Abstract
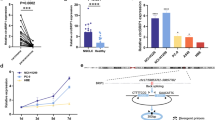
Thyroid cancer (TC) is the most common endocrine malignancy and its incidence has increased over the last few decades. As has been revealed by a number of studies, TC tissue’s micro-RNA (miRNA) profile may reflect histological features and the clinical behavior of tumor. However, alteration of the miRNA profile of plasma exosomes associated with TC development has to date not been explored. We isolated exosomes from plasma and assayed their characteristics using laser diffraction particle size analysis, atomic force microscopy, and western blotting. Next, we profiled cancer-associated miRNAs in plasma exosomes obtained from papillary TC patients, before and after surgical removal of the tumor. The diagnostic value of selected miRNAs was evaluated in a large cohort of patients displaying different statuses of thyroid nodule disease. MiRNA assessment was performed by RT-qPCR. In total, 60 patients with different types of thyroid nodal pathology were included in the study. Our results revealed that the development of papillary TC is associated with specific changes in exosomal miRNA profiles; this phenomenon can be used for differential diagnostics. MiRNA-31 was found to be over-represented in the plasma exosomes of patients with papillary TC vs. benign tumors, while miRNA-21 helped to distinguish between benign tumors and follicular TC. MiRNA-21 and MiRNA-181a-5p were found to be expressed reciprocally in the exosomes of patients with papillary and follicular TC, and their comparative assessment may help to distinguish between these types of TC with 100 % sensitivity and 77 % specificity.






Similar content being viewed by others
References
Gharib H. Changing trends in thyroid practice: understanding nodular thyroid disease. Endocr Pract. 2004;10(1):31–9. doi:10.4158/EP.10.1.31.
Dean DS, Gharib H. Epidemiology of thyroid nodules. Best Pract Res Clin Endocrinol Metab. 2008;22(6):901–11. doi:10.1016/j.beem.2008.09.019.
Kato MA, Fahey 3rd TJ. Molecular markers in thyroid cancer diagnostics. Surg Clin North Am. 2009;89(5):1139–55. doi:10.1016/j.suc.2009.06.012.
Wang CC, Friedman L, Kennedy GC, Wang H, Kebebew E, Steward DL, et al. A large multicenter correlation study of thyroid nodule cytopathology and histopathology. Thyroid. 2011;21(3):243–51. doi:10.1089/thy.2010.0243.
Yip L, Kelly L, Shuai Y, Armstrong MJ, Nikiforov YE, Carty SE, et al. MicroRNA signature distinguishes the degree of aggressiveness of papillary thyroid carcinoma. Ann Surg Oncol. 2011;18(7):2035–41. doi:10.1245/s10434-011-1733-0.
Di Leva G, Garofalo M, Croce CM. MicroRNAs in cancer. Annu Rev Pathol. 2014;9:287–314. doi:10.1146/annurev-pathol-012513-104715.
Lee JC, Gundara JS, Glover A, Serpell J, Sidhu SB. MicroRNA expression profiles in the management of papillary thyroid cancer. Oncologist. 2014;19(11):1141–7. doi:10.1634/theoncologist.2014-0135.
Cancer Genome Atlas Research N. Integrated genomic characterization of papillary thyroid carcinoma. Cell. 2014;159(3):676–90. doi:10.1016/j.cell.2014.09.050.
Dettmer MS, Perren A, Moch H, Komminoth P, Nikiforov YE, Nikiforova MN. MicroRNA profile of poorly differentiated thyroid carcinomas: new diagnostic and prognostic insights. J Mol Endocrinol. 2014;52(2):181–9. doi:10.1530/JME-13-0266.
Zhang Y, Zhong Q, Chen X, Fang J, Huang Z. Diagnostic value of microRNAs in discriminating malignant thyroid nodules from benign ones on fine-needle aspiration samples. Tumour Biol. 2014;35(9):9343–53. doi:10.1007/s13277-014-2209-1.
Schwarzenbach H, Nishida N, Calin GA, Pantel K. Clinical relevance of circulating cell-free microRNAs in cancer. Nat Rev Clin Oncol. 2014;11(3):145–56. doi:10.1038/nrclinonc.2014.5.
Yu S, Liu Y, Wang J, Guo Z, Zhang Q, Yu F, et al. Circulating microRNA profiles as potential biomarkers for diagnosis of papillary thyroid carcinoma. J Clin Endocrinol Metab. 2012;97(6):2084–92. doi:10.1210/jc.2011-3059.
Lee JC, Zhao JT, Clifton-Bligh RJ, Gill A, Gundara JS, Ip JC, et al. MicroRNA-222 and microRNA-146b are tissue and circulating biomarkers of recurrent papillary thyroid cancer. Cancer. 2013;119(24):4358–65. doi:10.1002/cncr.28254.
Cantara S, Pilli T, Sebastiani G, Cevenini G, Busonero G, Cardinale S, et al. Circulating miRNA95 and miRNA190 are sensitive markers for the differential diagnosis of thyroid nodules in a Caucasian population. J Clin Endocrinol Metab. 2014;99(11):4190–8. doi:10.1210/jc.2014-1923.
