Abstract
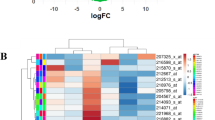
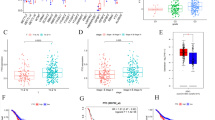
Cervical cancer is the third most common cancer and the fourth leading cause of cancer deaths among women in the world. The discovery of vital diagnostic and therapeutic markers against cervical squamous cell carcinoma (CSCC) would broaden our understanding on the molecular basis of CSCC. In this study, we thoroughly analyzed the transcriptome of CSCC and matched adjacent nontumor (ATN) tissue. RNA sequencing was performed to screen the differentially expressed genes (DEGs) of three pairs of CSCC and ATN tissues. Functional enrichment analysis was used to uncover the biological functions of DEGs. Protein interaction network was carried out to reveal interaction of DEGs. Quantitative real-time PCR was conducted to validate the expression of DEGs. Immunohistochemistry was used to detect the relationship between clinicopathological parameters of CSCC and DEGs. There were a total of 347 significantly common DEGs in the three paired examples, including 104 consistent upregulated and 148 consistent downregulated DEGs. The 347 DEGs were categorized into 73 functional categories by Gene Ontology (GO) analysis. The Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis suggested six significantly signal pathways. The protein interaction network uncovered three important DEGs, including retinol dehydrogenase 12 (RDH12), ubiquitin D (UBD), and serum amyloid A1 (SAA1). We found that RDH12 expression was decreased in 74.5 % of CSCC tissues. RDH12 expression was negatively associated with tumor size and depth of cervical invasion. The UBD was overexpressed in 61.7 % of CSCC tissues and was positively related with tumor size and lymphatic metastasis. The SAA1 protein was overexpressed in 57.4 % of CSCC tissues and was positively related with clinicopathological parameters of tumor size, lymphatic metastasis, and depth of cervical invasion. The RDH12, UBD, and SAA1 genes might participate in the progression of CSCC.




Similar content being viewed by others
References
Jemal A, Bray F, Center M, Ferlay J, Ward E, Forman D. Global cancer statistics. CA Cancer J Clin. 2011;61:69–90.
Mathew A, George PS. Trends in incidence and mortality rates of squamous cell carcinoma and adenocarcinoma of cervix-worldwide. Asian Pac J Cancer Prev. 2009;10:645–50.
McLaughlin-Drubin ME, Meyers J, Munger K. Cancer associated human papil-lomaviruses. Curr Opin Virol. 2012;2:459–66.
Moody CA, Laimins LA. Human papillomavirus oncoproteins: pathways to transformation. Nat Rev Cancer. 2010;10:550–60.
Grabowska AK, Riemer AB. The invisible enemy—how human papillomaviruses avoid recognition and clearance by the host immune system. Open Virol J. 2012;6:249–56.
Plummer M, Schiffman M, Castle PE, Maucort-Boulch D, Wheeler CM, ALTS. Group. A 2-year prospective study of human papillomavirus persistence among women with a cytological diagnosis of atypical squamous cells of undetermined significance or low-grade squamous intraepithelial lesion. J Infect Dis. 2007;195:1582–9.
de Freitas AC, Coimbra EC, Leitão MD. Molecular targets of HPV oncoproteins: potential biomarkers for cervical carcinogenesis. Biochim Biophys Acta. 1845;2014:91–103.
Wang YL, Ding XP, Xiong ZR, Liao H. Differential gene expression profile in cervical cancer and parenchyma infected with human papillomavirus 16 screened by cDNA microarray. Eur J Gynaecol Oncol. 2013;34:132–7.
Gautier L, Cope L, Bolstad BM, Irizarry RA. Affy-analysis of affymetrix genechip data at the probe level. Bioinformatics. 2004;20:307–15.
Marioni JC, Mason CE, Mane SM, Stephens M, Gilad Y. RNA-seq: an assessment of technical reproducibility and comparison with gene expression arrays. Genome Res. 2008;18:1509–17.
Zhang XM, Ma ZW, Wang Q, Wang JN, Yang JW, Li XD, et al. A new RNA-seq method to detect the transcription and non-codingRNA in prostate cancer. Pathol Oncol Res. 2014;20(1):43–50.
