Abstract
The present review focuses on the role of miRNAs in the control of white adipose tissue browning, a process which describes the recruitment of adipocytes showing features of brown adipocytes in white adipose tissue. MicroRNAs (miRNAs) are a class of short non-coding RNAs (19–22 nucleotides) involved in gene regulation. Although the main effect of miRNAs is the inhibition of the translational machinery, thereby preventing the production of the protein product, the activation of protein translation has also been described in the literature. In addition to modifying translation, miRNAs binding to its target mRNAs also trigger the recruitment and association of mRNA decay factors, leading to mRNA destabilization, degradation, and thus to the decrease in expression levels. Although a great number of miRNAs have been reported to potentially regulate genes that play important roles in the browning process, only a reduced number of studies have demonstrated experimentally an effect on this process associated to changes in miRNA expressions, so far. These studies have shown, by using either primary adipocyte cultures or experimental models of mice (KO mice, mice overexpressing a specific miRNA) that miR-196a, miR-26 and miR-30 are needed for browning process development. By contrast, miR-155, miR-133, miR-27b and miR-34 act as negative regulators of this process. Further studies are needed to fully describe the miRNA network-involved white adipose tissue browning regulation.

Similar content being viewed by others
References
Baek D, Villén J, Shin C, Camargo FD, Gygi SP, Bartel DP (2008) The impact of microRNAs on protein output. Nature 455:64–71
Bartel DP (2009) MicroRNAs: target recognition and regulatory functions. Cell 136:215–233
Bhaskaran M, Mohan M (2014) MicroRNAs: history, biogenesis, and their evolving role in animal development and disease. Vet Pathol 51:759–774
Cannon B, Nedergaard J (2004) Brown adipose tissue: function and physiological significance. Physiol Rev 84:277–359
Carey AL, Vorlander C, Reddy-Luthmoodoo M, Natoli AK, Formosa MF, Bertovic DA, Anderson MJ, Duffy SJ, Kingwell BA (2014) Reduced UCP-1 content in in vitro differentiated beige/brite adipocytes derived from preadipocytes of human subcutaneous white adipose tissues in obesity. PLoS One 9:e91997
Ceppi M, Pereira PM, Dunand-Sauthier I, Barras E, Reith W, Santos MA, Pierre P (2009) MicroRNA-155 modulates the interleukin-1 signaling pathway in activated human monocyte-derived dendritic cells. Proc Natl Acad Sci U S A 106:2735–2740
Chen Y, Siegel F, Kipschull S, Haas B, Fröhlich H, Meister G, Pfeifer A (2013) miR-155 regulates differentiation of brown and beige adipocytes via a bistable circuit. Nat Commun 4:1769
Chi SW, Zang JB, Mele A, Darnell RB (2009) Argonaute HITS-CLIP decodes microRNA-mRNA interaction maps. Nature 460:479–486
Costinean S, Sandhu SK, Pedersen IM, Tili E, Trotta R, Perrotti D, Ciarlariello D, Neviani P, Harb J, Kauffman LR, Shidham A, Croce CM (2009) Src homology 2 domain-containing inositol-5-phosphatase and CCAAT enhancer-binding protein beta are targeted by miR-155 in B cells of Emicro-MiR-155 transgenic mice. Blood 114:1374–1382
D’Alessandra Y, Devanna P, Limana F, Straino S, Di Carlo A, Brambilla PG, Rubino M, Carena MC, Spazzafumo L, De Simone M, Micheli B, Biglioli P, Achilli F, Martelli F, Maggiolini S, Marenzi G, Pompilio G, Capogrossi MC (2010) Circulating microRNAs are new and sensitive biomarkers of myocardial infarction. Eur Heart J 31:2765–2773