Lee YS, Lim YS, Lee JC, Wang SG, Park HY, Kim SY, et al. Differential expression levels of plasma-derived miR-146b and miR-155 in papillary thyroid cancer. Oral Oncol. 2015;51(1):77–83. doi:10.1016/j.oraloncology.2014.10.006.
Sato-Kuwabara Y, Melo SA, Soares FA, Calin GA. The fusion of two worlds: non-coding RNAs and extracellular vesicles—diagnostic and therapeutic implications (review). Int J Oncol. 2015;46(1):17–27. doi:10.3892/ijo.2014.2712.
Chen WX, Cai YQ, Lv MM, Chen L, Zhong SL, Ma TF, et al. Exosomes from docetaxel-resistant breast cancer cells alter chemosensitivity by delivering microRNAs. Tumour Biol. 2014;35(10):9649–59. doi:10.1007/s13277-014-2242-0.
Hannafon BN, Carpenter KJ, Berry WL, Janknecht R, Dooley WC, Ding WQ. Exosome-mediated microRNA signaling from breast cancer cells is altered by the anti-angiogenesis agent docosahexaenoic acid (DHA). Mol Cancer. 2015;14:133. doi:10.1186/s12943-015-0400-7.
Melo SA, Sugimoto H, O’Connell JT, Kato N, Villanueva A, Vidal A, et al. Cancer exosomes perform cell-independent microRNA biogenesis and promote tumorigenesis. Cancer Cell. 2014;26(5):707–21. doi:10.1016/j.ccell.2014.09.005.
Ogata-Kawata H, Izumiya M, Kurioka D, Honma Y, Yamada Y, Furuta K, et al. Circulating exosomal microRNAs as biomarkers of colon cancer. PLoS One. 2014;9(4):e92921. doi:10.1371/journal.pone.0092921.
Fujita Y, Kuwano K, Ochiya T, Takeshita F. The impact of extracellular vesicle-encapsulated circulating microRNAs in lung cancer research. BioMed Res Int. 2014;2014:486413. doi:10.1155/2014/486413.
Que R, Ding G, Chen J, Cao L. Analysis of serum exosomal microRNAs and clinicopathologic features of patients with pancreatic adenocarcinoma. World J Surg Oncol. 2013;11:219. doi:10.1186/1477-7819-11-219.
Zoller M. Pancreatic cancer diagnosis by free and exosomal miRNA. World J Gastrointest Pathophysiol. 2013;4(4):74–90. doi:10.4291/wjgp.v4.i4.74.
Whiteside TL. The potential of tumor-derived exosomes for noninvasive cancer monitoring. Expert Rev Mol Diagn. 2015;1293:310.
Lee JC, Zhao JT, Gundara J, Serpell J, Bach LA, Sidhu S. Papillary thyroid cancer-derived exosomes contain miRNA-146b and miRNA-222. J Surg Res. 2015;196(1):39–48. doi:10.1016/j.jss.2015.02.027.
Mestdagh P, Van Vlierberghe P, De Weer A, Muth D, Westermann F, Speleman F, et al. A novel and universal method for microRNA RT-qPCR data normalization. Genome Biol. 2009;10(6):R64. doi:10.1186/gb-2009-10-6-r64.
Benes V, Castoldi M. Expression profiling of microRNA using real-time quantitative PCR, how to use it and what is available. Methods. 2010;50(4):244–9. doi:10.1016/j.ymeth.2010.01.026.
Ferraz C, Lorenz S, Wojtas B, Bornstein SR, Paschke R, Eszlinger M. Inverse correlation of miRNA and cell cycle-associated genes suggests influence of miRNA on benign thyroid nodule tumorigenesis. J Clin Endocrinol Metab. 2013;98(1):E8–16. doi:10.1210/jc.2012-2564.
Suresh R, Sethi S, Ali S, Giorgadze T, Sarkar FH. Differential expression of MicroRNAs in papillary thyroid carcinoma and their role in racial disparity. J Cancer Sci Ther. 2015;7(5):145–54. doi:10.4172/1948-5956.1000340.
Xiong Y, Kotian S, Zeiger MA, Zhang L, Kebebew E. miR-126-3p inhibits thyroid cancer cell growth and metastasis, and is associated with aggressive thyroid cancer. PLoS One. 2015;10(8):e0130496. doi:10.1371/journal.pone.0130496.
Boufraqech M, Zhang L, Jain M, Patel D, Ellis R, Xiong Y, et al. miR-145 suppresses thyroid cancer growth and metastasis and targets AKT3. Endocr Relat Cancer. 2014;21(4):517–31. doi:10.1530/ERC-14-0077.
Gu Y, Li D, Luo Q, Wei C, Song H, Hua K, et al. MicroRNA-145 inhibits human papillary cancer TPC1 cell proliferation by targeting DUSP6. Int J Clin Exp Med. 2015;8(6):8590–8.