Wu Y, Wang X, Wu F, Huang R, Xue F, Liang G, et al. Transcriptome profiling of the cancer, adjacent non-tumor and distant normal tissues from a colorectal cancer patient by deep sequencing. PLoS One. 2012;7:e41001.
Stephens PJ, Tarpey PS, Davies H, Von Loo P, Greenman C, Wedge DC, et al. The landscape of cancer genes and mutational processes in breast cancer. Nature. 2012;486:400–4.
Trapnell C, Williams BA, Pertea G, Mortazavi A, Kwa G, van Baren MJ, et al. Transcript assembly and quantification by RNA-seq reveals unannotated transcripts and inform switching during cell differentiation. Nat Biotechnol. 2010;28:511–5.
Zhang W, Edwards A, Fan W, Fang Z, Deininger P, Zhang K. Inferring the expression variability of human transposable element-derived exons by linear model analysis of deep RNA sequencing data. BMC Genomics. 2013;14:584.
da Huang W, Sherman BT, Lempicki RA. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc. 2009;4:44–57.
Morimoto I, Sasaki Y, Ishida S, Imai K, Tokino T. Identification of the osteopontin gene as a direct target of TP53. Genes Chromosome Cancer. 2002;33:270–8.
Wong YF, Cheung TH, Tsao GS, Lo KW, Yim SF, Wang VW, et al. Genome-wide gene expression profiling of cervical cancer in Hong Kong women by oligonucleotide microarray. Int J Cancer. 2006;118:2461–9.
Peng X, Wu Z, Yu L, Li J, Xu W, Chan HC, et al. Overexpression of cystic fibrosis transmembrane conductance regulator (CFTR) is associated with humancervical cancer malignancy, progression and prognosis. Gynecol Oncol. 2012;125:470–6.
Lee J, Yoon YS, Chung JH. Epigenetic silencing of the WNT antagonist DICKKOPF-1 in cervical cancer cell lines. Gynecol Oncol. 2008;109:270–4.
Yamaki M, Sugiura K, Muro Y, Shimoyama Y, Tomita Y. Epidermal growth factor receptor tyrosine kinase inhibitors induce CCL2 and CCL5 via reduction in IL-1R2 in keratinocytes. Exp Dermatol. 2010;19:730–5.
Sun YX, Fang M, Wang J, Cooper CR, Pienta KJ, Taichman RS. Expression and activation of alpha v beta 3 integrins by SDF-1/CXC12 increases the aggressiveness of prostate cancer cells. Prostate. 2007;67:61–73.
Eke I, Koch U, Hehlgans S, Sandfort V, Stanchi F, Zips D, et al. PINCH1 regulates Akt1 activation and enhances radioresistance by inhibiting PP1alpha. J Clin Invest. 2010;120:2516–27.
Ahmed KM, Zhang H, Park CC. NF-κB Regulates radioresistance mediated by β1-integrin in three-dimensional culture of breast cancer cells. Cancer Res. 2013;73:3737–48.
Ou J, Luan W, Deng J, Sa R, Liang H. αV integrin induces multicellular radioresistance in human nasopharyngeal carcinoma via activating SAPK/JNK pathway. PLoS One. 2012;7:e38737.
Zhou SL, Dai Z, Zhou ZJ, Chen Q, Wang Z, Xiao YS, et al. CXCL5 contributes to tumour metastasis and recurrence of intrahepatic cholangiocarcinoma by recruiting infiltrative intratumoural neutrophils. Carcinogenesis. 2014;35:597–605.
Pitteri SJ, Kelly-Spratt KS, Gurley KE, Kennedy J, Buson TB, Chin A, et al. Tumor microenvironment-derived proteins dominate the plasma proteome response during breast cancer induction and progression. Cancer Res. 2011;71:5090–100.
Kang JU, Koo SH, Kwon KC, Park JW, Kim JM. Identification of novel candidate target genes, including EPHB3, MASP1 and SST at 3q26.2-q29 in squamous cell carcinoma of the lung. BMC Cancer. 2009;9:237.
Liu YS, Luo XY, Li QR, Li H, Li C, Ni H, et al. Shotgun and targeted proteomics reveal that pre-surgery serum levels of LRG1, SAA, and C4BP may refine prognosis of resected squamous cell lung cancer. J Mol Cell Biol. 2012;4:344–7.