Elsen M, Raschke S, Eckel J (2014) Browning of white fat: does irisin play a role in humans? J Endocrinol 222:R25–38
Forman J (2010) The code within the code: microRNAs target coding region. In: HA C (ed)Cell cycle, p 1533-1541
Fu T, Seok S, Choi S, Huang Z, Suino-Powell K, Xu HE, Kemper B, Kemper JK (2014) MicroRNA 34a inhibits beige and brown fat formation in obesity in part by suppressing adipocyte fibroblast growth factor 21 signaling and SIRT1 function. Mol Cell Biol 34:4130–4142
Gabriely G, Wurdinger T, Kesari S, Esau CC, Burchard J, Linsley PS, Krichevsky AM (2008) MicroRNA 21 promotes glioma invasion by targeting matrix metalloproteinase regulators. Mol Cell Biol 28:5369–5380
Garzon R, Calin GA, Croce CM (2009) MicroRNAs in cancer. Annu Rev Med 60:167–179
Gennarino VA, Sardiello M, Avellino R, Meola N, Maselli V, Anand S, Cutillo L, Ballabio A, Banfi S (2009) MicroRNA target prediction by expression analysis of host genes. Genome Res 19:481–490
Georges SA, Biery MC, Kim SY, Schelter JM, Guo J, Chang AN, Jackson AL, Carleton MO, Linsley PS, Cleary MA, Chau BN (2008) Coordinated regulation of cell cycle transcripts by p53-inducible microRNAs, miR-192 and miR-215. Cancer Res 68:10105–10112
Grimson A, Farh KK, Johnston WK, Garrett-Engele P, Lim LP, Bartel DP (2007) MicroRNA targeting specificity in mammals: determinants beyond seed pairing. Mol Cell 27:91–105
Hafner M, Landthaler M, Burger L, Khorshid M, Hausser J, Berninger P, Rothballer A, Ascano M, Jungkamp AC, Munschauer M, Ulrich A, Wardle GS, Dewell S, Zavolan M, Tuschl T (2010) Transcriptome-wide identification of RNA-binding protein and microRNA target sites by PAR-CLIP. Cell 141:129–141
He M, Xu Z, Ding T, Kuang DM, Zheng L (2009) MicroRNA-155 regulates inflammatory cytokine production in tumor-associated macrophages via targeting C/EBPbeta. Cell Mol Immunol 6:343–352
He J, Zhang JF, Yi C, Lv Q, Xie WD, Li JN, Wan G, Cui K, Kung HF, Yang J, Yang BB, Zhang Y (2010) miRNA-mediated functional changes through co-regulating function related genes. PLoS One 5:e13558
Helwak A, Kudla G, Dudnakova T, Tollervey D (2013) Mapping the human miRNA interactome by CLASH reveals frequent noncanonical binding. Cell 153:654–665
Himms-Hagen J, Melnyk A, Zingaretti MC, Ceresi E, Barbatelli G, Cinti S (2000) Multilocular fat cells in WAT of CL-316243-treated rats derive directly from white adipocytes. Am J Physiol Cell Physiol 279:C670–681
Hu W, Wang X, Ding X, Li Y, Zhang X, Xie P, Yang J, Wang S (2012) MicroRNA-141 represses HBV replication by targeting PPARA. PLoS One 7:e34165
Hu F, Wang M, Xiao T, Yin B, He L, Meng W, Dong M, Liu F (2015) miR-30 promotes thermogenesis and the development of beige fat by targeting RIP140. Diabetes 64:2056–2068
Iliopoulos D, Malizos KN, Oikonomou P, Tsezou A (2008) Integrative microRNA and proteomic approaches identify novel osteoarthritis genes and their collaborative metabolic and inflammatory networks. PLoS One 3:e3740
Jespersen NZ, Larsen TJ, Peijs L, Daugaard S, Homøe P, Loft A, de Jong J, Mathur N, Cannon B, Nedergaard J, Pedersen BK, Møller K, Scheele C (2013) A classical brown adipose tissue mRNA signature partly overlaps with brite in the supraclavicular region of adult humans. Cell Metab 17:798–805
Kang T, Lu W, Xu W, Anderson L, Bacanamwo M, Thompson W, Chen YE, Liu D (2013) MicroRNA-27 (miR-27) targets prohibitin and impairs adipocyte differentiation and mitochondrial function in human adipose-derived stem cells. J Biol Chem 288:34394–34402
