Abstract
Calanus glacialis, a major contributor to zooplankton biomass in the Arctic shelf seas, is a key link between primary production and higher trophic levels that may be sensitive to climate warming. The aim of this study was to explore genetic variation in contemporary populations of this species to infer possible changes during the Quaternary period, and to assess its population structure in both space and time. Calanus glacialis was sampled in the fjords of Spitsbergen (Hornsund and Kongsfjorden) in 2003, 2004, 2006, 2009 and 2012. The sequence of a mitochondrial marker, belonging to the ND5 gene, selected for the study was 1249 base pairs long and distinguished 75 unique haplotypes among 140 individuals that formed three main clades. There was no detectable pattern in the distribution of haplotypes by geographic distance or over time. Interestingly, a Bayesian skyline plot suggested that a 1000-fold increase in population size occurred approximately 10,000 years before present, suggesting a species expansion after the Last Glacial Maximum.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
A large part of zooplankton biomass in the Arctic shelf seas is formed by Calanus glacialis (Fleminger and Hulseman, 1977; Blachowiak-Samolyk et al. 2008; Weydmann et al. 2013), a lipid-rich calanoid grazer. In the lipid-based Arctic food web, it is an essential link between the low-energy microalgae and higher trophic levels (Lee and Hirota 1973; Falk-Petersen et al. 2009). Its life cycle is between one (MacLellan 1967; Weydmann et al. 2013) and three years (Kosobokova 1999), depending on the region and environmental conditions, although C. glacialis typically has a 2-year life span (Hirche and Kwaśniewski 1997). The areas of its occurrence, including peripheral seas of the Arctic Ocean and adjacent regions of the North Atlantic and Pacific Oceans (Jashnov 1970; Conover 1988), are now facing intensive modifications from an unprecedented combination of environmental changes, such as increasing ocean temperatures and reduction in sea ice extent, caused by climate warming (IPCC 2014), with the record high Atlantic Water temperature and salinity in 2006 (Walczowski et al. 2012). With the Arctic region likely to continue warming more rapidly than the global mean (IPCC 2014), changes are expected to affect Arctic marine biota. For example, the loss of sea ice represents a loss of critical habitat for ice-related species, such as C. glacialis, that needs energy from the ice algal bloom to fuel its reproduction (Søreide et al. 2010). Major changes in the function of the Arctic marine ecosystem are now anticipated.
The capacity for populations to evolve in response to environmental changes is based on genetic diversity, which encompasses the variation among individuals within a population and the genetic variation among populations (Gray 1997; Kenchington et al. 2003; Reed and Frankham 2003). Climatic changes during the Quaternary period in Arctic regions, with repeated glacial and interglacial periods causing cyclical expansions and contractions of species, have shaped their genetic variation and genealogies. During this time, some populations and lineages became extinct, while others underwent bottlenecks and founder events. Mitochondrial markers, which provide suitably variable sequences, are among the most favored for tracking such events during the Quaternary (Hewitt 2004).
To adequately assess patterns of genetic diversity at the population level, fast-evolving markers should be used, particularly for the study of animal populations that have expanded substantially since the Last Glacial Maximum 10,000–14,000 years ago (Baker 2000). To date, few Arctic species have been studied in detail. The lack of adequate polymorphic makers was one of the factors limiting genetic research in Calanus spp., although several microsatellite markers have recently been published (Provan et al. 2007; Provan et al. 2009; Parent et al. 2012; Weydmann et al. 2014). Provan et al. (2009) used microsatellite markers and mitochondrial cytochrome b gene (CYTB) in Calanus finmarchicus, revealing no significant genetic differentiation at the inter-population level or across the species’ range, in either nuclear or mitochondrial data sets. The authors postulated that these results indicated high levels of dispersal and a constant effective population size over the period 359,000–566,000 years before present, and suggested that C. finmarchicus possessed the capacity to track changes in available habitat, a feature that may be of crucial importance for the species’ ability to cope with the current period of global climate change. However, similar studies have not been conducted on its Arctic sibling, C. glacialis. Using the 16S ribosomal RNA gene, Nelson et al. (2009) defined two genetically distinct C. glacialis populations—an Arctic and a North Pacific (Bering Sea) population—although the latter was not reproductively established in the Arctic Ocean. The authors suggested that climate warming could increase opportunities for southern organisms to become established in the Arctic. In contrast, Weydmann et al. (2016), on the basis of microsatellite markers, reported a panmictic population of C. glacialis with large-scale gene flow around the Arctic.
