Abstract
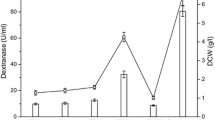
Dextranase from Chaetomium gracile is generally considered safe for use in the sugarcane industry. Herein, a truncated and codon-optimised α-dextranase gene from C. gracile was successfully cloned and expressed in Saccharomyces cerevisiae for the first time. The optimum conditions of fermentation was achieved when the maximum dextranase activity reached to 58.45 U/mL after 48 h in shake flasks. The optimal pH and temperature were 5.5 and 60 °C, respectively. The recombinant dextranase remained stable between pH 4 and 6 and temperature between 55 and 60 °C. The findings in the present study could facilitate large-scale production of food-grade recombinant dextranase for use in the sugar industry.





Similar content being viewed by others
References
Armando, M.R., M.A. Galvagno, C.A. Dogi, P. Cerrutti, A.M. Dalcero, and L.R. Cavaglieri. 2013. Statistical optimization of culture conditions for biomass production of probiotic gut-borne Saccharomyces cerevisiae strain able to reduce fumonisin B1. Journal of Applied Microbiology 114(5): 1338–1346. https://doi.org/10.1111/jam.12144.
Bertrand, E., G. Pierre, C. Delattre, C. Gardarin, N. Bridiau, T. Maugard, A. Strancar, and P. Michaud. 2014. Dextranase immobilization on epoxy CIM (R) disk for the production of isomaltooligosaccharides from dextran. Carbohydrate Polymers 111: 707–713. https://doi.org/10.1016/j.carbpol.2014.04.100.
Chen, Z.W., W. Xu, W.L. Zhang, T. Zhang, B. Jiang, and W.M. Mu. 2018. Characterization of a thermostable recombinant l-rhamnose isomerase from Caldicellulosiruptor obsidiansis OB47 and its application for the production of l-fructose and l-rhamnulose. Journal of the Science of Food and Agriculture 98(6): 2184–2193. https://doi.org/10.1002/jsfa.8703.
Chen, L., C. Yu, X.S. Zhou, and Y.X. Zhang. 2009. Rational introduction of disulfide bond to enhance optimal temperature of Lipomyces starkeyi alpha-dextranase expressed in Pichia pastoris. Journal of Microbiology and Biotechnology 19(12): 1506–1513. https://doi.org/10.4014/jmb.0902.0096.
Chen, L., X.S. Zhou, W.M. Fan, and Y.X. Zhang. 2008. Expression, purification and characterization of a recombinant Lipomyces starkey dextranase in Pichia pastoris. Protein Expression and Purification 58(1): 87–93. https://doi.org/10.1016/j.pep.2007.10.021.
Eggleston, Gillian. 2009. Application of dextranases in sugarcane factory: Overcoming practical problems. Sugar Tech 11(2): 135–141. https://doi.org/10.1007/s12355-009-0020-x.
Eggleston, G., and A. Monge. 2005. Optimization of sugarcane factory application of commercial dextranases. Process Biochemistry 40(5): 1881–1894. https://doi.org/10.1016/j.procbio.2004.06.025.
Fraga Vidal, Reinaldo, Aidin Martinez, Claire Moulis, Pierre Escalier, Sandrine Morel, Magali Remaud-Simeon, and Pierre Monsan. 2011. A novel dextransucrase is produced by Leuconostoc citreum strain B/110-1-2: An isolate used for the industrial production of dextran and dextran derivatives. Journal of Industrial Microbiology and Biotechnology 38(9): 1499–1506. https://doi.org/10.1007/s10295-010-0936-x.
Igarashi, T., H. Morisaki, and N. Goto. 2004. Molecular characterization of dextranase from Streptococcus rattus. Microbiology and Immunology 48(3): 155–162. https://doi.org/10.1111/j.1348-0421.2004.tb03501.x.
