (2008)
Clauser, K.R., Baker, P., Burlingame, A.L.: Role of accurate mass measurement (±10 ppm) in protein identification strategies employing MS or MS/MS and database searching. Anal. Chem. 71, 287–2882 (1999)
Conrads, T.P., Anderson, G.A., Veenstra, T.D., Paša-Tolić, L., Smith, R.D.: Utility of accurate mass tags for proteome-wide protein identification. Anal. Chem. 72, 3349–3354 (2000)
Eriksson, J., Chait, B.T., Fenyő, D.: A statistical basis for testing the significance of mass spectrometric protein identification results. Anal. Chem. 72, 999–1005 (2000)
Cooper, C.A., Harrison, M.J., Wilkins, M.R., Packer, N.H.: GlycoSuiteDB: a new curated relational database of glycoprotein glycan structures and their biological sources. Nucleic Acids Res. 29, 332–335 (2001)
Cooper, C.A., Joshi, H.J., Harrison, M.J., Wilkins, M.R., Packer, N.H.: GlycoSuiteDB: a curated relational database of glycoprotein glycan structures and their biological sources. 2003 update. Nucleic Acids Res 31, 511–513 (2003)
Acknowledgments
The authors acknowledge support for this work in part by the Faculty of Medicine (UNSW) and grants from the Australian Research Council (Cancer Institute NSW, National Collaborative Research Infrastructure Scheme).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(DOC 827 kb)
Rights and permissions
About this article
Cite this article
Hart-Smith, G., Raftery, M.J. Detection and Characterization of Low Abundance Glycopeptides Via Higher-Energy C-Trap Dissociation and Orbitrap Mass Analysis. J. Am. Soc. Mass Spectrom. 23, 124–140 (2012). https://doi.org/10.1007/s13361-011-0273-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13361-011-0273-y