Abstract
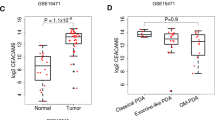
Even though cell–cell adhesion molecule carcinoembryonic antigen-related cell adhesion molecule 1 (CEACAM1) is extensively studied since the discovery, the role of CEACAM1 in different cancers is not completely clarified. In the present study, we examined CEACAM1 expression and its association with patient survival in various cancers by analysis of multiple databases. Oncomine database analysis revealed that CEACAM1 expression was upregulated in lung and pancreatic cancers, but downregulated in colorectal and head and neck cancers. PrognoScan and Kaplan‑Meier analyses showed that colorectal cancer patients as well as head and neck cancer patients with high CEACAM1 expression exhibited a higher overall survival rate. STRING analysis identified CEACAM3, CEACAM8, FN1, etc. as CEACAM1 interactors. Gene alteration analysis showed that CEACAM1 mutation predominantly occurred in the N-terminal. Coexpression analysis demonstrated that CEACAM1 had distinct coexpressed genes in different cancers, but KRT protein was consistently coexpressed with CEACAM1 in diverse cancer types. All the observations supported that CEACAM1 can serve as a diagnostic marker for some cancers, such as pancreatic cancer. And high CEACAM1 expression provides a better prognosis for some cancers, such as colorectal and head and neck cancers.















Similar content being viewed by others
References
Beauchemin N, Arabzadeh A (2013) Carcinoembryonic antigen-related cell adhesion molecules (CEACAMs) in cancer progression and metastasis. Cancer Metast Rev 32:643–671. https://doi.org/10.1007/s10555-013-9444-6
Beauchemin N, Kunath T, Robitaille J et al (1997) Association of biliary glycoprotein with protein tyrosine phosphatase SHP-1 in malignant colon epithelial cells. Oncogene 14:783–790. https://doi.org/10.1038/sj.onc.1200888
Beauchemin N, Draber P, Dveksler G et al (1999) Redefined nomenclature for members of the carcinoembryonic antigen family. Exp Cell Res 252:243–249. https://doi.org/10.1006/excr.1999.4610
Bhandari V, Hoey C, Liu LY et al (2019) Molecular landmarks of tumor hypoxia across cancer types. Nat Genet 51:308–318. https://doi.org/10.1038/s41588-018-0318-2
Blake SJ, Stannard K, Liu J et al (2016) Suppression of metastases using a new lymphocyte checkpoint target for cancer immunotherapy. Cancer Discov 6:446–459. https://doi.org/10.1158/2159-8290.Cd-15-0944
Bray F, Ferlay J, Soerjomataram I et al (2018) Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 68:394–424. https://doi.org/10.3322/caac.21492
Calinescu A, Turcu G, Nedelcu RI et al (2018) On the dual role of carcinoembryonic antigen-related cell adhesion molecule 1 (CEACAM1) in human malignancies. J Immunol Res 2018:1–8. https://doi.org/10.1155/2018/7169081
Campbell JD, Alexandrov A, Kim J et al (2016) Distinct patterns of somatic genome alterations in lung adenocarcinomas and squamous cell carcinomas. Nat Genet 48:607–616. https://doi.org/10.1038/ng.3564
Cerami E, Gao J, Dogrusoz U et al (2012) The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov 2:960–960. https://doi.org/10.1158/2159-8290.Cd-12-0326
Dankner M, Gray-Owen SD, Huang YH et al (2017) CEACAM1 as a multi-purpose target for cancer immunotherapy. Oncoimmunology 6:e1328336. https://doi.org/10.1080/2162402x.2017.1328336
Ellrott K, Bailey MH, Saksena G et al (2018) Scalable open science approach for mutation calling of tumor exomes using multiple genomic pipelines. Cell Syst 6:271–281. https://doi.org/10.1016/j.cels.2018.03.002
Gao JJ, Aksoy BA, Dogrusoz U et al (2013) Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal 6:pl1. https://doi.org/10.1126/scisignal.2004088
Gao QS, Liang WW, Foltz SM et al (2018) Driver fusions and their implications in the development and treatment of human cancers. Cell Rep 23:227–238. https://doi.org/10.1016/j.celrep.2018.03.050
Gardner EE, Lok BH, Schneeberger VE et al (2017) Chemosensitive relapse in small cell lung cancer proceeds through an EZH2-SLFN11 axis. Cancer Cell 31:286–299. https://doi.org/10.1016/j.ccell.2017.01.006