Kang, H.K., S.H. Kim, J.Y. Park, X.J. Jin, D.K. Oh, S.S. Kang, and K. Doman. 2005. Cloning and characterization of a dextranase gene from Lipomyces starkeyi and its expression in Saccharomyces cerevisiae. Yeast 22(15): 1239–1248. https://doi.org/10.1002/yea.1311.
Kang, H.K., J.Y. Park, J.S. Ahn, S.H. Kim, and D. Kim. 2009. Cloning of a gene encoding dextranase from Lipomyces starkeyi and its expression in Pichia pastoris. Journal of Microbiology and Biotechnology 19(2): 172–177. https://doi.org/10.4014/jmb.0802.100.
Khalikova, E., P. Susi, and T. Korpela. 2005. Microbial dextran-hydrolyzing enzymes: Fundamentals and applications. Microbiology and Molecular Biology Reviews 69(2): 306–325. https://doi.org/10.1128/Jmrb.69.2.306-325.2005.
Kwak, Suryang, and Yong-Su Jin. 2017. Production of fuels and chemicals from xylose by engineered Saccharomyces cerevisiae: A review and perspective. Microbial Cell Factories 16: 82. https://doi.org/10.1186/s12934-017-0694-9.
Li, X., S.H. Millson, R.D. Coker, and I.H. Evans. 2006. Cloning and expression of Penicillium minioluteum dextranase in Saccharomyces cerevisiae and its exploitation as a reporter in the detection of mycotoxins. Biotechnology Letters 28(23): 1955–1964. https://doi.org/10.1007/s10529-006-9183-7.
Li, Aimin, Zengshan Liu, LuYu. Qianxue Li, Dacheng Wang, and Xuming Deng. 2007. Construction and characterization of bidirectional expression vectors in Saccharomyces cerevisiae. FEMS Yeast Research 8(1): 6–9. https://doi.org/10.1111/j.1567-1364.2007.00335.x.
Li, K., H.Q. Lu, F.X. Hang, S.B. Li, and J.D. Liu. 2017. Improved dextranase production by Chaetomium gracile through optimization of carbon source and fermentation parameters. Sugar Tech 19(4): 432–437. https://doi.org/10.1007/s12355-016-0476-4.
Lian, Jiazhang, Shekhar Mishra, and Huimin Zhao. 2018. Recent advances in metabolic engineering of Saccharomyces cerevisiae: New tools and their applications. Metabolic Engineering. https://doi.org/10.1016/j.ymben.2018.04.011.
Liu, Z.H., K.E.J. Tyo, J.L. Martinez, D. Petranovic, and J. Nielsen. 2012. Different expression systems for production of recombinant proteins in Saccharomyces cerevisiae. Biotechnology and Bioengineering 109(5): 1259–1268. https://doi.org/10.1002/bit.24409.
Liu, J.D., W.P. Zhang, G.C. Du, J. Chen, and J.W. Zhou. 2013. Overproduction of geraniol by enhanced precursor supply in Saccharomyces cerevisiae. Journal of Biotechnology 168(4): 446–451. https://doi.org/10.1016/j.jbiotec.2013.10.017.
Ljungdahl, Per O., and Bertrand Daignan-Fornier. 2012. Regulation of amino acid, nucleotide, and phosphate metabolism in Saccharomyces cerevisiae. Genetics 190(3): 885–929. https://doi.org/10.1534/genetics.111.133306.
Luo, G., J.H. Tian, H.Q. Huang, and L. An. 2018. Improving heterologous expression of porcine follicle-stimulating hormone in Pichia pastoris by integrating molecular strategies and culture condition optimization. Applied Microbiology and Biotechnology 102(20): 8867–8882. https://doi.org/10.1007/s00253-018-9260-6.
Macauley-Patrick, S., M.L. Fazenda, B. McNeil, and L.M. Harvey. 2005. Heterologous protein production using the Pichia pastoris expression system. Yeast 22(4): 249–270. https://doi.org/10.1002/yea.1208.
