Abstract
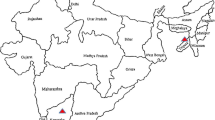
Samples from toria plants (Brassica rapa L. subsp. dichotoma (Roxb.)) exhibiting phyllody, virescence, witches broom, extensive malformation of floral parts, formation of bladder like siliquae and flower sterility were collected from four different locations in India. Sequencing and phylogenetic analysis of the 16S rRNA, a part of 23S rRNA, partial sec A genes, rp gene and 16S–23S intergenic spacer region indicated that the phytoplasmas associated with toria phyllody (TP) symptoms were identical and belonged to 16SrIX phytoplasma Pigeon pea witches’-broom (PPWB) group. The iPhyClassifier generated virtual RFLP pattern of 1.25 kb 16S rDNA sequences indicated that TP phytoplasma belongs to 16SrIX-C phytoplasma subgroup. Complete 23S rRNA gene of TP phytoplasma had 2,787 nucleotides and is the first sequence of 16SrIX phytoplasma group. Restriction digestion of 16S rDNA and 23S rDNA PCR products has also shown that TP phytoplasmas from all the four locations in India were identical. Toria is a previously unreported host for a phytoplasma in16SrIX-C subgroup.





Similar content being viewed by others
References
Arnaud G, Malembic-Maher S, Salar P, Bonnet P, Maixner M, Marcone C, Boudon-Padieu E, Foissac X. Multilocus sequence typing confirms the close genetic interrelatedness of three distinct flavescence dore’e phytoplasma strain clusters and group 16SrV phytoplasmas infecting grapevine and alder in Europe. Appl Environ Microbiol. 2007;73:4001–10.
Azadvar M (2010). Biological and molecular characterization of phytoplasma associated with Brassica rapa in India. PhD Thesis, Indian Agricultural Research Institute, New Delhi.
Azadvar M, Baranwal VK, Yadava DK. First report of a 16SrIX (Pigeon pea witches’ broom) phytoplasma associated with toria (Brassica rapa cv. toria) phyllody disease in India. New Dis Rep. 2009;20:27.
Bertaccini A, Vorácková Z, Vibio M, Fránová J, Navrátil M, Špak J, Nebesárová J. Comparison of phytoplasmas infecting winter oilseed rape in the Czech Republic with Italian Brassica phytoplasmas and their relationship to the aster yellows group. Plant Pathol. 1998;47:317–24.
Bhowmik TP. Oilseed Brassicas constraints and their management. New Delhi: CBS Publishers and Distributors; 2003, p. 254.
Bindra OS, Bakhetia DRC. A note on natural incidence of sesamum phyllody virus disease in Brassica spp. Ludhiana J Res. 1967;4:406–8.
Botti S, Bertaccini A. Variability and functional role of chromosomal sequences in 16SrI-B subgroup phytoplasmas including aster yellows and related strains. J Appl Microbiol. 2003;94:103–10.
Davis RE, Dally E, Zhao Y, Lee IM, Jomantiene R, Detweiler AJ, Putnam ML. First report of a new subgroup 16SrIX-E, ‘Candidatus Phytoplasma phoenicium’-related, phytoplasma associated with juniper witches’ broom disease in Oregon. New Dis Rep. 2009;20:35.
Firrao G, Gibb K, Streten C. Short taxonomic guide to the genus ‘Candidatus phytoplasma’. J Plant Pathol. 2005;87:249–263
Guo VH, Cheng ZM, Walla JA. Amplification of the 23S rRNA gene and its application in differentiation and detection of phytoplasmas. Can J Plant Pathol. 2000;22:380–6.
Hall TA. BioEdit: a user-friendly biological sequence alignment editor and analysis program for windows 95/98/NT. Nucleic Acids Symp Ser. 1999;41:95–8.
Hodgetts J, Boonham N, Mumford R, Harrison N, Dickinson M. Phytoplasma phylogenetics based on analysis of secA and 23S rRNA gene sequences for improved resolution of candidate species of ‘Candidatus Phytoplasma’. Int J Syst Evol Microbiol. 2008;58:1826–37.
Kaushik CD, Tripathi NN, Vir S. Effect of date of sowing on toria phyllody incidence and estimation of losses. Haryana Agric Univ J Res. 1978;8:28–30.
Lee IM, Zhao Y, Bottner KD. SecY gene sequence analysis for finer differentiation of diverse strains in the aster yellows phytoplasma group. Mol Cell Probes. 2006;20:87–91.