Here we aimed to estimate genetic variation in contemporary populations of Calanus glacialis to examine possible changes during the Quaternary, especially after the Last Glacial Maximum, when major latitudinal species range shifts occurred. Additionally, we wished to assess the recent population structure of this key Arctic zooplankton species at both geographic and temporal scales. To this end, we chose two Spitsbergen fjords (Svalbard Archipelago) that are contrasting in terms of water masses, in addition to the availability of samples from a time series collected between 2003 and 2012. Finally, we based our study on a newly developed mitochondrial marker, chosen based on the length of mitochondrial genes, and intermediate intra- and interspecies polymorphism, which is greater than those used to date.
Material and methods
Study area
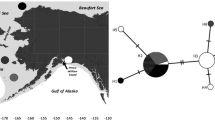
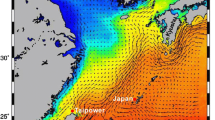
Our study area covered two fjords in the Atlantic sector of the Arctic Ocean. Hornsund is a medium-sized fjord located in the southwest part of Spitsbergen (Fig. 1). The fjord is under the influence of the cold coastal South Cape Current and warmer, more saline West Spitsbergen Current. The inner fjord basin, Brepollen, is isolated from the main basin by an underwater sill establishing a reservoir of winter cooled water throughout all seasons (Swerpel 1985), where a local population of C. glacialis was reported to exist (Weydmann and Kwaśniewski 2008).
Kongsfjorden is an open fjord situated on the west coast of Spitsbergen. Due to the absence of a sill at the entrance, the fjord faces strong pulsed influxes of relatively warm Atlantic water (Cottier et al. 2005). Despite the fjord’s location at 79°N latitude, the fauna of Kongsfjorden is of a rather sub-arctic character due to the strong influence of the West Spitsbergen Current and advection processes (Kwaśniewski et al. 2003; Walkusz et al. 2009).
Sampling
Zooplankton samples were collected from the fjords during the summers of 2003 (Hornsund), 2004 (Kongsfjorden), 2006 (Hornsund), 2009 (Kongsfjorden), and 2012 (both fjords) (Table 1) during the Arctic cruises of the R/V Oceania, using a WP-2 mesozooplankton net (0.25 m2 mouth opening; 180 μm mesh size), and were preserved in 96% ethanol, which was changed 24 h after sampling.
DNA extraction and amplification
In total, 140 Calanus glacialis individuals of the fifth copepodite stage and adult females were identified to the species level based on the prosome length (Weydmann and Kwasniewski 2008) and characteristic morphological features (Brodskii et al. 1983) and were retrieved from the mixed zooplankton samples. Their genomic DNA was extracted using the Sherlock AX kit (A&A Biotechnology).
Specific PCR primers (popF: 5’-AAGATACTTGGTATATTTCTGACACC-3’, popR: 5’-ATATTTATGTTGATTCTCAGCCC-3’) and a third sequencing primer (popR2 5’-TTCACAATATAAAAGATTACC-3’) were designed using sequences available in the NCBI database of sequence read archives (SRA, accession numbers SRR1793125, SRR1791606, SRR1791605, SRR1791524, SRR1791525) (Ramos et al. 2015). The PCR product, covering 1465 base pairs (bp) of the mitochondrial ND5 gene encoding the fifth subunit of the respiratory chain complex I (NADH dehydrogenase subunit 5), was obtained. This fragment was chosen based on the length of mitochondrial genes, their intermediate intra- and interspecies polymorphism in comparison to conserved CYTB, cytochrome oxidase subunit I (COI) and highly variable NADH dehydrogenase subunits 3 and 4 (ND3 and ND4) genes in the copepod subclass (Minxiao et al. 2011).
The final reaction volume for PCR amplification was 10 μl, with approximately 5 ng of total DNA, 0.5 μM of popF and popR primers, dNTPs at 200 μM each, 2 mM MgCl2 and 0.5 U of DyNAzyme EXT DNA Polymerase (Thermo Fisher Scientific), in a buffer supplied by the manufacturer. The PCR amplification protocol was as follows: initial denaturation at 95°C for 5 min, followed by 30 cycles of denaturation at 94°C for 30 s, annealing at 56°C for 30 s, and extension at 72°C for 2 min. The final extension lasted 5 min (TProfessional Gradient Cycler from Biometra).