Mao, H.Y., M.H. Qiu, X.F. Chen, H. Verweij, and Y.Q. Fan. 2018. Fabrication and in situ fouling mitigation of a supported carbon nanotube/gamma-alumina ultrafiltration membrane. Journal of Membrane Science 550: 26–35. https://doi.org/10.1016/j.memsci.2017.12.050.
Martinez-Moreno, R., P. Morales, R. Gonzalez, A. Mas, and G. Beltran. 2012. Biomass production and alcoholic fermentation performance of Saccharomyces cerevisiae as a function of nitrogen source. FEMS Yeast Research 12(4): 477–485. https://doi.org/10.1111/j.1567-1364.2012.00802.x.
Purushe, Shweta, Divya Prakash, Neelu N. Nawani, Prashant Dhakephalkar, and Balasaheb Kapadnis. 2012. Biocatalytic potential of an alkalophilic and thermophilic dextranase as a remedial measure for dextran removal during sugar manufacture. Bioresource Technology 115: 2–7. https://doi.org/10.1016/j.biortech.2012.01.002.
Rerngsamran, P., P. Temjitpukdee, N. Assavasirijinda, S. Chareonpornwattana, and S. Thaniyavarn. 2014. Cloning, characterization, and heterologous expression of a dextranase gene from Penicillium pinophilum SMCU3-14. Scienceasia 40(6): 405–413. https://doi.org/10.2306/scienceasia1513-1874.2014.40.405.
Roca, H., B. Garcia, E. Rodriguez, D. Mateu, L. Coroas, J. Cremata, R. Garcia, T. Pons, and J. Delgado. 1996. Cloning of the Penicillium minioluteum gene encoding dextranase and its expression in Pichia pastoris. Yeast 12(12): 1187–1200. https://doi.org/10.1002/(Sici)1097-0061(19960930)12:12%3c1187:Aid-Yea986%3e3.0.Co;2-U.
Sikorski, Robert S., and Philip Hieter. 1989. A system of shuttle vectors and yeast host strains designed for efficient manipulation of DNA in Saccharomyces cerevisiae. Genetics 122: 19–27.
Virgen-Ortíz, J.J., V. Ibarra-Junquera, P. Escalante-Minakata, J. de Ornelas-Paz, J.A. Osuna-Castro, and A. González-Potes. 2015. Kinetics and thermodynamic of the purified dextranase from Chaetomium erraticum. Journal of Molecular Catalysis B: Enzymatic 122: 80–86. https://doi.org/10.1016/j.molcatb.2015.08.020.
Zhang, Yu-Qi, Ruo-Han Li, Hong-Bin Zhang, Wu Min, and Hu Xue-Qin. 2017. Purification, characterization, and application of a thermostable dextranase from Talaromyces pinophilus. Journal of Industrial Microbiology and Biotechnology 44(2): 317–327. https://doi.org/10.1007/s10295-016-1886-8.
Zhang, Z.D., J.D. Liu, S.Y. Ma, H.Q. Lu, F.X. Hang, P. Huang, and K. Li. 2018. Enhancement of catalytic performance of alpha-dextranase from Chaetomium gracile through optimization and suitable shear force. Sugar Tech 20(1): 78–87. https://doi.org/10.1007/s12355-017-0540-8.
Acknowledgements
This work was supported by the National Natural Science Foundation of China (31460026), the National Natural Science Foundation of Guangxi Province (2018GXNSFAA050126), the Bosch Young Teachers Innovation Training Project (BRP180215) and the Fundamental Research Fund of Guangxi University (XJZ140293).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical Statement
This article does not contain any studies with human participants or animals performed by any of the authors.
Informed Consent
Informed consent was obtained from all individual participants included in the study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Liu, J., Sun, Q., Yin, H. et al. Optimal Fermentation of Saccharomyces cerevisiae Expressing a Dextranase from Chaetomium gracile. Sugar Tech 22, 171–178 (2020). https://doi.org/10.1007/s12355-019-00746-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12355-019-00746-5