Abstract
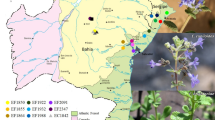
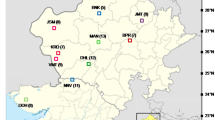
Three molecular marker systems, Random Amplified Polymorphic DNA (RAPD), Inter-Simple Sequence Repeats (ISSR) and Sequence-Related Amplified Polymorphism (SRAP) were employed to investigate the genetic structure and diversity among the 14 natural populations of Butea monosperma collected from different geographical regions of India. Detected by 17 RAPD, 15 ISSR and 11 SRAP primer combinations, the proportions of polymorphic bands were 84.2 %, 77.2 % and 91.9 %, respectively, and the mean Nei’s genetic distances among the populations were 0.13, 0.10 and 0.13, respectively. Partitioning of genetic variability by Analysis of molecular variance (AMOVA) revealed that the high genetic diversity was distributed within the populations. AMOVA also revealed that the coefficient of gene differentiation among populations based on FST was very high irrespective of markers used. The overall gene flow among populations (Nm) was very low. Cophenetic correlation coefficients of Nei’s distance values and clustering pattern by Mental test were statistically significant for all three marker systems used but poor fit for ISSR data than for RAPD, SRAP and combined data set of all three markers. For all markers, a high similarity in dendrogram topologies was obtained, although some differences were observed with ISSR. The dendrogram obtained by RAPD, SRAP and combined data set of all three markers reflect relationship of most of the populations according to their geographic distribution.



Similar content being viewed by others
References
Andrianoelina O, Rakotondraoelina H, Ramamonjisoa L, Maley J, Danthu P, Bouvet JM (2006) Genetic diversity of Dalbergia monticola (Fabaceae) an endangered tree species in the fragmented oriental forest of Madagascar. Biodivers Conserv 15:1109–1128
Aparajita S, Senapati SK, Rout GR (2008) Identification and genetic relationships among nine Albizzia species based on morphological and molecular markers. Plant Biosyst 142:30–39
Awasthi AK, Nagaraja GM, Naik GV, Kanginakudru S, Thangavelu K, Nagaraju J (2004) Genetic diversity and relationships in mulberry (genus Morus) as revealed by RAPD and ISSR marker assays. BMC Genet 5:1–9
Budak HRC, Shearman I, Parmaksiz RE, Gaussoin TP, Riosdan DI (2004) Molecular characterization of Buffalograss germplasm using sequence-related amplified polymorphism markers. Theor Appl Genet 108:328–334
Burli DA, Khade AB (2007) A comprehensive review on Butea monosperma (Lam.) Kuntze. Pharmacogn Rev 2:333–337
Casiva PV, Saidman BO, Vilardi JC, Cialdella AM (2002) First comparative phenetic studies of Argentinean species of Acacia (Fabaceae), using morphometric, isozymal, and RAPD approaches. Am J Bot 89:843–853
EspÓsito MA, Martin EA, Cravero VP, Cointry E (2007) Characterization of pea accessions by SRAP’s markers. Sci Hortic-Amsterdam 113:329–335
Excoffier L, Smouse PE, Quattro JM (1992) Analysis of molecular variance inferred from metric distances among DNA haplotypes: Application to human mitochondrial DNA data. Genetics 131:479–491
Ferriol M, Picó B, Fernández de Córdova P, Nuez F (2004) Molecular diversity of a germplasm collection of squash (cucurbita moschata) determined by SRAP and AFLP markers. Crop Sci 44:653–664
Hamrick JL, Godt MJW (1989) Allozyme diversity in plant species. In: Brown AHD, Clegg MT, Kahler AL, Weir BS (eds) Plant population genetics, breeding, and genetic resources. Sinauer, Massachusetts, pp 43–63
Hamrick JL, Godt MJW, Murawski DA, Loveless MD (1991) Correlations between species traits and allozyme diversity: implications for conservation biology. In: Falk D, Holsinger K (eds) Genetics and conservation of rare plants. Oxford Press, London, pp 75–86
Heider B, Andersson MS, Schultze-Kraft R (2007) RAPD variation among North Vietnamese Flemingia macrophylla (Willd.) Kuntze ex Merr. Accessions. Biodivers Conserv 16:1617–1631
Juarez-Munoz J, Carrillo-Castañeda G, Arreguín R, Rubluo A (2002) Inter and intragenetic variation of four wild populations of Prosopis using RAPD-PCR fingerprints. Biodivers Conserv 11:921–930
Juchum FS, Leal JB, Santos LM, Almeida MP, Ahnert D, Corrêa RX (2007) Evaluation of genetic diversity in a natural rosewood population (Dalbergia nigra Vell. Allemão ex Benth.) using RAPD markers. Genet Mol Res 6:543–553
Kimura M, Weiss GH (1964) The stepping-stone model of population structure and the decrease of genetic correlation with distance. Genetics 49:561–576
Lacerda DR, Acedo MDP, Lemos Filho JP, Lovato MB (2001) Genetic diversity and structure of natural populations of Plathymeni reticulate (Mimosoideae), a tropical tree from the Brazilian Cerrado. Mol Ecol 10:1143–1152