PCR products were separated by a 1% agarose gel electrophoresis in 0.5X TBE buffer and visualized with ethidium bromide in UV light. Products which showed a strong band of the correct size were selected for sequencing. DNA concentration was estimated based on gel images, and the products were cleaned using Exonuclease I and alkaline phosphatase treatment (Werle et al. 1994). Sequencing was performed by Sanger technology, in both directions, using all three primers (Macrogen, Inc.).
Bioinformatic analysis
The raw sequence reads were assembled using Staden Package software (Staden 1996). The resulting partial ND5 sequences were aligned in MEGA6 (Tamura et al. 2013) using the ClustalW (Larkin et al. 2007) algorithm and trimmed to the same length of 1249 base pairs. The alignment was straightforward, as there were no indels in the sequences. All sequences were deposited in GenBank under accession numbers MF447532 - MF447671.
General diversity indices were calculated in DnaSP (Librado and Rozas 2009): haplotype diversity (Hd), which is expected to be high for organisms with large effective population sizes (Hd close to 1); nucleotide diversity (π), which is expected to be within 1% for intraspecies mitochondrial polymorphism; and Tajima’s D statistics used to test the departure of haplotype distribution from neutral expectations. To check for genetic differentiation among samples, analysis of molecular variance (AMOVA), including population pairwise fixation indices (ΦST, with no. of permutations for significance = 1000) was calculated in Arlequin 3.5 (Excoffier and Lischer 2010). A minimum spanning network (MSN) of all observed haplotypes was built using a median-joining algorithm (Bandelt et al. 1999) implemented in Network software (fluxus-engineering.com). This type of analysis is more appropriate for population-level data than classic phylogenetic tree building, and allows quick visual inspection of existing relationships between genetic diversity and other factors (such as geographic or temporal scales). To elucidate the demographic history of the studied population of C. glacialis, analysis of population size changes was performed in BEAST 2.4.5 (Bouckaert et al. 2014) using a Bayesian skyline plot (BSP) reconstruction approach. The best-fit model of substitutions (HKY+G) as well as clock model (relaxed uncorrelated lognormal clock) was selected using Bayes factor comparison (Baele et al. 2012). No constrains were used; therefore, the obtained plots were scaled in mutational units. The Markov chain Monte Carlo (MCMC) was run for 10 million generations, in four replicates. The default 25% of initial (burn-in) generations was discarded after inspection of the results in Tracer v1.6 (Rambaut et al. 2014). All runs converged at the same solution; hence the resulting log and tree files were combined. The effective sample size (ESS) of all parameters exceeded 300, ensuring that the results of the analysis were meaningful. BSP was created in Tracer using combined tree and log files.
Results
A 1249-bp long fragment of mitochondrial DNA, encoding part of the ND5 gene, was sequenced in six samples of C. glacialis (Table 1). The diversity indices showed overall high haplotype diversity (Hd), which was apparently associated with low nucleotide diversity (π) in the studied population of C. glacialis (Table 2). There were 75 haplotypes among 140 sequenced individuals (Hd = 0.892). Despite this appreciable number of haplotypes, the overall nucleotide diversity was very low, at the level of π = 0.004. There was no genetic differentiation between tested pairs of populations (population pairwise ΦST did not differ significantly from zero); hence there was no evidence of any population genetic structure among the compared samples (p > 0.05; Table 3). Various groupings were checked for possible higher-level structuring using AMOVA, but no significant fixation indices were recovered, regardless of the grouping tested (data not shown). The MSN of all haplotypes (Fig. 2) was relatively simple and well-resolved, with three closely related haplotypes surrounded by several minor-frequency variants. However, there was no visible trend in the distribution of haplotypes by geographic location (Fig. 2a) or year of sampling (Fig. 2b). Taking into account the lack of structuring, all subsequent analyses were run on a combined set of all 140 sequences obtained.
The minimum spanning networks (MSN) of ND5 haplotypes of Calanus glacialis from the fjords of Spitsbergen. Circle diameters are proportional to the number of individuals bearing each haplotype, and lines connecting circles are roughly proportional to the number of mutational steps connecting haplotypes. a Distribution of haplotypes between sampling locations; number of individuals is also shown here. b Distribution of haplotypes between sampling years; exact numbers of mutational steps are shown here
Tajima’s D test statistic was significantly negative (D = −2.51, p < 0.05), indicating an excess of rare variants and hinting at possible recent population expansion. To test this interpretation and to elucidate the demographic history of the studied population of C. glacialis, BSP analysis was performed (Fig. 3). The resulting plot indicates a strong increase in population size occurring at approximately the time sufficient to accumulate between 6×10−4 and 9×10−4 substitutions per site, with a relatively wide CI of 4×10−4 substitutions.
