Abstract
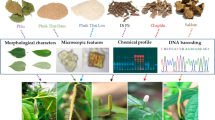
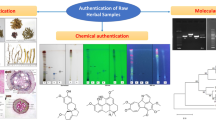
Saraca asoca (Roxb.) De Wilde is an important medicinal plant from the Western Ghats of India, traditionally used in treatment of various gynecological disorders. Increasing commercial demand and decreasing numbers has resulted in this plant becoming endangered with crude drug materials being extensively substituted/adulterated with other plant species. The present study was undertaken with the objective of development and evaluation of multivariate cluster analysis of ISSR fingerprints against rbcL-based DNA barcodes as tool to understand the relationships and to differentiate common adulterants and substituents from S. asoca. ISSR-based Hierarchical Cluster Analysis was carried out on 41 samples of S. asoca and 5 each of the 5 common substituent/adulterant plants and the clustering patterns were evaluated against DNA-sequence-based barcoding of rbcL region of their plastids. Factorial analysis and Principal Coordinate Analysis revealed distinct groups of genetic pools of respective taxa thereby confirming the utility of ISSR fingerprinting as a useful tool for differentiation between the genuine and the adulterants/substituents. NCBI-BLAST search on DNA barcode rbcL region confirmed the results of ISSR assays. Therefore, our study demonstrated the utility of simple, cost-effective method of ISSR fingerprinting coupled with rbcL barcoding in differentiating this important medicinal plant from its common adulterants/substituents.
Graphical Abstract






Similar content being viewed by others
Abbreviations
- ISSR:
-
Inter Simple Sequence Repeat
- rbcL :
-
Ribulose-1, 5-bisphosphate carboxylase/oxygenase large subunit
- PCoA:
-
Principal Coordinate Analysis
- HCA:
-
Hierarchical Cluster Analysis
- CP:
-
Cophenetic Correlation Coefficient
- UPGMA:
-
Unweighted Pair Group Method with Arithmetic Mean
References
Anonymous (2005) Saraca asoca. In: Gupta AK, Tandon N, Sharma M (eds) Quality standards of indian medicinal plants, 2nd edn. Indian Council of Medical Research (ICMR), New Delhi, pp 201–208
Bodin SS, Kim JS, Kim JH (2016) Phylogenetic inferences and the evolution of plastid DNA in Campynemataceae and the Mycoheterotrophic Corsia dispar DL Jones and B Gray (Corsiaceae). Plant Mol Biol Rep 34(1):192–210
Briskin DP (2000) Medicinal plants and phytomedicines: linking plant biochemistry and physiology to human health. Plant Physiol 124(2):507–514
Carvalho FA, Renner SS (2012) A dated phylogeny of the papaya family (Caricaceae) reveals the crop’s closest relatives and the family’s biogeographic history. Mol Phylogenet Evol 65(1):46–53
Chow YL, Quon HH (1968) Chemical constituents of the heartwood of Mesua ferrea. Phytochemistry 7(10):1871–1874
Ellstrand NC (2014) Is gene flow the most important evolutionary force in plants? Am J Bot 101(5):737–753
Farag MA, Sakna ST, El-fiky NM, Shabana MM, Wessjohann LA (2015) Phytochemical, antioxidant and antidiabetic evaluation of eight Bauhinia L species from Egypt using UHPLC–PDA–qTOF-MS and chemometrics. Phytochemistry 119:41–50
Feng SG, Lu JJ, Gao L, Liu JJ, Wang HZ (2014) Molecular phylogeny analysis and species identification of Dendrobium (Orchidaceae) in China. Biochem Genet 52(3–4):127–136
Fu YX, Li WH (1993) Statistical tests of neutrality of mutations. Genetics 133(3):693–709
Garcia-Vallve S, Palau J, Romeu A (1999) Horizontal gene transfer in glycosyl hydrolases inferred from codon usage in Escherichia coli and Bacillus subtilis. Mol Biol Evol 16(9):1125–1134