Acunzo M, Romano G, Wernicke D, Croce CM. MicroRNA and cancer—a brief overview. Adv Biol Regul. 2015;57:1–9. doi:10.1016/j.jbior.2014.09.013.
Kosaka N. Decoding the secret of cancer by means of extracellular vesicles. J Clin Med. 2016;5(2). doi:10.3390/jcm5020022.
Swierniak M, Wojcicka A, Czetwertynska M, Stachlewska E, Maciag M, Wiechno W, et al. In-depth characterization of the microRNA transcriptome in normal thyroid and papillary thyroid carcinoma. J Clin Endocrinol Metab. 2013;98(8):E1401–9. doi:10.1210/jc.2013-1214.
Nikiforova MN, Tseng GC, Steward D, Diorio D, Nikiforov YE. MicroRNA expression profiling of thyroid tumors: biological significance and diagnostic utility. J Clin Endocrinol Metab. 2008;93(5):1600–8. doi:10.1210/jc.2007-2696.
Agretti P, Ferrarini E, Rago T, Candelieri A, De Marco G, Dimida A, et al. MicroRNA expression profile helps to distinguish benign nodules from papillary thyroid carcinomas starting from cells of fine-needle aspiration. Eur J Endocrinol. 2012;167(3):393–400. doi:10.1530/EJE-12-0400.
Whiteside TL. The potential of tumor-derived exosomes for noninvasive cancer monitoring. Expert Rev Mol Diagn. 2015;15(10):1293–310. doi:10.1586/14737159.2015.1071666.
Lowry MC, Gallagher WM, O’Driscoll L. The role of exosomes in breast cancer. Clin Chem. 2015;61(12):1457–65. doi:10.1373/clinchem.2015.240028.
Taverna S, Giallombardo M, Gil-Bazo I, Carreca AP, Castiglia M, Chacartegui J, et al. Exosomes isolation and characterization in serum is feasible in non-small cell lung cancer patients: critical analysis of evidence and potential role in clinical practice. Oncotarget. 2016. 10.18632/oncotarget.7638.
Chevillet JR, Kang Q, Ruf IK, Briggs HA, Vojtech LN, Hughes SM, et al. Quantitative and stoichiometric analysis of the microRNA content of exosomes. Proc Natl Acad Sci U S A. 2014;111(41):14888–93. doi:10.1073/pnas.1408301111.
Koga K, Matsumoto K, Akiyoshi T, Kubo M, Yamanaka N, Tasaki A, et al. Purification, characterization and biological significance of tumor-derived exosomes. Anticancer Res. 2005;25(6A):3703–7.
Lozupone F, Kont V, Logozzi A, Talpsepp K, Oja T, Kubo A, et al. TM9SF4 level of expression on exosomes as new marker of malignancy in human cancer. First scientific meeting of ISEV-International Society for Extracellular Vesicles April 18-21 2012. Gothenburg: University of Gothenburg; 2012. p. 19.
Melo SA, Luecke LB, Kahlert C, Fernandez AF, Gammon ST, Kaye J, et al. Glypican-1 identifies cancer exosomes and detects early pancreatic cancer. Nature. 2015;523(7559):177–82. doi:10.1038/nature14581.
Ferracin M, Lupini L, Salamon I, Saccenti E, Zanzi MV, Rocchi A, et al. Absolute quantification of cell-free microRNAs in cancer patients. Oncotarget. 2015;6(16):14545–55.
Madhavan B, Yue S, Galli U, Rana S, Gross W, Muller M, et al. Combined evaluation of a panel of protein and miRNA serum-exosome biomarkers for pancreatic cancer diagnosis increases sensitivity and specificity. Int J Cancer. 2015;136(11):2616–27. doi:10.1002/ijc.29324.
Aragon Han P, Weng CH, Khawaja HT, Nagarajan N, Schneider EB, Umbricht CB, et al. MicroRNA expression and association with clinicopathologic features in papillary thyroid cancer: a systematic review. Thyroid. 2015. doi:10.1089/thy.2015.0193.
Dettmer M, Perren A, Moch H, Komminoth P, Nikiforov YE, Nikiforova MN. Comprehensive MicroRNA expression profiling identifies novel markers in follicular variant of papillary thyroid carcinoma. Thyroid. 2013;23(11):1383–9. doi:10.1089/thy.2012.0632.
Acknowledgments
This study was supported by Oncosystem Ltd. for (to R.S. and A.M.) and by grants from the Helmsley Trust Fund (to H.G.H).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The study had Ethical Committee of N.N. Petrov Institute of Oncology approval. Informed consent was obtained from all individual participants included in the study.
Conflicts of interest
None
Rights and permissions
About this article
Cite this article
Samsonov, R., Burdakov, V., Shtam, T. et al. Plasma exosomal miR-21 and miR-181a differentiates follicular from papillary thyroid cancer. Tumor Biol. 37, 12011–12021 (2016). https://doi.org/10.1007/s13277-016-5065-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13277-016-5065-3