The demographic history of the Arctic population of Calanus glacialis based on a Bayesian skyline plot (BSP) reconstruction approach. The black line depicts median values, and the shaded region represents 95% highest posterior density. Both axes are scaled in uncalibrated, mutational units; if divided by the true substitution rate, they would produce time (x-axis) or the effective population size (y-axis)
Facing the complete lack of possible calibration points, the dating of this event can only be highly provisional. However, in order to fall within the postglacial limit, the expansion start (9×10−4 substitutions) would have to be inferred at no more than 2×104 years before present (the Last Glacial Maximum). Accordingly, to fit our data within the confidence limits, the substitution rate would have to be in the range of 3.5–5.5% per MY ([9×10−4− 2×10−4]/2×104 substitutions per site per year for the lower limit). Assuming a lower substitution rate would push the expansion event out of the interglacial.
Discussion
Our study revealed no evidence of genetic structure in Calanus glacialis among the fjords compared nor among different years of sampling, regardless of their classification as warm (2006) or cold (2003, 2004). The results are similar to those by Weydmann et al. (2016), who reported a lack of genetic structure in C. glacialis from seven locations distributed around the Arctic (Svalbard fjords, White Sea, and Amundsen Gulf), sampled in 2008 and 2009, in support of the hypothesis that large-scale effective dispersal and gene flow driven by ocean currents allows for the free exchange of planktonic copepods in the Arctic. Therefore, we believe the results would be similar even if we had sampled more sites around the Arctic. There is also numerous evidence of panmictic populations and/or high gene flow of planktonic copepods across extensive geographic ranges of the Northern Hemisphere, which has been reported for the Atlantic Calanus finmarchicus (Provan et al. 2009), Pacific Calanus sinicus (Huang et al. 2014), cosmopolitan Clausocalanus arcuicornis (Blanco-Bercial et al. 2011), and Arctic Pseudocalanus minutus (Aarbakke et al. 2014; Questel et al. 2016). At the same time, to our knowledge, there is only one study confirming the existence of two populations of C. glacialis, in the Arctic and the North Pacific (Bering Sea), although the latter was not reproductively established in the Arctic Ocean (Nelson et al. 2009).
Although there was no connection to locations or time in the distribution of C. glacialis haplotypes, one interesting feature of the observed topology was the existence of star-like elements in the MSN: single, dominant haplotypes connected by short branches with several low-frequency haplotypes (Fig. 2). Such structures usually indicate recent expansion of the clades represented by them (Network software documentation, www.fluxus-engineering.com). Recent population expansions are also known to leave certain traces in the observed diversity indices. The excess of rare polymorphism is expected in such situations, leading to significantly negative Tajima’s D test statistic (Tajima 1989), which was the case in our study. This excess can also be caused by selection acting on the studied marker; however, in the case of a mitochondrial marker it is usually assumed that demographic processes are responsible for this phenomenon (Grant 2015).
Multilocus data are known to be better for inferring demographic histories, particularly when combined with ancient DNA sampling (Grant 2015). Unfortunately, for various technical reasons, such data are currently unavailable for Calanus species. Active marker development is ongoing (Smolina et al. 2014; Weydmann et al. 2014), but is hampered by the atypical genome organization in Calanus. Also, using a single mitochondrial gene has some advantages: as a maternally inherited, haploid genome, it has a smaller effective population size and is more prone to bottleneck effects (Hartl and Clark 2007). Therefore, the expansion seen in our data was not necessarily preceded by a very strong bottleneck.
The rapid change recorded on the BSP plot (Fig. 3) confirms that the studied population had undergone an expansion. The observed high haplotype diversity is indicative of a large effective population size, typically expected for a planktonic marine invertebrate. The causes of the observed pattern can be attributed to the bottleneck experienced by C. glacialis during the Pleistocene glaciation and the following rapid expansion of this species after the Last Glacial Maximum, when the Arctic was exposed to warming, resulting in a transition from full glacial conditions to widespread interglacial conditions attained approximately 10,000 years ago. Such expansion is common for many Arctic species, which survived in a few refugia and very rapidly recolonized their current ranges after deglaciation; their current genetic diversity depends largely on the number of refugia and effective population sizes of the surviving populations (Hewitt 2000; Hewitt 2004). Marine species like C. glacialis are additionally affected by ocean currents that contribute to mixing processes between their populations.