Li G, Quiros CF (2001) Sequence-related amplified polymorphism (SRAP), a new marker system based on a simple PCR reaction: its application to mapping and gene tagging in Brassica. Theor Appl Genet 103:455–461
Lloyd DG (1992) Self- and cross-fertilization in plants. II. The selection of self-fertilization. Int J Plant Sci 153:370–380
Mantel N (1967) The detection of disease clustering and a generalized regression approach. Cancer Res 27:209–220
Mazumder PM, Das MK, Das S, Das S (2011) Butea monosperma (Lam) Kuntze-A comprehensive review. Int J Pha Sci Nanotech 4(2):1390–1393
Medri C, Ruas PM, Higa AR, Murakami M, De Fátima RC (2003) Effects of forest management on the genetic diversity in a population of Araucaria angustifolia (bert.) O. Kuntze. Silvae Genet 52:202–205
Mitra AP (1988) The wealth of India: a dictionary of Indian raw materials & industrial products: raw materials, vol. 2: B. Publications & Information Directorate (CSIR), New Delhi
Nei M, Li WH (1979) Mathematical model for studying genetic variation in terms of restriction endonucleases. Proc Natl Acad Sci USA 76:5269–5273
Pal P, Bose S (2011) Phytopharmacological and Phytochemical Reviews of Butea monosperma. Int J Pha Biomed Sci 2(3):1374–1388
Pither R, Shore JS, Kellman M (2003) Genetic diversity of the tropical tree Terminalia amazonia (Combretaceae) in naturally fragmented populations. Heredity 91:307–313
Qiu YL, Parks CR (1994) Disparity of allozyme variation levels in three Mangnolia (Mangnoliaceae) species from the southeastern United States. Am J Bot 81:1300–1308
Rohlf FJ (1992) NTSYS-pc. Numerical taxonomy and multivariate analysis system, version 1.7. Exeter Software, New York
Saghai-Maroof MA, Soliman KM, Jorgensen RA, Allard RW (1984) Ribosomal DNA spacer-length polymorphisms in barley: Mendelian inheritance, chromosomal location and population dynamics. Proc Natl Acad Sci USA 81:8014–8018
Schierenbeck KA, Skupski M, Lieberman D, Lieberman M (1997) Population structure and genetic diversity in four tropical tree species in Costa Rica. Mol Ecol 6:137–144
Schnabel A, Hamrick JL (1990) Nonrandom associations between sex and 6-phosphogluconate dehydrogenase isozyme genotypes in Gleditsia triacanthos L. J Hered 81:230–233
Schneider S, Roessli D, Excoffier L (2000) ARLEQUIN, version 2.0: a software for population genetics data analysis. Genetic and Biometry Laboratory, Department of Anthropology, University of Geneva, Switzerland
Semagn S, Bjornstad A, Ndjiondjop MN (2006) An overview of molecular markers methods for plants. Afr J Biotechnol 5(25):2540–2568
Sharma KK, Ramani R (1999) An update on synoptic catalogue of lac insects (Homoptera: Tachardidae). J Bombay Nat Hist Soc 96:438–443
Sneath PHA, Sokal RR (1973) Numerical taxonomy. W.H. Freeman and Company, San Francisco
Song C, Bao M (2006) Genetic diversity of RAPD marker for natural Davidia involucrata populations. Front For China 1:95–99
Storfar A (1996) Quantitative genetics: a promising approach for the assessment genetic variation in endangered species. Trends Ecol Evol 11:343–348
Tandon R, Shivanna KR, Mohan Ram HY (2003) Reproductive biology of Butea monosperma (Fabaceae). Ann Bot 92:715–723
Vaishali, Khan S, Sharma V (2008) RAPD based assessment of genetic diversity of Butea monosperma from different agro-ecological regions of India. Indian J Biotechnol 7:320–327
Wang LL, Ping ZL, Qin GY, Xia WM, Ming CL, Lan YJ, Yan W, Min YF, Zhi WL (2008) DNA fingerprinting and genetic diversity analysis of late-bolting radish cultivars with RAPD, ISSR and SRAP markers. Sci Hortic-Amsterdam 116:240–247
Wright S (1943) Isolation by distance. Genetics 28:114–138
Wright S (1951) The genetical structure of populations. Ann Eug 15:323–354
Xue X, Wang Y, Korpelainen H, Li C (2007) Genetic diversity of Picea asperata populations based on RAPDs. Plant Biol 9:101–108
Acknowledgements
Authors are highly thankful and gratefully acknowledge Dr. KV Bhat, Principal Scientist, National Bureau of Plant Genetic Resources (NBPGR) New Delhi for helping in data analysis and to Department of Biotechnology, New Delhi (India) for financial support to carry out the present study.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Vashishtha, A., Jehan, T. & Lakhanpaul, S. Genetic diversity and population structure of Butea monosperma (Lam.) Taub.- a potential medicinal legume tree. Physiol Mol Biol Plants 19, 389–397 (2013). https://doi.org/10.1007/s12298-013-0170-x
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12298-013-0170-x