Lee IM, Zhao Y, Davis RE. Prospects to multiple gene-based systems for differentiation and classification of phytoplasmas. In: Weintraub PG, Jones P, editors. Phytoplasmas genomes, plant hosts and vectors. Wallingford: CAB International; 2010. p. 51–63.
Maliogka VI, Tsialtas JT, Papantoniou A, Efthimiou K, Katis NI. First report of a phytoplasma associated with an oilseed rape disease in Greece. Plant Pathol. 2009;58:792.
Marcone C, Lee IM, Davis RE, Ragozzino A, Seemüller E. Classification of aster yellows-group phytoplasmas based on combined analyses of rRNA and tuf gene sequences. Int J Syst Evol Microbiol. 2000;50:1703–13.
Marcone C, Ragozzino C, Camele I, Rana GL, Seemüller E. Updating and extending genetic characterization and classification of phytoplasmas from wild and cultivated plants in southern Italy. J Plant Pathol. 2001;83:133–8.
Martini M, Lee IM, Bottner KD, Zhao Y, Botti S, Bertaccini A, Harrison NA, Carraro L, Marcone C, Khan AJ, Osler R. Ribosomal protein gene based phylogeny for finer differentiation and classification of phytoplasmas. Int J Syst Evol Microbiol. 2007;57:2037–51.
Olivier CY, Séguin-Swartz G, Hegedus D. First report of “Candidatus Phytoplasma asteris” related strains in Brassica rapa in Saskatchewan, Canada. Plant Dis. 2006;90:832.
Salehi M, Izadpanah K, Heydarnejad J. Characterization of a new almond witches’-broom phytoplasma in Iran. J Phytopathol. 2006;154:386–91.
Sambrook J, Russell DW. Molecular cloning: a laboratory manual. Cold Spring Harbor: Cold Spring Harbor Laboratory; 2001.
Schneider B, Seemüller E, Smart CD, Kirkpatrick BC. Phylogenetic classification of plant pathogenic mycoplasma like organisms or phytoplasmas. In: Razin R, Tully JG, editors. Molecular and diagnostic procedures in mycoplasmology, vol. I. San Diego: Academic Press; 1995. p. 369–80.
Streten C, Gibb KS. Genetic variation in ‘Candidatus Phytoplasma australiense’. Plant Pathol. 2005;54:8–14.
Tamura K, Dudley J, Nei M, Kumar S. MEGA 4.0: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Mol Biol Evol. 2007;24:1596–9.
Thompson JD, Higgins DG, Gibson TJ. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, positions-specific gap penalties, and weight matrix choice. Nucleic Acids Res. 1994;22:4673–80.
Verdin E, Salar P, Danet JL, Choueiri E, Jreijiri F, El Zammar S, Gélie B, Bové JM, Garnier M. ‘Candidatus Phytoplasma phoenicium’ sp. nov., a novel phytoplasma associated with an emerging lethal disease of almond trees in Lebanon and Iran. Int J Syst Evol Microbiol. 2003;53:833–8.
Wang K, Hiruki C. Molecular characterization and classification of phytoplasmas associated with canola yellows and a new phytoplasma strain associated with dandelions. Plant Dis. 2001;85:76–9.
Wei W, Davis RE, Lee IM, Zhao Y. Computer-simulated RFLP analysis of 16S rRNA genes: identification of ten new phytoplasma groups. Int J Syst Evol Microbiol. 2007;57:1855–67.
Zhao Y, Wei W, Lee IM, Shao J, Suo X, Davis RE. Construction of an interactive online phytoplasma classification tool, iPhyClassifier, and its application in analysis of the peach X-disease phytoplasma group (16SrIII). Int J Syst Evol Microbiol. 2009;59:2582–93.
Zhao Y, Wei W, Davis RE, Lee IM. Recent advances in 16S rRNA gene-based phytoplasma differentiation, classification and taxonomy. In: Weintraub PG, Jones P, editors. Phytoplasmas genomes, plant hosts and vectors. Wallingford: CAB International; 2010. p. 64–92.
Acknowledgment
We thank Dr. R.K. Jain, Head Division of Plant Pathology, for providing the facilities and Dr. D.K. Yadava, Division of Genetics, IARI, New Delhi for his kind help in sample collection.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Azadvar, M., Baranwal, V.K. Molecular Characterization and Phylogeny of a Phytoplasma Associated with Phyllody Disease of toria (Brassica rapa L. subsp. dichotoma (Roxb.)) in India. Indian J. Virol. 21, 133–139 (2010). https://doi.org/10.1007/s13337-011-0023-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13337-011-0023-6