The question remains whether this explanation is plausible and the estimated substitution rates are acceptable. The typically assumed general mitochondrial substitution rates are based on separating pairs of shrimp species by the emerging Isthmus of Panama (Knowlton et al. 1993; Knowlton and Weigt 1998) and a relatively short fragment of the conserved COX1 gene. It is assumed that other crustaceans, including Calanus species, accumulate mitochondrial substitutions at a similar pace, resulting in the 1.4–2.2% increase in overall divergence per million years (Papadopoulos et al. 2005). The substitution rate needed to attribute the observed expansion in C. glacialis to the last interglacial is 3.5% per million years, leading to the accumulation of divergence at the speed of at least 7% per million years, a value seemingly much higher but still in the same order of magnitude. At least two factors must be considered, each acting in favor of the increased substitution rate. First, the published data are dealing with the most conservative COX1 gene, while the ND5 sequence is most likely evolving much faster. Recent mitogenomic analysis of Metacrangonyctidae crustaceans (Pons et al. 2014) have shown that ND5 is among the fastest-evolving mitochondrial genes and accumulates substitutions about twice as fast as COX1.
Direct estimates of the substitution rate are rare, but Haag-Liautard et al. (2008) measured the mitochondrial mutation rate in Drosophila. The obtained value of 6.2 × 10−8 per site per fly generation would correspond to 3.1 × 10−8 per site per year for C. glacialis (assuming the 2-year generation time). That would fit our requirement reasonably well, further indicating that the assumed substitution rate is quite plausible.
Another important consideration is the apparent time dependency of molecular rate estimates (Ho et al. 2005), frequently leading to large errors in calibrating recent events by using rate estimates derived from phylogenetic species separations (Grant 2015). These effects are difficult to mea-sure, but they both act in the same direction: towards the increase in an apparent substitution rate and the placement of the expansion event within the current interglacial. In line with this view is the notion that, when dealing with contemporary sequence data, only the last major expansion event can be noted after even a moderate bottleneck (Grant 2015); the information about the past demographic events is lost. It is difficult to imagine that the Last Glacial Maximum had no effect on the demography of C. glacialis, and only such an assumption would allow us to interpret the observed expansion as an earlier event.
Why would the Arctic ice-associated species benefit from the interglacial conditions? One possible answer is that the thinning of sea ice affected the primary production regime, allowing for the earlier ice algal bloom and subsequent phytoplankton bloom after the ice melts, which are required for the early maturation, reproduction and growth of this key Arctic grazer (Niehoff et al. 2002; Søreide et al. 2010). Another possible explanation is that during the glacial, the sea level was lower and shelves were mostly covered by grounded ice, so there was no optimal habitat for C. glacialis, which is primarily a shelf and shelf break species, probably not as successful in the basins (Ji et al. 2012). On the other hand, interglacial conditions enhance thermohaline circulation in the subpolar North Atlantic (Sarnthein et al. 1994; McManus et al. 2002), thus providing better conditions for transporting zooplankton. In conclusion, we hypothesize that the recent interglaciation opened up the Arctic Ocean’s shelves, changed the circulation of ocean currents and accelerated the blooms, and the combination of these jointly enabled the postglacial expansion of this Arctic keystone copepod.