Gerstel D, Wegwitz F, Jannasch K et al (2011) CEACAM1 creates a pro-angiogenic tumor microenvironment that supports tumor vessel maturation. Oncogene 30:4275–4288. https://doi.org/10.1038/onc.2011.146
Giannakis M, Mu XJ, Shukla SA et al (2016) Genomic correlates of immune-cell infiltrates in colorectal carcinoma. Cell Rep 17:1206–1206. https://doi.org/10.1016/j.celrep.2016.10.009
Giulietti M, Occhipinti G, Principato G et al (2016) Weighted gene co-expression network analysis reveals key genes involved in pancreatic ductal adenocarcinoma development. Cell Oncol 39:379–388. https://doi.org/10.1007/s13402-016-0283-7
Gyorffy B, Lanczky A, Eklund AC et al (2010) An online survival analysis tool to rapidly assess the effect of 22,277 genes on breast cancer prognosis using microarray data of 1,809 patients. Breast Cancer Res Tr 123:725–731. https://doi.org/10.1007/s10549-009-0674-9
Helfrich I, Singer BB (2019) Size matters: the functional role of the ceacam1 isoform signature and its impact for nk cell-mediated killing in melanoma. Cancers 11:356. https://doi.org/10.3390/cancers11030356
Ho AS, Kannan K, Roy DM et al (2013) The mutational landscape of adenoid cystic carcinoma. Nat Genet 45:791. https://doi.org/10.1038/ng.2643
Hoadley KA, Yau C, Hinoue T et al (2018) Cell-of-origin patterns dominate the molecular classification of 10,000 tumors from 33 types of cancer. Cell 173:291–304. https://doi.org/10.1016/j.cell.2018.03.022
Horst AK, Wagener C (2004) CEA-Related CAMs. Handb Exp Pharmacol. https://doi.org/10.1007/978-3-540-68170-0_10
Huang YH, Zhu C, Kondo Y et al (2015) CEACAM1 regulates TIM-3-mediated tolerance and exhaustion. Nature 517:386–U566. https://doi.org/10.1038/nature13848
Huber M, Izzi L, Grondin P et al (1999) The carboxyl-terminal region of biliary glycoprotein controls its tyrosine phosphorylation and association with protein-tyrosine phosphatases SHP-1 and SHP-2 in epithelial cells. J Biol Chem 274:335–344. https://doi.org/10.1074/jbc.274.1.335
Ieda J, Yokoyama S, Tamura K et al (2011) Re-expression of CEACAM1 long cytoplasmic domain isoform is associated with invasion and migration of colorectal cancer. Int J Cancer 129:1351–1361. https://doi.org/10.1002/ijc.26072
Imielinski M, Berger AH, Hammerman PS et al (2012) Mapping the hallmarks of lung adenocarcinoma with massively parallel sequencing. Cell 150:1107–1120. https://doi.org/10.1016/j.cell.2012.08.029
Jamal-Hanjani M, Wilson GA, McGranahan N et al (2017) Tracking the evolution of non-small-cell lung cancer. New Engl J Med 376:2109–2121. https://doi.org/10.1056/NEJMoa1616288
Kammerer R, Zimmermann W (2010) Coevolution of activating and inhibitory receptors within mammalian carcinoembryonic antigen families. BMC Biol 8:12. https://doi.org/10.1186/1741-7007-8-12
Lefebvre C, Bachelot T, Filleron T et al (2016) Mutational profile of metastatic breast cancers: a retrospective analysis. PLoS Med 13:e1002201. https://doi.org/10.1371/journal.pmed.1002201
Leung N, Turbide C, Olson M et al (2006) Deletion of the carcinoembryonic antigen-related cell adhesion molecule 1 (Ceacam1) gene contributes to colon tumor progression in a murine model of carcinogenesis. Oncogene 25:5527–5536. https://doi.org/10.1038/sj.onc.1209541
Liu JF, Lichtenberg T, Hoadley KA et al (2018) An integrated TCGA pan-cancer clinical data resource to drive high-quality survival outcome analytics. Cell 173:400–416. https://doi.org/10.1016/j.cell.2018.02.052
McLeod RL, Angagaw MH, Baral TN et al (2018) Characterization of murine CEACAM1 in vivo reveals low expression on CD8(+) T cells and no tumor growth modulating activity by anti-CEACAM1 mAb CC1. Oncotarget 9:34459–34470. https://doi.org/10.18632/oncotarget.26108
Mizuno H, Kitada K, Nakai K et al (2009) PrognoScan: a new database for meta-analysis of the prognostic value of genes. Bmc Med Genomics 2:18. https://doi.org/10.1186/1755-8794-2-18
Morin RD, Mendez-Lago M, Mungall AJ et al (2011) Frequent mutation of histone-modifying genes in non-Hodgkin lymphoma. Nature 476:298–303. https://doi.org/10.1038/nature10351
Nagy A, Lanczky A, Menyhart O et al (2018) Validation of miRNA prognostic power in hepatocellular carcinoma using expression data of independent datasets. Sci Rep 8:9227. https://doi.org/10.1038/s41598-018-27521-y