Karbiener M, Fischer C, Nowitsch S, Opriessnig P, Papak C, Ailhaud G, Dani C, Amri EZ, Scheideler M (2009) microRNA miR-27b impairs human adipocyte differentiation and targets PPARgamma. Biochem Biophys Res Commun 390:247–251
Karbiener M, Pisani DF, Frontini A, Oberreiter LM, Lang E, Vegiopoulos A, Mössenböck K, Bernhardt GA, Mayr T, Hildner F, Grillari J, Ailhaud G, Herzig S, Cinti S, Amri EZ, Scheideler M (2014) MicroRNA-26 family is required for human adipogenesis and drives characteristics of brown adipocytes. Stem Cells 32:1578–1590
Kim YJ, Bae SW, Yu SS, Bae YC, Jung JS (2009) miR-196a regulates proliferation and osteogenic differentiation in mesenchymal stem cells derived from human adipose tissue. J Bone Miner Res 24:816–825
Kim SY, Kim AY, Lee HW, Son YH, Lee GY, Lee JW, Lee YS, Kim JB (2010) miR-27a is a negative regulator of adipocyte differentiation via suppressing PPARgamma expression. Biochem Biophys Res Commun 392:323–328
Kong X, Yu J, Bi J, Qi H, Di W, Wu L, Wang L, Zha J, Lv S, Zhang F, Li Y, Hu F, Liu F, Zhou H, Liu J, Ding G (2015) Glucocorticoids transcriptionally regulate miR-27b expression promoting body fat accumulation via suppressing the browning of white adipose tissue. Diabetes 64:393–404
Kosaka N, Ochiya T (2011) Unraveling the mystery of cancer by secretory microRNA: horizontal microRNA transfer between living cells. Front Genet 2:97
Krol J, Loedige I, Filipowicz W (2010) The widespread regulation of microRNA biogenesis, function and decay. Nat Rev Genet 11:597–610
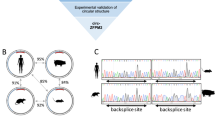
Kuhn DE, Martin MM, Feldman DS, Terry AV, Nuovo GJ, Elton TS (2008) Experimental validation of miRNA targets. Methods 44:47–54
Lee JJ, Drakaki A, Iliopoulos D, Struhl K (2012) MiR-27b targets PPARγ to inhibit growth, tumor progression and the inflammatory response in neuroblastoma cells. Oncogene 31:3818–3825
Lee P, Greenfield JR, Ho KK, Fulham MJ (2010) A critical appraisal of the prevalence and metabolic significance of brown adipose tissue in adult humans. Am J Physiol Endocrinol Metab 299:E601–606
Lee EK, Lee MJ, Abdelmohsen K, Kim W, Kim MM, Srikantan S, Martindale JL, Hutchison ER, Kim HH, Marasa BS, Selimyan R, Egan JM, Smith SR, Fried SK, Gorospe M (2011) miR-130 suppresses adipogenesis by inhibiting peroxisome proliferator-activated receptor gamma expression. Mol Cell Biol 31:626–638
Li Y, Zhang M, Chen H, Dong Z, Ganapathy V, Thangaraju M, Huang S (2010) Ratio of miR-196s to HOXC8 messenger RNA correlates with breast cancer cell migration and metastasis. Cancer Res 70:7894–7904
Lin Q, Gao Z, Alarcon RM, Ye J, Yun Z (2009) A role of miR-27 in the regulation of adipogenesis. FEBS J 276:2348–2358
Liu W, Bi P, Shan T, Yang X, Yin H, Wang YX, Liu N, Rudnicki MA, Kuang S (2013) miR-133a regulates adipocyte browning in vivo. miR-133a regulates adipocyte browning in vivo 9:e1003626
Lorenzen JM, Martino F, Thum T (2012) Epigenetic modifications in cardiovascular disease. Basic Res Cardiol 107:245
Martinelli R, Nardelli C, Pilone V, Buonomo T, Liguori R, Castanò I, Buono P, Masone S, Persico G, Forestieri P, Pastore L, Sacchetti L (2010) miR-519d overexpression is associated with human obesity. Obesity (Silver Spring) 18:2170–2176
Mori M, Nakagami H, Rodriguez-Araujo G, Nimura K, Kaneda Y (2012) Essential role for miR-196a in brown adipogenesis of white fat progenitor cells. PLoS Biol 10:e1001314
Mueller DW, Bosserhoff AK (2011) MicroRNA miR-196a controls melanoma-associated genes by regulating HOX-C8 expression. Int J Cancer 129:1064–1074