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucl Acids Symp Ser 41:95–98
Hegde S, Hegde HV, Jalalpure SS, Peram MR, Pai SR, Roy S (2017a) Resolving identification issues of Saraca asoca from its adulterant and commercial samples using phytochemical markers. Pharmacognosy Mag 13:S266–S272
Hegde S, Pai SR, Roy S (2017b) Combination of DNA isolation and RP-HPLC analysis method for bark samples of Saraca asoca and its adulterant. 3 Biotech 7(3):208
Jensen TL, Stubbendeck RM, Barnes AL, Wichers KM, Keller BJ, Bailey CA (2012) Use of phylogenetics for the purpose of quality control in the green algae genus Chlorella. FASEB J 26:9833
Jordan GE, Piel WH (2008) PhyloWidget: web-based visualizations for the tree of life. Bioinformatics 24(14):1641–1642
Jorde PE, Ryman N (1995) Temporal allele frequency change and estimation of effective size in populations with overlapping generations. Genetics 139(2):1077–1090
Khare CP (2007) Indian medicinal plants: an illustrated dictionary. Springer, Heidelberg
Kress WJ, Wurdack KJ, Zimmer EA, Weigt LA, Janzen DH (2005) Use of DNA barcodes to identify flowering plants. Proc Natl Acad Sci USA 102(23):8369–8374
Kress WJ, Erickson DL, Jones FA, Swenson NG, Perez R et al (2009) Plant DNA barcodes and a community phylogeny of a tropical forest dynamics plot in Panama. Proc Natl Acad Sci USA 106(44):18621–18626
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 70 for bigger data sets. Mol Biol Evol 33(7):1870–1874
Levin RA, Wagner WL, Hoch PC et al (2003) Family-level relationships of onagraceae based on chloroplast rbcL and ndhF data. Am J Bot 90(1):107–115
Mooi E, Sarstedt MA (2011) Concise guide to market research: cluster analysis. Springer, Heidelberg, pp 237–284
Nadkarni AK (1976) Indian meteria medica, vol 1. Popular Prakashan, Mumbai
Nei M (1987) Molecular evolutionary genetics. Columbia Univ Press, New York
Palmblad M, Deelder AM (2012) Molecular phylogenetics by direct comparison of tandem mass spectra. Rapid Commun Mass Spectrom 26(7):728–732
Patra A, Deya AK, Kundu AB, Purushothaman KK, Saraswathy A (1992) Shoreaphenol, a polyphenol from Shorea robusta. Phytochemistry 31(7):2561–2562
Peakall ROD, Smouse PE (2012) GenAlEx 6.5: genetic analysis in excel. Population genetic software for teaching and research-an update. Bioinformatics 28(19):2537–2539
Pendkar SK, Hegde S, Nayak SU, Hegde H, Kholkute SD, Roy S (2016) Detection of adulteration by Wedelia calendulacea in Eclipta alba through ISSR and RAPD Markers. Planta Med Int Open 3(02):e43–e46
Perrier X, Jacquemoud-Collet, JP (2006) DARwin software. http://darwin.cirad.fr/
Rasmussen HN, Rasmussen OF, Andersen JK, Olsen JE (1994) Specific detection of pathogenic Yersinia enterocolitica by two-step PCR using hot-start and DMSO. Mol Cell Probes 8(2):99–108
Richards E, Reichardt M, Rogers S (1994) Preparation of genomic DNA from plant tissue. In: Ausubel FM, Brent R, Kingston RE, Moore DD, Seidman JG, Smith JA, Struhl K (Eds), Curr Protoc Mol Biol, John Wiley & Sons, New York, USA
Rozas J, Sánchez-Del Barrio JC, Messeguer X, Rozas R (2003) DnaSP, DNA polymorphism analyses by the coalescent and other methods. Bioinformatics 19(18):2496–2497
Sarin YK (1996) Illustrated Herbs of Ayurveda New Delhi, India. Council of Scientific and Industrial Research (CSIR) and Indian Council of Medical Research (ICMR), New Delhi
Schaal BA, Hayworth DA, Olsen KM, Rauscher JT, Smith WA (1998) Phylogeographic studies in plants: problems and prospects. Mol Ecol 7(4):465–474