Samarut E, Rochette-Egly C. Nuclear retinoic acid receptors: conductors of the retinoic acid symphony during development. Mol Cell Endocrinol. 2012;3488:348–60.
Skrypnyk N, Chen X, Hu W, Su Y, Mont S, Yang S, et al. PPARα activation can help prevent and treat non-small cell lung cancer. Cancer Res. 2014;74:621–31.
Su TR, Lin JJ, Tsai CC, Huang TK, Yang ZY, Wu MO, et al. Inhibition of melanogenesis by gallic acid: possible involvement of the PI3K/Akt, MEK/ERK and Wnt/β-catenin signaling pathways in B16F10 cells. Int J Mol Sci. 2013;14:20443–58.
Duester G. Families of retinoid dehydrogenases regulating vitamin A function: production of visual pigment and retinoic acid. Eur J Biochem. 2000;267:4315–24.
de Thé H, Chomienne C, Lanotte M, Degos L, Dejean A. The t(15;17) translocation of acute promyelocytic leukaemia fuses the retinoic acid receptor alpha gene to a novel transcribed locus. Nature. 1990;347:558–61.
Kropotova ES, Zinov’eva OL, Zyrianova AF, Choĭnzonov EL, Afanas’ev SG, Cherdyntseva NV, et al. Expression of genes involved in retinoic acid biosynthesis in human gastric cancer. Mol Biol. 2013;47:317–30.
Chen J, Yang L, Chen H, Yuan T, Liu M, Chen P. Recombinat adenovirus encoding FAT10 small interfering RNA inhibits HCC growth in vitro and in vivo. Exp Mol Pathol. 2014;96:207–11.
Gao Y, Theng SS, Zhuo J, Teo WB, Ren J, Lee CG. FAT10, an ubiquitin-like protein, confers malignant properties in non-tumorigenic and tumorigenic cells. Carcinogenesis. 2014;35:923–34.
Olsen HG, Skovgaard K, Nielsen OL, Leifsson PS, Jensen HE, Lburg T, et al. Organization and biology of the porcine serum amyloid A (SAA) gene cluster: isoform specific responses to bacterial infection. PLoS One. 2013;8:e76695.
Uhlar CM, Whitehead AS. Serum amyloid A, the major vertebrate acute-phase reactant. Eur J Biochem. 1999;9:501–23.
Lu SY, Rodriguez M, Liao WS. YY1 represses rat serum amyloid A1 gene transcription and is antagonized by NF-kappa B during acute-phase response. Mol Cell Biol. 1994;9:6523–63.
Furlaneto CJ, Campa A. A novel function of serum amyloid a: a potent stimulus for the release of tumor necrosis factor-alpha, interleukin-1beta, and interleukin-8 by human blood neutrophil. Biochem Biophys Res Commun. 2000;9:405–8.
Connolly M, Marrelli A, Blades M, McCormick J, Maderna P, Godson C, et al. Acute serum amyloid a induces migration, angiogenesis, and inflammation in synovial cells in vitro and in a human rheumatoid arthritis/SCID mouse chimera model. J Immunol. 2010;9:6427–37.
Lee HY, Kim MK, Park KS, Bae YH, Yun J, Park JI, et al. Serum amyloid A stimulates matrix-metalloproteinase-9 upregulation via formyl peptide receptor like-1-mediated signaling in human monocytic cells. Biochem Biophys Res Commun. 2005;9:989–98.
Acknowledgments
This research is supported by the National Natural Science Foundation of China (81472446).
Conflicts of interest
None
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
S1
The list of differentially expressed genes. (XLS 227 kb)
S2
The list of functional categories of differentially expressed genes. (XLS 83 kb)
S3
Protein-protein interaction network construction of CSCC. The 13 cervical cancer related genes from NCI are filled with blue. Three novel cervical cancer related genes identified in our study are filled with red. The other detected DEGs in the PPI network are filled with green. Each link between two nodes corresponds to a direct interaction between the two genes according to HPRD database (GIF 2948 kb)
Rights and permissions
About this article
Cite this article
Peng, G., Dan, W., Jun, W. et al. Transcriptome profiling of the cancer and adjacent nontumor tissues from cervical squamous cell carcinoma patients by RNA sequencing. Tumor Biol. 36, 3309–3317 (2015). https://doi.org/10.1007/s13277-014-2963-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13277-014-2963-0