Ouellet V, Labbé SM, Blondin DP, Phoenix S, Guérin B, Haman F, Turcotte EE, Richard D, Carpentier AC (2012) Brown adipose tissue oxidative metabolism contributes to energy expenditure during acute cold exposure in humans. J Clin Invest 122:545–552
Petrovic N, Walden TB, Shabalina IG, Timmons JA, Cannon B, Nedergaard J (2010) Chronic peroxisome proliferator-activated receptor gamma (PPARgamma) activation of epididymally derived white adipocyte cultures reveals a population of thermogenically competent, UCP1-containing adipocytes molecularly distinct from classic brown adipocytes. J Biol Chem 285:7153–7164
Rao DS, O’Connell RM, Chaudhuri AA, Garcia-Flores Y, Geiger TL, Baltimore D (2010) MicroRNA-34a perturbs B lymphocyte development by repressing the forkhead box transcription factor Foxp1. Immunity 33:48–59
Romao JM, Jin W, Dodson MV, Hausman GJ, Moore SS, Guan LL (2011) MicroRNA regulation in mammalian adipogenesis. Exp Biol Med (Maywood) 236:997–1004
Rosenwald M, Perdikari A, Rülicke T, Wolfrum C (2013) Bi-directional interconversion of brite and white adipocytes. Nat Cell Biol 15:659–667
Rothwell NJ, Stock MJ (1983) Luxuskonsumption, diet-induced thermogenesis and brown fat: the case in favour. Clin Sci (Lond) 64:19–23
Sanchez-Gurmaches J, Hung CM, Sparks CA, Tang Y, Li H, Guertin DA (2012) PTEN loss in the Myf5 lineage redistributes body fat and reveals subsets of white adipocytes that arise from Myf5 precursors. Cell Metab 16:348–362
Schimanski CC, Frerichs K, Rahman F, Berger M, Lang H, Galle PR, Moehler M, Gockel I (2009) High miR-196a levels promote the oncogenic phenotype of colorectal cancer cells. World J Gastroenterol 15:2089–2096
Seale P, Bjork B, Yang W, Kajimura S, Chin S, Kuang S, Scimè A, Devarakonda S, Conroe HM, Erdjument-Bromage H, Tempst P, Rudnicki MA, Beier DR, Spiegelman BM (2008) PRDM16 controls a brown fat/skeletal muscle switch. Nature 454:961–967
Selbach M, Schwanhäusser B, Thierfelder N, Fang Z, Khanin R, Rajewsky N (2008) Widespread changes in protein synthesis induced by microRNAs. Nature 455:58–63
Shan T, Liang X, Bi P, Zhang P, Liu W, Kuang S (2013) Distinct populations of adipogenic and myogenic Myf5-lineage progenitors in white adipose tissues. J Lipid Res 54:2214–2224
Spiegelman BM (2013) Banting Lecture 2012: Regulation of adipogenesis: toward new therapeutics for metabolic disease. Diabetes 62:1774–1782
Sun L, Trajkovski M (2014) MiR-27 orchestrates the transcriptional regulation of brown adipogenesis. Metabolism 63:272–282
Tanzer A, Stadler PF (2006) Evolution of microRNAs. Methods Mol Biol 342:335–350
Tavazoie SF, Alarcón C, Oskarsson T, Padua D, Wang Q, Bos PD, Gerald WL, Massagué J (2008) Endogenous human microRNAs that suppress breast cancer metastasis. Nature 451:147–152
Timmons JA, Wennmalm K, Larsson O, Walden TB, Lassmann T, Petrovic N, Hamilton DL, Gimeno RE, Wahlestedt C, Baar K, Nedergaard J, Cannon B (2007) Myogenic gene expression signature establishes that brown and white adipocytes originate from distinct cell lineages. Proc Natl Acad Sci U S A 104:4401–4406
Trajkovski M, Ahmed K, Esau CC, Stoffel M (2012) MyomiR-133 regulates brown fat differentiation through Prdm16. Nat Cell Biol 14:1330–1335
van Marken Lichtenbelt WD, Vanhommerig JW, Smulders NM, Drossaerts JM, Kemerink GJ, Bouvy ND, Schrauwen P, Teule GJ (2009) Cold-activated brown adipose tissue in healthy men. N Engl J Med 360:1500–1508
van Rooij E (2011) The art of microRNA research. Circ Res 108:219–234
Vasudevan S, Tong Y, Steitz JA (2007) Switching from repression to activation: microRNAs can up-regulate translation. Science 318:1931–1934