Shukla S, Hegde S, Kumar A, Chaudhary G, Tewari SK, Upreti DK, Pal M (2018) Fatty acid composition and antibacterial potential of Cassia tora (leaves and stem) collected from different geographic areas of India. J Food Drug Anal 26(1):107–111
Singh S, Krishna THA, Kamalraj S, Kuriakose GC, Valayil JM, Jayabaskaran C (2015) Phytomedicinal importance of Saraca asoca (Ashoka): an exciting past, an emerging present and a promising future. Curr Sci 109(10):1790–1801
Smillie TJ, Khan IA (2010) A comprehensive approach to identifying and authenticating botanical products. Clin Pharmacol Ther 87(2):175–186
Tandon N, Yadav SS (2017) Contributions of Indian Council of Medical Research (ICMR) in the area of Medicinal plants/Traditional medicine. J Ethnopharmacol 197:39–45
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22(22):4673–4680
Urumarudappa SKJ, Gogna N, Newmaster SG, Venkatarangaiah K, Subramanyam R, Saroja SG, Gudasalamani R, Dorai K, Ramanan US (2016) DNA barcoding and NMR spectroscopy-based assessment of species adulteration in the raw herbal trade of Saraca asoca (Roxb) Willd, an important medicinal plant. Int J Legal Med 130(6):1457–1470
Wan CY, Wilkins TA (1993) Spermidine facilitates PCR amplification of target DNA. PCR Methods Appl 3(3):208–210
Wang H, Khera P, Huang B, Yuan M, Katam R, Zhuang W, Harris-Shultz K, Moore KM, Culbreath AK, Zhang X, Varshney RK, Xie L, Guo B (2016) Analysis of genetic diversity and population structure of peanut cultivars and breeding lines from China, India and the US using simple sequence repeat markers. J Integr Plant Biol 58(5):452–465
Wolf JB, Brodie ED III, Cheverud JM, Moore AJ, Wade MJ (1998) Evolutionary consequences of indirect genetic effects. Trends Ecol Evol 13(2):64–69
Acknowledgements
Authors are thankful to the Director-in-Charge, ICMR-NITM, formerly RMRC, Belagavi, India, and Indian Council of Medical Research (Government of India), for supporting this study. Authors are also thankful to Dr. Vidya S. Gupta, Biochemical Science Division, National Chemical Laboratory, Pune, for technical suggestions and Bio-Medical Informatics Centre of ICMR at NITM, Belagavi, for informatics support. Authors extend thanks to Pramod Kumar and Jotiba B. Palekar, ICMR-NITM, Belagavi, Anil Bisht (Dehradun), Jayendrasinh Chavda (Gujarat) and Aparna J (Kerala), India, for help during sample collection. SH is grateful to ICMR for funding.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
This study was funded by The Indian Council of Medical Research, New Delhi (Grant No. 45/53/2013/BMS/TRM; IRIS ID: 2013-18920).
Conflict of interest
All authors declare that there is no conflict of interest.
Ethical approval
Not applicable.
Electronic supplementary material
Below is the link to the electronic supplementary material.
13205_2018_1175_MOESM1_ESM.pptx
Fig. 1a: ISSR amplification profile from genomic DNA of 41 individuals of Saraca asoca; M: 100+500bp Molecular weight markers; NC: Negative control. Fig. 1b: ISSR amplification profile from genomic DNA samples of BV (1-5), SR (1-5), PL (1-5), MF (1-5) and TO (1-5); M: 100+500bp Molecular weight markers; NC: Negative control (PPTX 1666 kb)
Rights and permissions
About this article
Cite this article
Hegde, S., Saini, A., Hegde, H.V. et al. Molecular identification of Saraca asoca from its substituents and adulterants. 3 Biotech 8, 161 (2018). https://doi.org/10.1007/s13205-018-1175-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13205-018-1175-5