Abstract
The rosa monophyletic group of transposons is a group of transposable element with characteristics of encoding a DD41D motif in the catalytic domain. However, biology and evolutionary history of this monophyletic group are still poorly understood. In this study, we report the first description for the presence of a rosa transposon in the silkworm Bombyx mori. Further analyses confirmed that this element in the silkworm genome had recently amplified and might still be capable of transposition. In addition, we present evidence, based on searches of publicly available insect genomes, that a new clade of the rosa monophyletic group was identified. Interestingly, analysis of their three dimensional structures suggested that these proteins showed highly similar protein structures with that of the Mos1 transposase. These results provided useful insights into the functionality of these transposases and their structural and functional deviations from other transposases in the Tc1/mariner superfamily. Meanwhile, sequence and phylogenetic analysis confirmed that DD41D and maT elements might represent another independent large group of the Tc1/mariner superfamily. Importantly, the result of the comparison of terminal inverted repeats (TIRs) validated that DD41D and maT elements almost had identical consensus terminal sequences (5′-CAGGGTGNSNCA-3′), implying they might have similar cleavage sites or patterns during the process of their transposition. In a word, this study will enrich and expand our knowledge of the Tc1/mariner superfamily.





Similar content being viewed by others
References
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Aravind L, Landsman D (1998) AT-hook motifs identified in a wide variety of DNA-binding proteins. Nucleic Acids Res 26:4413–4421
Auge′-Gouillou C, Hamelin MH, Demattei MV, Periquet G, Bigot Y (2001) The ITR binding domain of the mariner Mos1 transposase. Mol Genet Genom 265:58–65
Brillet B, Bigot Y, Augé-Gouillou C (2007) Assembly of the Tc1 and mariner transposition initiationcomplexes depends on the origins of their transposase DNAbinding domains. Genetica 130:105–120
Bryan G, Garza D, Hartl D (1990) Insertion and excision of the transposable element mariner in Drosophila. Genetics 125:103–114
Capy P, Vitalis R, Langin T, Higuet D, Bazin C (1996) Relationships between transposable elements based upon the integrase-transposase domains: is there a common ancestor? J Mol Evol 42:359–368
Clark KJ, Carlson DF, Leaver MJ, Foster LK, Fahrenkrug SC (2009) Passport, a native Tc1 transposon from flatfish, is functionally active in vertebrate cells. Nucleic Acids Res 37:1239–1247
Claudianos C, Brownlie J, Russell R, Oakeshott J, Whyard S (2002) maT:a clade of transposons intermediate between mariner and Tc1. Mol Biol Evol 19:2101–2109
Collins J, Forbes E, Anderson P (1989) The Tc3 family of transposable genetic elements in Caenorhabditis elegans. Genetics 121:47–55
Cui Z, Geurts AM, Liu G, Kaufman CD, Hackett PB (2002) Structure–function analysis of the inverted terminal repeats of the Sleeping Beauty transposon. J Mol Biol 318:1221–1235
Daboussi MJ, Langin T, Brygoo Y (1992) Fot1, a new family of fungal transposable elements. Mol Gen Genet 232:12–16
Delauriere L, Chenais B, Hardivillier Y, Gauvry L, Casse N (2009) Mariner transposons as genetic tools in vertebrate cells. Genetica 137:9–17
Duan J, Li R, Cheng D, Fan W, Zha X, Cheng T, Wu Y, Wang J, Mita K, Xiang Z, Xia Q (2010) SilkDB v20: a platform for silkworm (Bombyx mori) genome biology. Nucleic Acids Res 38:D453–D456
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32:1792–1797
Emmons SW, Yesner L, Ruan K, Katzenberg D (1983) Evidence for a transposon in Caenorhabditis elegans. Cell 32:55–65
Feschotte C, Pritham EJ (2007) DNA transposons and the evolution of eukaryotic genomes. Annu Rev Genet 41:331–368
Franz G, Savakis C (1991) Minos, a new transposable element from Drosophila hydei, is a member of the Tc1-like family of transposons. Nucleic Acids Res 19:6646
Gomulski LM, Torti C, Bonizzoni M, Moralli D, Raimondi E, Capy P, Gasperi G, d Malacrida AR (2001) A new basal subfamily of mariner elements in Ceratitis rosa and other tephritid flies. J Mol Evol 53:597–606
Haymer DS, Marsh JL (1986) Germ line and somatic instability of a white mutation in Drosophila mauritiana due to a transposable genetic element. Dev Genetics 6:281–291
Izsvak Z, Khare D, Behlke J, Heinemann U, Plasterk RH, Ivics Z (2002) Involvement of a bifunctional, paired-likeDNA-binding domain and a transpositional enhancer inSleeping Beauty transposition. J Biol Chem 277:34581–34588
