Abstract
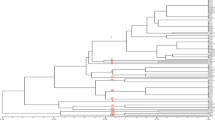
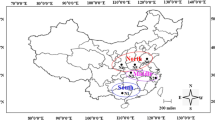
AG-A belongs to the binucleate Rhizoctonia (BNR) anastomosis group (AG) of the Ceratobasidium teleomorph, which parasitizes the roots of many plant species. Ninety nine isolate species of AG-A were obtained from Tibet, Sichuan, and Yunnan Province in China. All isolates were divided into three types based on their cultural characteristics. Type I: abundant aerial mycelia, dense hyphae, loose sclerotia; Type II: abundant aerial mycelia, no sclerotia. Type III: sparse aerial mycelium and no sclerotia. All of the isolates infected the seedlings of Chinese mustard and Chinese cabbage, causing the formation of lesions on the stem and a brown discoloration of the roots. Sequence analysis of the 5.8S rDNA-ITS showed a similarity of 98–100% among the isolates. Inter Simple Sequence Repeat (ISSR) was used to detect genetic variation in binucleate Rhizoctonia spp. Forty two AG-A isolates were amplified using 15 random primers. From a total of 164 bands, 144 bands (87.8%) were polymorphic in the 42 tested isolates. A dendrogram showing genetic relationships between the isolates was constructed using unweighted pair-group averages based on genetic distances. According to the dendrogram, the 42 tested isolates could be aligned into three clusters with a genetic similarity coefficient of 0.29, the first clusters including 27 isolates with III of culture characteristics on PDA; the second clusters included eight isolates with I of cultural characteristics on PDA; the third cluster included seven isolates with II of cultural characteristics on PDA. The results of ISSR analysis showed an association between the hosts of these isolates. Our results showed that ISSR analysis can reveal more molecular variation among isolates of AG-A than sequence analysis using the 5.8S rDNA-ITS.





Similar content being viewed by others
References
Guleria, S., Aggarwal, R., Thind, T. S., & Sharma, T. R. (2007). Morphological and pathological variability in rice isolates of Rhizoctonia solani and molecular analysis of their genetic variability. Journal of Phytopathology, 155, 654–661.
Hillis, D. M., & Dixon, M. T. (1991). Ribosomal DNA: Molecular evolution and phylogenetic inference. The Quarterly Review of Biology, 66, 411–453.
Hong, S. B., Go, S. J., Ryu, J. C., Kim, W. G., & Kim, I. S. (1998). Differentiation of intraspecific groups within Korean isolates of Rhizoctonia solani using PCR-RFLP of ribosomal DNA. Korean Journal of Plant Pathology, 14, 157–163. (in Korean, with English abstract).
Hyakumachi, M., Priyatmojo, A., Kubota, M., & Fukui, H. (2005). New anastomosis groups, AG-T and AG-U, of binucleate Rhizoctonia causing root and stem rot of cut-flower and miniature roses. Phytopathology, 95, 784–792.
Ichielevich-Auster, M., Sneh, B., Barash, I., & Koltin, Y. (1985). Pathogenicity, host specificity and anastomosis groups of Rhizoctonia spp. isolated from soils in Israel. Phytoparasitica, 13, 103–112.
Kuninaga, S., Natsuaki, T., & Takeuchi, T. (1997). Sequence variation of the rDNA- ITS regions within and between anastomosis groups in Rhizoctonia solani. Current Genetics, 32, 237–243.
Kuzoff, R. K., Sweere, J. A., Soltis, D. E., Soltis, P. S., & Zimmer, E. A. (1998). The phylogenetic potential of entire 26S rDNA sequences in plants. Molecular Biology and Evolution, 15, 251–263.
Larson, A. (1991). Evolutionary analysis of length-variable sequences: Divergent domains of ribosomal RNA. In M. M. Miyamoto & J. Cracraft (Eds.), Phylogenetic analysis of DNA sequences (pp. 221–248). New York, NY: Oxford University Press.
Mahdiyeh, K., Naser, S., & Masoud, S. (2009). Genetic diversity of Iranian AG1-IA isolates of Rhizoctonia solani, the cause of rice sheath blight, using morphological and molecular markers. Journal of Phytopathology, 157, 708–714.
Martin, S. B. (1988). Identification, isolation frequency, and pathogenicity of anastomosis groups of binucleate Rhizoctonia spp. from strawberry roots. Phytopathology, 78, 379–384.
Mazzola, M. (1997). Identification and pathogenicity of Rhizoctonia spp. isolated from apple roots and orchard soils. Phytopathology, 87, 582–587.
