Abstract
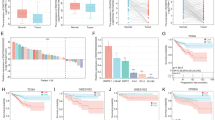
Prostate cancer is a big killer in many regions especially American men, and this year, the diagnosed rate rises rapidly. We aimed to find the biomarker or any changing in prostate cancer patients. With the development of next generation sequencing, much genomic alteration has been found. Here, basing on the RNA-seq result of human prostate cancer tissue, we tried to find the transcription or non-coding RNA expressed differentially between normal tissue and prostate cancer tissue. 10 T sample data is the RNA-seq data for prostate cancer tissue in this study, we found the differential gene is TFF3-Trefoil factor 3, which was more than seven fold change from prostate cancer tissue to normal tissue, and the most outstanding transcript is C15orf21. Additionally, 9 lncRNAs were found according our method. Finally, we found the many important non-coding RNA related to prostate cancer, some of them were long non-coding RNA (lncRNA).




Similar content being viewed by others
References
Berger MF, Lawrence MS, Demichelis F, Drier Y, Cibulskis K, Sivachenko AY, Sboner A, Esgueva R, Pflueger D, Sougnez C (2011) The genomic complexity of primary human prostate cancer. Nature 470(7333):214–220
Barbieri CE, Baca SC, Lawrence MS, Demichelis F, Blattner M, Theurillat JP, White TA, Stojanov P, Van Allen E, Stransky N (2012) Exome sequencing identifies recurrent SPOP, FOXA1 and MED12 mutations in prostate cancer. Nat Genet 44(6):685–689
Grasso CS, Wu YM, Robinson DR, Cao X, Dhanasekaran SM, Khan AP, Quist MJ, Jing X, Lonigro RJ, Brenner JC (2012) The mutational landscape of lethal castration-resistant prostate cancer. Nature 487(7406):239–243
Morin RD, Bainbridge M, Fejes A, Hirst M, Krzywinski M, Pugh TJ, McDonald H, Varhol R, Jones SJM, Marra MA (2008) Profiling the HeLa S3 transcriptome using randomly primed cDNA and massively parallel short-read sequencing. Biotechniques 45(1):81–94
Maher CA, Kumar-Sinha C, Cao X, Kalyana-Sundaram S, Han B, Jing X, Sam L, Barrette T, Palanisamy N, Chinnaiyan AM (2009) Transcriptome sequencing to detect gene fusions in cancer. Nature 458(7234):97–101
Pflueger D, Terry S, Sboner A, Habegger L, Esgueva R, Lin PC, Svensson MA, Kitabayashi N, Moss BJ, MacDonald TY (2011) Discovery of non-ETS gene fusions in human prostate cancer using next-generation RNA sequencing. Genome Res 21(1):56–67
Sahu A, Iyer MK, Chinnaiyan AM (2012) Insights into Chinese prostate cancer with RNA-seq. Cell Res 22(5):786–788
Mattick JS (2001) Non-coding RNAs: the architects of eukaryotic complexity. EMBO reports 2(11):986–991
Stefani G, Slack FJ (2008) Small non-coding RNAs in animal development. Nat Rev Mol Cell Biol 9(3):219–230
Mercer TR, Dinger ME, Mattick JS (2009) Long non-coding RNAs: insights into functions. Nat Rev Genet 10(3):155–159
Gupta RA, Shah N, Wang KC, Kim J, Horlings HM, Wong DJ, Tsai MC, Hung T, Argani P, Rinn JL (2010) Long non-coding RNA HOTAIR reprograms chromatin state to promote cancer metastasis. Nature 464(7291):1071–1076
Ren S, Peng Z, Mao JH, Yu Y, Yin C, Gao X, Cui Z, Zhang J, Yi K, Xu W (2012) RNA-seq analysis of prostate cancer in the Chinese population identifies recurrent gene fusions, cancer-associated long noncoding RNAs and aberrant alternative splicings. Cell Res 22(5):806–821
Leinonen R, Akhtar R, Birney E, Bower L, Cerdeno-Tárraga A, Cheng Y, Cleland I, Faruque N, Goodgame N, Gibson R (2011) The European nucleotide archive. Nucleic Acids Res 39(suppl 1):D28–D31
Trapnell C, Pachter L, Salzberg SL (2009) TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25(9):1105–1111
Trapnell C, Roberts A, Goff L, Pertea G, Kim D, Kelley DR, Pimentel H, Salzberg SL, Rinn JL, Pachter L (2012) Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat Protoc 7(3):562–578
Jiang H, Wong WH (2009) Statistical inferences for isoform expression in RNA-Seq. Bioinformatics 25(8):1026–1032
Toung JM, Morley M, Li M, Cheung VG (2011) RNA-sequence analysis of human B-cells. Genome Res 21(6):991–998
Garraway IP, Seligson D, Said J, Horvath S, Reiter RE (2004) Trefoil factor 3 is overexpressed in human prostate cancer. Prostate 61(3):209–214
Laxman B, Morris DS, Yu J, Siddiqui J, Cao J, Mehra R, Lonigro RJ, Tsodikov A, Wei JT, Tomlins SA (2008) A first-generation multiplex biomarker analysis of urine for the early detection of prostate cancer. Cancer Res 68(3):645
Vestergaard EM, Borre M, Poulsen SS, Nexø E, Tørring N (2006) Plasma levels of trefoil factors are increased in patients with advanced prostate cancer. Clin Cancer Res 12(3):807–812
Ather MH, Abbas F, Faruqui N, Israr M, Pervez S (2004) Expression of pS2 in prostate cancer correlates with grade and Chromogranin A expression but not with stage. BMC Urol 4(1):14
Tomlins SA, Laxman B, Dhanasekaran SM, Helgeson BE, Cao X, Morris DS, Menon A, Jing X, Cao Q, Han B (2007) Distinct classes of chromosomal rearrangements create oncogenic ETS gene fusions in prostate cancer. Nature 448(7153):595–599
Cabili MN, Trapnell C, Goff L, Koziol M, Tazon-Vega B, Regev A, Rinn JL (2011) Integrative annotation of human large intergenic noncoding RNAs reveals global properties and specific subclasses. Genes Dev 25(18):1915–1927
Conflict of Interest
The authors have no conflict of interest to declare.
Author information
Authors and Affiliations
Corresponding author
Additional information
Xiao-Ming Zhang, Zhong-Wei Ma and Qiang Wang contributed equally to this work as the co-first author.
Rights and permissions
About this article
Cite this article
Zhang, XM., Ma, ZW., Wang, Q. et al. A New RNA-Seq Method to Detect the Transcription and Non-coding RNA in Prostate Cancer. Pathol. Oncol. Res. 20, 43–50 (2014). https://doi.org/10.1007/s12253-013-9618-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12253-013-9618-0