Jacobson JW, Medhora MM, Hartl DL (1986) Molecular structure of a somatically unstable transposable element in Drosophila. Proc Natl Acad Sci USA 83:8684–8688
Jee SH, Kim GE, Hong SH, Seo SB, Shim JK, Park SC, Choo JK (2007) Characterization of EamaT1, a member of maT family of transposable elements from the earthworm Eisenia andrei (Annelida, Oligochaeta). Mol Genet Genomics 278:479–486
Jurka J, Kapitonov VV, Pavlicek A, Klonowski P, Kohany O, Walichiewicz J (2005) Repbase Update, a database of eukaryotic repetitive elements. Cytogenet Genome Res 110:462–467
Kalendar R, Lee D, Schulman AH (2014) FastPCR software for PCR, in silico PCR, and oligonucleotide assembly and analysis. Methods Mol Biol 1116:271–302
Kelley LA, Mezulis S, Yates CM, Wass MN, Sternberg MJ (2015) The Phyre2 web portal for protein modeling, prediction and analysis. Nat Protoc 10:845–858
Kumaresan and Mathavan (2004) Molecular diversity and phylogenetic analysis of mariner-like transposons in the genome of the silkworm Bombyx mori. Insect Mol Biol 2004(13):259–271
Lampe DJ (2010) Bacterial genetic methods to explore the biology of mariner transposons. Genetica 138:499–508
Langin T, Capy P, Daboussi MJ (1995) The transposable element impala, a fungal member of the Tc1-mariner superfamily. Mol Gen Genet 246:19–28
Liu Y, Yang G (2014) Tc1-like transposable elements in plant genomes. Mob DNA 5:17
Lohe A, Sullivan D, Hartl D (1996) Genetic evidence for subunit interactions in the transposase of the transposable element mariner. Genetics 144:1087–1095
McGuffin LJ, Bryson K, Jones DT (2000) The PSIPRED protein structure prediction server. Bioinformatics 16(4):404–405
Munoz-Lopez M, Siddique A, Bischerour J, Lorite P, Chalmers R, Palomeque T (2008) Transposition of Mboumar-9: identification of a new naturally active mariner-family transposon. J Mol Evol 382:567–572
Nicholas KB, Nicholas HB, d Deerfield DW (1997) GeneDoc: analysis and visualization of genetic variation. EMBNEW News 4:14
Pietrokovski S, Henikoff S (1997) A helix-turn-helix DNA-binding motif predicted for transposases of DNA transposons. Mol Gen Genet 254:689–695
Rice P, Longden I, Bleasby A (2000) EMBOSS: the European molecular biology open software suite. Trends Genet 16:276–277
Richardson JM, Colloms SD, Finnegan DJ, Walkinshaw MD (2009) Molecular architecture of the Mos1 paired-end complex: the structural basis of DNA transposition in a eukaryote. Cell 138:1096–1108
Robertson HM, Lampe DJ (1995) Recent horizontal transfer of a mariner transposable element among and between Diptera and Neuroptera. Mol Biol Evol 12:850–862
Shao H, Tu Z (2001) Expanding the diversity of the IS630-Tc1-mariner superfamily: discovery of a unique DD37E transposon and reclassification of the DD37D and DD39D transposons. Genetics 159:1103–1115
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
van Pouderoyen G, Ketting RF, Perrakis A, Plasterk RH (1997) Sixma TK (1997), Crystal structure of the specific DNA-binding domain of Tc3 transposase of C. elegans in complex with transposon DNA. EMBO J 160:6044–6054
Xia X, Xie Z (2001) DAMBE: software package for data analysis in molecular biology and evolution. J Hered 92:371–373
Acknowledgments
This work is supported by the National Natural Science Foundation of China (31560308 to ZHH, 31260632 to ZXG and 31401106 to HMJ), the National Natural Science Foundation in Jiangxi Province of China (20122BAB204018 to ZXG) and Doctoral start-up funding of Jiujiang University (8879424 to ZHH).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare no conflict of interest.
Additional information
Hua-Hao Zhang and Yi-Hong Shen contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Fig. S1
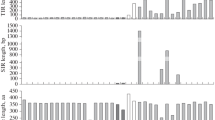
The frequency distributions (ordinate) of sequence identities (abscissa) of rosa_Bm. The percentage identity is based on BLAST search, using the consensus as the query. The sequences shorter than 100 bp were excluded. The black line represents the tendency estimated by a simple moving average of the percentage identity scores. Supplementary material 1 (PDF 106 kb)
Rights and permissions
About this article
Cite this article
Zhang, HH., Shen, YH., Xiong, XM. et al. Identification and evolutionary history of the DD41D transposons in insects. Genes Genom 38, 109–117 (2016). https://doi.org/10.1007/s13258-015-0356-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13258-015-0356-4