References
Aarbakke ONS, Bucklin A, Halsband C, Norrbin F (2014) Comparative phylogeography and demographic history of five sibling species of Pseudocalanus (Copepoda: Calanoida) in the North Atlantic Ocean. J Exp Mar Biol Ecol 461:479–488. doi:10.1016/j.jembe.2014.10.006
Baele G, Lemey P, Bedford T, Rambaut A, Suchard MA, Alekseyenko AV (2012) Improving the accuracy of Demographic and Molecular Clock Model comparison while accommodating phylogenetic uncertainty. Mol Biol Evol 29:2157–2167. doi:10.1093/molbev/mss084
Baker A (ed) (2000) Molecular Methods in Ecology. Blackwell Science
Bandelt HJ, Forster P, Röhl A (1999) Median-joining networks for inferring intraspecific phylogenies. Mol Biol Evol 16:37–48
Blachowiak-Samolyk K, Søreide JE, Kwaśniewski S, Sundfjord A, Hop H, Falk-Petersen S et al (2008) Hydrodynamic control of mesozooplankton abundance and biomass in northern Svalbard waters (79–81°N). Deep-Sea Res II 55:2210–2224. doi:10.1016/j.dsr2.2008.05.018
Blanco-Bercial L, Álvarez-Marqués F, Bucklin A (2011) Comparative phylogeography and connectivity of sibling species of the marine copepod Clausocalanus (Calanoida). J Exp Mar Biol Ecol 404:108–115
Bouckaert R, Heled J, Kühnert D, Vaughan T, Wu C-H, Xie D, Suchard MA, Rambaut A, Drummond AJ (2014) BEAST 2: A software platform for Bayesian evolutionary analysis. PLoS Comput Biol 10:e1003537. doi:10.1371/journal.pcbi.1003537
Brodskii KA, Vyshkvartseva NV, Kos MS, Markhatseva EL (1983) Copepods (Copepoda: Calanoida) of the seas of the USSR and adjacent waters. Vol. 1. Keys to the fauna of the USSR No 135. In Russian
Conover RJ (1988) Comparative life histories in the genera Calanus and Neocalanus in high latitudes of the northern hemisphere. Hydrobiologia 167(168):127–142. doi:10.1007/BF00026299
Cottier FR, Tverberg V, Inall ME, Svendsen H, Nilsen F, Griffiths C (2005) Water mass modification in an Arctic fjord through cross-shelf exchange: the seasonal hydrography of Kongsfjord, Svalbard. J Geophysical Res 110:C12005. doi:10.1029/2004JC002757
Excoffier L, Lischer HEL (2010) Arlequin suite ver 35: A new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Resour 10:564–567. doi:10.1111/j.1755-0998.2010.02847.x
Falk-Petersen S, Mayzaud P, Kattner G, Sargent JR (2009) Lipids and life strategy of Arctic Calanus. Mar Biol Res 5:18-39. doi:10.1080/17451000802512267
Fleminger A, Hulseman K (1977) Geographical range and taxonomic divergence in North Atlantic Calanus (C helgolandicus, C finmarchicus and C glacialis). Mar Biol 40:233–248. doi:10.1007/BF00390879
Grant WS (2015) Problems and cautions with sequence mismatch analysis and Bayesian skyline plots to infer historical demography. J Hered 106:333–346. doi:10.1093/jhered/esv020
Gray JS (1997) Marine biodiversity: patterns, threats and conservation needs. Biodivers Conserv 6:153–175. doi:10.1023/A:1018335901847
Haag-Liautard C, Coffey N, Houle D, Lynch M, Charlesworth B et al (2008) Direct Estimation of the Mitochondrial DNA Mutation Rate in Drosophila melanogaster. PLOS Biology 6(8):e204. doi:10.1371/journal.pbio.0060204
Hartl DL, Clark AG (2007) Principles of Population Genetics. Sinauer Associates, Inc
Hewitt GM (2000) The genetic legacy of the Quaternary ice ages. Nature 45:907–913. doi:10.1038/35016000
Hewitt GM (2004) Genetic consequences of climatic oscillations in the Quaternary. Philos Trans R Soc Lond B Biol Sci 359:183–195. doi:10.1098/rstb.2003.1388
Hirche HJ, Kwaśniewski S (1997) Distribution, reproduction and development of Calanus species in the Northeast Water in relation to environmental conditions. J Mar Syst 10:299–317. doi:10.1016/S0924-7963(96)00057-7
Ho SYW, Phillips MJ, Cooper A, Drummond AJ (2005) Time dependency of molecular rate estimates and systematic overestimation of recent divergence times. Mol Biol Evol 22:1561–1568. doi:10.1093/molbev/msi145
Huang Y, Liu G, Chen X (2014) Molecular phylogeography and population genetic structure of the planktonic copepod Calanus sinicus Brodsky in the coastal waters of China. Acta Oceanol Sin 33:74–84. doi:10.1007/s13131-014-0542-2