Raeder, V., & Broda, P. (1985). Rapid preparation of DNA from filaments fungi. Letters in Applied Microbiology, 1, 17–20.
Salazar, O., Julian, M. C., & Rubio, V. (2000). Primers based on specific rDNA-ITS sequences for PCR detection of Rhizoctonia solani, R. solani AG2 subgroups and ecological types, and binucleate Rhizoctonia. Mycological Research, 104, 281–285.
Sharon, M., Kuninaga, S., Hyakumachi, M., Naito, S., & Sneh, B. (2008). Classification of Rhizoctonia spp. using rDNA-ITS sequence analysis supports the genetic basis of the classical anastomosis grouping. Mycoscience, 49, 93–114.
Sneh, B., Sharon, M., Kuninaga, S., & Hyakumachi, M. (2006). The advancing identification and classification of Rhizoctonia spp. using molecular and biotechnological methods compared with the classical anastomosis grouping. Mycoscience, 47, 299–316.
Sneh, B., Burpee, L., & Ogoshi, A. (1998). Identification of Rhizoctonia species. St. Paul, MN, USA: The American Phytopathological Society.
Spatafora, J. W., & Blackwell, M. (1993). Molecular systematics of unitunicate perithecial ascomycetes: The Clavicipitales–Hypocreales connection. Mycologia, 85, 912–922.
Stepansky, A., Kovalski, R., & Perl-Treves, I. (1999). Intraspecific classification of melons (Cucumis melo L.) in view of their phenotypic and molecular variation. Plant Systematics and Evolution, 217, 313–332.
Swann, E. C., & Taylor, J. W. (1993). Higher taxa of Basidiomycetes: An 18SrRNA gene perspective. Mycologia, 85, 923–936.
Toda, T., Hyakumachi, M., Suga, H., Kageyama, K., Tanaka, A., & Tani, T. (1999). Differentiation of Rhizoctonia AG-D isolates from turfgrass into subgroups I and II based on rDNA and RAPD analysis. European Journal of Plant Pathology, 105, 835–846.
Toda, T., Nasu, H., Kageyama, K., & Hyakumachi, M. (1998). Genetic identification of web-blight fungus (Rhizoctonia solani AG1) obtained from European pear using RFLP of rDNA-ITS and RAPD analyses. Research Bulletin of the Faculty College of Agriculture Gifu University, 63, 1–9.
Vilgalys, R., & Sun, B. L. (1994). Ancient and recent patterns of geographic speciation in the oyster mushroom Pleurotus revealed by phylogenetic analysis of ribosomal DNA sequences. Proceedings of the National Academy of Sciences of the United States of America, 91, 4599–4603.
Yang, G. H., Chen, H. R., Naito, S., & Ogoshi, A. (2005a). Characterization of Rhizoctonia solania AG 4HG-III causing stem canker and wirestem on Green amaranth and Chinese amaranth. Journal of Phytopathology, 153, 185–187.
Yang, G. H., Chen, H. R., Naito, S., Ogoshi, A., & Deng, Y. L. (2005b). First report of AG-A of binucleate Rhizoctonia in China, pathogenic to soybean, pea, snap bean and pak choy. Journal of Phytopathology, 153, 333–336.
Yang, G. H., Chen, H. R., Naito, S., Wu, J. Y., He, X. H., & Duan, C. F. (2005c). Occurrence of foliar rot of pak choy and Chinese mustard caused by Rhizoctonia solani AG-1 IB in China. Journal of General Plant Pathology, 71, 377–379.
Zhang, M., & Dernoeden, P. H. (1995). Facilitating anastomosis grouping of Rhizoctonia solani isolates from cool season turfgrasses. Hortscience, 30, 1260–1262.
Acknowledgments
The research was supported by National Natural Science Fund (30660006) and Science and technology projects in Yunnan Tobacco Company (09YN006). We appreciate the assistance of Dr. Robert L. Conner of the Morden Research Station of Agriculture and Agri-Food Canada and Prof. Michael A. Fullen of The University of Wolverhampton, UK, who reviewed this manuscript very carefully.
Author information
Authors and Affiliations
Corresponding author
Additional information
L. P. Lei is a Co-first author.
Rights and permissions
About this article
Cite this article
Li, Y.Q., Lei, L.P., Dong, W.H. et al. Molecular diversity of binucleate Rhizoctonia AG-A in China. Phytoparasitica 39, 461–470 (2011). https://doi.org/10.1007/s12600-011-0187-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12600-011-0187-z