Waldén TB, Hansen IR, Timmons JA, Cannon B, Nedergaard J (2012) Recruited vs. nonrecruited molecular signatures of brown, “brite,” and white adipose tissues. Am J Physiol Endocrinol Metab 302:E19–31
Wang B, Hsu SH, Frankel W, Ghoshal K, Jacob ST (2012) Stat3-mediated activation of microRNA-23a suppresses gluconeogenesis in hepatocellular carcinoma by down-regulating glucose-6-phosphatase and peroxisome proliferator-activated receptor gamma, coactivator 1 alpha. Hepatology 56:186–197
Worm J, Stenvang J, Petri A, Frederiksen KS, Obad S, Elmén J, Hedtjärn M, Straarup EM, Hansen JB, Kauppinen S (2009) Silencing of microRNA-155 in mice during acute inflammatory response leads to derepression of c/ebp Beta and down-regulation of G-CSF. Nucleic Acids Res 37:5784–5792
Wu J, Boström P, Sparks LM, Ye L, Choi JH, Giang AH, Khandekar M, Virtanen KA, Nuutila P, Schaart G, Huang K, Tu H, van Marken Lichtenbelt WD, Hoeks J, Enerbäck S, Schrauwen P, Spiegelman BM (2012) Beige adipocytes are a distinct type of thermogenic fat cell in mouse and human. Cell 150:366–376
Yang Z, Bian C, Zhou H, Huang S, Wang S, Liao L, Zhao RC (2011) MicroRNA hsa-miR-138 inhibits adipogenic differentiation of human adipose tissue-derived mesenchymal stem cells through adenovirus EID-1. Stem Cells Dev 20:259–267
Yekta S, Shih IH, Bartel DP (2004) MicroRNA-directed cleavage of HOXB8 mRNA. Science 304:594–596
Yin Q, McBride J, Fewell C, Lacey M, Wang X, Lin Z, Cameron J, Flemington EK (2008) MicroRNA-155 is an Epstein-Barr virus-induced gene that modulates Epstein-Barr virus-regulated gene expression pathways. J Virol 82:5295–5306
Yin Q, Wang X, Fewell C, Cameron J, Zhu H, Baddoo M, Lin Z, Flemington EK (2010) MicroRNA miR-155 inhibits bone morphogenetic protein (BMP) signaling and BMP-mediated Epstein-Barr virus reactivation. J Virol 84:6318–6327
Ying SY, Chang DC, Miller JD, Lin SL (2006) The microRNA: overview of the RNA gene that modulates gene functions. Methods Mol Biol 342:1–18
Zampetaki A, Kiechl S, Drozdov I, Willeit P, Mayr U, Prokopi M, Mayr A, Weger S, Oberhollenzer F, Bonora E, Shah A, Willeit J, Mayr M (2010) Plasma microRNA profiling reveals loss of endothelial miR-126 and other microRNAs in type 2 diabetes. Circ Res 107:810–817
Zaragosi LE, Wdziekonski B, Brigand KL, Villageois P, Mari B, Waldmann R, Dani C, Barbry P (2011) Small RNA sequencing reveals miR-642a-3p as a novel adipocyte-specific microRNA and miR-30 as a key regulator of human adipogenesis. Genome Biol 12:R64
Zheng L, Lv GC, Sheng J, Yang YD (2010) Effect of miRNA-10b in regulating cellular steatosis level by targeting PPAR-alpha expression, a novel mechanism for the pathogenesis of NAFLD. J Gastroenterol Hepatol 25:156–163
Zhou J, Wang KC, Wu W, Subramaniam S, Shyy JY, Chiu JJ, Li JY, Chien S (2011) MicroRNA-21 targets peroxisome proliferators-activated receptor-alpha in an autoregulatory loop to modulate flow-induced endothelial inflammation. Proc Natl Acad Sci U S A 108:10355–10360
Acknowledgments
This study was supported by grants from Instituto de Salud Carlos III (CIBERobn), the Government of the Basque Country (IT-512-13 and S-PR13UN004 RESBRITE), and University of the Basque Country (UPV/EHU) (ELDUNANOTEK UFI11/32).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Rights and permissions
About this article
Cite this article
Arias, N., Aguirre, L., Fernández-Quintela, A. et al. MicroRNAs involved in the browning process of adipocytes. J Physiol Biochem 72, 509–521 (2016). https://doi.org/10.1007/s13105-015-0459-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13105-015-0459-z