IPCC (2014) Climate Change 2014: Synthesis report. Contribution of Working Groups I, II and III to the Fifth Assessment Report of the Intergovernmental Panel on Climate Change [Core Writing Team, Pachauri RK, Meyer LA, editors]. IPCC Geneva. doi: 10013/epic.45156
Ji R, Ashjian CJ, Campbell RG et al (2012) Life history and biogeography of Calanus copepods in the Arctic Ocean: an individual-based modelling study. Prog Oceanogr 96:40–56. doi:10.1016/j.pocean.2011.10.001
Kenchington E, Heino M, Nielsen EE (2003) Managing marine genetic diversity: time for action? ICES J Mar Sci 60:1172–1176. doi:10.1016/S1054-3139(03)00136-X
Knowlton N, Weigt LA (1998) New dates and new rates for divergence across the Isthmus of Panama. Proc R Soc Lond B Biol Sci 265:2257–2263. doi:10.1098/rspb.1998.0568
Knowlton N, Weigt LA, Solorzano LA, Mills DK, Bermingham E (1993) Divergence in proteins, mitochondrial DNA, and reproductive compatibility across the Isthmus of Panama. Science 260:1629–1632
Kosobokova KN (1999) The reproductive cycle and life history of the Arctic copepod Calanus glacialis in the White Sea. Polar Biol 22:254–263. doi:10.1007/s003000050418
Kwaśniewski S, Hop H, Falk-Petersen S, Pedersen G (2003) Distribution of Calanus species in Kongsfjorden, a glacial fjord in Svalbard. J Plankton Res 25:1–20. doi:10.1093/plankt/25.1.1
Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H et al (2007) Clustal W and Clustal X version 20. Bioinformatics 23:2947–2948. doi:10.1093/bioinformatics/btm404
Lee RF, Hirota J (1973) Wax esters in tropical zooplankton and nekton and the geographical distribution of wax esters in marine copepods. Limnol Oceanogr 18:227–239. doi:10.4319/lo.1973.18.2.0227
Librado P, Rozas J (2009) DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25:1451–1452. doi:10.1093/bioinformatics/btp187
MacLellan DC (1967) The annual cycle of certain calanoid species in west Greenland. Can J Zool 45:101–115. doi:10.1139/z67-013
McManus J, Oppo D, Keigwin L, Cullen J, Bond G (2002) Thermohaline circulation and prolonged interglacial warmth in the North Atlantic. Quat Rev 58:17–21. doi:10.1006/qres.2002.2367
Minxiao W, Song S, Chaolun L, Xinm S (2011) Distinctive mitochondrial genome of Calanoid copepod Calanus sinicus with multiple large non-coding regions and reshuffled gene order: useful molecular markers for phylogenetic and population studies. BMC Genomics 27:12–73. doi:10.1186/1471-2164-12-73
Nelson RJ, Carmack EC, McLaughlin FA, Cooper GA (2009) Penetration of Pacific zooplankton into the western Arctic Ocean tracked with molecular population genetics. Mar Ecol Prog Ser 381:129–138. doi:10.3354/meps07940
Niehoff B, Madsen SD, Hansen BW, Nielsen TG (2002) Reproductive cycles of three dominant Calanus species in Disko Bay, West Greenland. Mar Biol 140:567–576. doi:10.1007/s00227-001-0731-3
Papadopoulos LN, Peijnenburg KTCA, Luttikhuizen PC (2005) Phylogeography of the calanoid copepods Calanus helgolandicus and C. euxinus suggests Pleistocene divergences between Atlantic, Mediterranean, and Black Sea populations. Mar Biol 147:1353–1365. doi:10.1007/s00227-005-0038-x
Parent GJ, Plourde S, Turgeon J (2012) Natural hybridization between Calanus finmarchicus and C. glacialis (Copepoda) in the Arctic and Northwest Atlantic. Limnol Oceanogr 57:1057–1066. doi:10.4319/lo.2012.57.4.1057
Pons J, Bauzà-Ribot MM, Jaume D, Juan C (2014) Next-generation sequencing, phylogenetic signal and comparative mitogenomic analyses in Metacrangonyctidae (Amphipoda: Crustacea). BMC Genomics 15:566. doi:10.1186/1471-2164-15-566
Provan J, Beatty GE, Keating SL, Maggs CA, Savidge G (2009) High dispersal potential has maintained long-term population stability in the North Atlantic copepod Calanus finmarchicus. Proc R Soc B 276:301–307. doi:10.1098/rspb.2008.1062
Provan J, Beatty GE, Maggs CA, Savidge G (2007) Expressed sequence tag-derived microsatellites for the cool-water marine copepod Calanus finmarchicus. Mol Ecol Notes 7:1369–1371. doi:10.1111/j.1471-8286.2007.01889.x