Nollau P, Scheller H, KonaHorstmann M et al (1997) Expression of CD66a (human C-CAM) and other members of the carcinoembryonic antigen gene family of adhesion molecules in human colorectal adenomas. Cancer Res 57:2354–2357
Pereira B, Chin SF, Rueda OM et al (2016) The somatic mutation profiles of 2,433 breast cancers refine their genomic and transcriptomic landscapes. Nat Commun 7:11479. https://doi.org/10.1038/ncomms11479
Rhodes DR, Yu J, Shanker K et al (2004) ONCOMINE: a cancer microarray database and integrated data-mining platform. Neoplasia 6:1–6. https://doi.org/10.1016/S1476-5586(04)80047-2
Rhodes DR, Kalyana-Sundaram S, Mahavisno V et al (2007) Oncomine 3.0: genes, pathways, and networks in a collection of 18,000 cancer gene expression profiles. Neoplasia 9:166–180. https://doi.org/10.1593/neo.07112
Sanchez-Vega F, Mina M, Armenia J et al (2018) The molecular landscape of oncogenic signaling pathways in The Cancer Genome Atlas. Cancer Res 78:3302. https://doi.org/10.1158/1538-7445.Am2018-3302
Seshagiri S, Stawiski EW, Durinck S et al (2012) Recurrent R-spondin fusions in colon cancer. Nature 488:660. https://doi.org/10.1038/nature11282
Shah SP, Roth A, Goya R et al (2012) The clonal and mutational evolution spectrum of primary triple-negative breast cancers. Nature 486:395–399. https://doi.org/10.1038/nature10933
Shi JF, Xu SX, He P et al (2014) Expression of carcinoembryonic antigen-related cell adhesion molecule 1(CEACAM1) and its correlation with angiogenesis in gastric cancer. Pathol Res Pract 210:473–476. https://doi.org/10.1016/j.prp.2014.03.014
Simeone DM, Ji B, Banerjee M et al (2007) CEACAM1, a novel serum biomarker for pancreatic cancer. Pancreas 34:436–443. https://doi.org/10.1097/MPA.0b013e3180333ae3
Szklarczyk D, Gable AL, Lyon D et al (2019) STRING v11: protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res 47:D607–D613. https://doi.org/10.1093/nar/gky1131
Taylor AM, Shih J, Ha G et al (2018) Genomic and functional approaches to understanding cancer aneuploidy. Cancer Cell 33:676–689. https://doi.org/10.1016/j.ccell.2018.03.007
Wang Y, Chen Y, Yan Y et al (2016) Loss of CEACAM1, a tumor-associated factor, attenuates post-infarction cardiac remodeling by inhibiting apoptosis. Sci Rep 6:21972. https://doi.org/10.1038/srep21972
Weng CY, Nguyen T, Shively JE (2016) miRNA-342 regulates CEACAM1-induced lumen formation in a three-dimensional model of mammary gland morphogenesis. J Biol Chem 291:16777–16786. https://doi.org/10.1074/jbc.M115.710152
Witkiewicz AK, McMillan EA, Balaji U et al (2015) Whole-exome sequencing of pancreatic cancer defines genetic diversity and therapeutic targets. Nat Commun 6:6744. https://doi.org/10.1038/ncomms7744
Zhang L, Zhou W, Velculescu VE et al (1997) Gene expression profiles in normal and cancer cells. Science 276:1268–1272. https://doi.org/10.1126/science.276.5316.1268
Zhu YC, Song DL, Song YL et al (2019) Interferon gamma induces inflammatory responses through the interaction of CEACAM1 and PI3K in airway epithelial cells. J Transl Med 17:147. https://doi.org/10.1186/s12967-019-1894-3
Acknowledgements
We acknowledge the data contributors of the TCGA, Oncomine, Kaplan–Meier plotter, Prognoscan, STRING and cBioPortal databases. The enormous data in these databases are the foundation of deep exploration, multidimensional analysis and delicate visualization across multiple cancers. Additionally, we acknowledge Natural Science Foundation of Zhejiang Province (No. LQ18C010005) for supporting this work.
Author information
Authors and Affiliations
Contributions
CW and YW designed the study, statistically analyzed the data and drafted the manuscript, CW and XH collected the data, YW revised the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest in the publication.
Rights and permissions
About this article
Cite this article
Weng, CY., Hu, XY. & Wang, YJ. Integrated analysis of gene expression, alteration and clinical significance of carcinoembryonic antigen-related cell adhesion molecule 1 in cancer. 3 Biotech 10, 132 (2020). https://doi.org/10.1007/s13205-020-2122-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13205-020-2122-9