Questel JM, Blanco-Bercial L, Hopcroft R, Bucklin A (2016) Phylogeography and connectivity of the Pseudocalanus (Copepoda: Calanoida) species complex in the eastern North Pacific and the Pacific Arctic Region. J Plankton Res 38:610–623. doi:10.1093/plankt/fbw025
Rambaut A, Suchard MA, Xie D, Drummond AJ (2014) Tracer v16 Available from http://beast.bio.ed.ac.uk/Tracer
Ramos AA, Weydmann A, Cox CJ, Canário AV, Serrão EA, Pearson GA (2015) A transcriptome resource for the copepod Calanus glacialis across a range of culture temperatures. Mar Genomics 23:27-29. doi: 10.1016/j.margen.2015.03.014
Reed DH, Frankham R (2003) Correlation between fitness and genetic diversity. Conserv Biol 17:230–237. doi:10.1046/j.1523-1739.2003.01236.x
Sarnthein M, Winn K, Jung SJA, Duplessy JC, Labeyrie L, Erlenkeuser H, Ganssen G (1994) Changes in east Atlantic deep water circulation over the last 30,000 years: Eight time slice reconstructions. Paleoceanography 9:209–267. doi:10.1029/93PA03301
Smolina I, Kollias S, Poortvliet M, Nielsen TG, Lindeque P, Castellani C, Moller EF, Blanco-Bercial L, Hoarau G (2014) Genome- and transcriptome-assisted development of nuclear insertion/deletion markers for Calanus species (Copepoda: Calanoida) identification. Mol Ecol Resour 14:1072–1079. doi:10.1111/1755-0998.12241
Søreide JE, Leu E, Berge J, Graeve M, Falk-Petersen S (2010) Timing of blooms, algal food quality and Calanus glacialis reproduction and growth in a changing Arctic. Global Change Biol 16:3154–3163. doi:10.1111/j.1365-2486.2010.02175.x
Staden R (1996) The Staden Sequence Analysis Package. Mol Biotechnol 5:233–241. doi:10.1007/BF02900361
Swerpel S (1985) The Hornsund Fiord: Water Masses. Pol Polar Res 6:475–496
Tajima F (1989) Statistical method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics 123:585–595
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: Molecular Evolutionary Genetics Analysis version 6.0. Mol Biol Evol 30:2725–2729 10.1093/molbev/mst197
Walczowski W, Piechura J, Goszczko I, Wieczorek P (2012) Changes of the Atlantic Water properties: an important factor of the European Arctic marine climate. ICES J Mar Sci 69:864–869. doi:10.1093/icesjms/fss068
Walkusz W, Kwaśniewski S, Falk-Petersen S, Hop H, Tverberg V, Wieczorek P, Węsławski JM (2009) Seasonal and spatial changes in the zooplankton community of Kongsfjorden, Svalbard. Polar Res 28:254–281. doi:10.1111/j.1751-8369.2009.00107.x
Werle E, Schneider C, Renner M, Volker M, Fiehn W (1994) Convenient single-step, one tube purification of PCR products for direct sequencing. Nucleic Acids Res 22:4354–4355
Weydmann A, Coelho NC, Ramos AA, Serrão EA, Pearson GA (2014) Microsatellite markers for the Arctic copepod Calanus glacialis and cross-amplification with C finmarchicus. Conserv Genet Resour 6:1003–1005. doi:10.1007/s12686-014-0269-6
Weydmann A, Coelho NC, Serrão EA, Burzyński A, Pearson GA (2016) Pan-Arctic population of the keystone copepod Calanus glacialis. Polar Biol 39:2311–2318. doi:10.1007/s00300-016-1898-x
Weydmann A, Kwasniewski S (2008) Distribution of Calanus populations in a glaciated fjord in the Arctic (Hornsund, Spitsbergen) – an interplay between biological and physical factors. Polar Biol 31:1023–1035. doi:10.1007/s00300-008-0441-0
Weydmann A, Søreide JE, Kwasniewski S, Leu E, Falk-Petersen S, Berge J (2013) Ice-related seasonality in zooplankton community composition in a high Arctic fjord. J Plankton Res 35:831–842. doi:10.1093/plankt/fbt031
Acknowledgements
This research project was supported by grants no. 2011/03/B/NZ8/02876 and GAME from the National Science Centre, the Polish Ministry of Science and Higher Education Iuventus Plus grant no. IP2014 050573 and by CCMAR/Multi/04326/2013 from FCT- PT.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by R. R. Hopcroft
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Weydmann, A., Przyłucka, A., Lubośny, M. et al. Postglacial expansion of the Arctic keystone copepod Calanus glacialis . Mar Biodiv 48, 1027–1035 (2018). https://doi.org/10.1007/s12526-017-0774-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12526-017-0774-4