Abstract
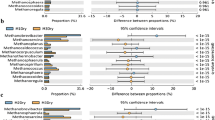
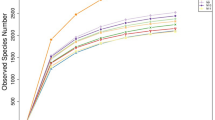
Molecular diversity of rumen archaeal populations from bovine rumen fluid incubated with or without condensed tannins was investigated using 16S rRNA gene libraries. The predominant order of rumen archaea in the 16S rRNA gene libraries of the control and condensed tannins treatment was found to belong to a novel group of rumen archaea that is distantly related to the order Thermoplasmatales, with 59.5% (15 phylotypes) and 81.43% (21 phylotypes) of the total clones from the control and treatment clone libraries, respectively. The 16S rRNA gene library of the control was found to have higher proportions of methanogens from the orders Methanomicrobiales (32%) and Methanobacteriales (8.5%) as compared to those found in the condensed tannins treatment clone library in both orders (16.88% and 1.68% respectively). The phylotype distributed in the order Methanosarcinales was only found in the control clone library. The study indicated that condensed tannins could alter the diversity of bovine rumen methanogens.
Similar content being viewed by others
References
Altschul, S.F., T.L. Madden, A.A. Schaffer, J. Zhang, Z. Zhang, W. Miller, and D.J. Lipman. 1997. Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402.
Amann, R.J., W. Ludwig, and K. Schleifer. 1995. Phylogenetic identification and in situ detection of individual microbial cells without cultivation. Microbiol. Mol. Biol. Rev. 59, 143–169.
Ashelford, K.E., N.A. Chuzhanova, J.C. Fry, A.J. Jones, and A.J Weightman. 2005. At least 1 in 20 16S rRNA sequence records currently held in public repositories is estimated to contain substantial anomalies. Appl. Environ. Microbiol. 71, 7724–7736.
Bapteste, E., C. Brochier, and Y. Boucher. 2005. Higher-level classification of the Archaea: evolution of methanogenesis and methanogens. Archaea 1, 353–363.
Benson, D.A., I. Karsch-Mizrachi, D.J. Lipman, J. Ostell, and D.L. Wheeler. 2008. GenBank. Nucleic Acids Res. 36, D25–D30.
Chaudhary, P.P. and S.K. Sirohi. 2009. Dominance of Methanomicrobium phylotype in methanogen population present in Murrah buffaloes (Bubalus bubalis). Lett. Appl. Microbiol. 49, 274–277.
Denman, S.E., N.W. Tomkins, and C.S. McSweeney. 2007. Quantitation and diversity analysis of ruminal methanogenic populations in response to the antimethanogenic compound bromochloromethane. FEMS Microbiol. Ecol. 62, 313–322.
FAO 2003. World Agriculture: Towards 2015/2030. An FAO Perspective. Rome.
Felsenstein, J. 1985. Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39, 783–791.
Hall, T.A. 1999. BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp. Ser. 41, 95–98.
Hess, H.D., M. Kreuzer, T.E. Díaz, C.E. Lascano, J.E. Carulla, C.R. Soliva, and A. Machmüller. 2003. Saponin rich tropical fruits affect fermentation and methanogenesis in faunated and defaunated rumen fluid. Anim. Feed Sci. Technol. 109, 79–94.
Hook, S.E., K.S. Northwood, A.D.G. Wright, and B.W. McBride. 2009. Long-term monensin supplementation does not significantly affect the quantity or diversity of methanogens in the rumen of the lactating dairy cow. Appl. Environ. Microbiol. 75, 374–380.
Irbis, C. and K. Ushida. 2004. Detection of methanogens and proteobacteria from a single cell of rumen ciliate protozoa. J. Gen. Appl. Microbiol. 50, 203–212.
IPCC. 2007. Summary for policymakers. In B. Metz, P.R. Bosch, R. Dave, and L.A. Meyer (eds.), Climate Change: Mitigation, Contribution of Working Group III to the Fourth Assessment Report of the Intergovernmental Panel on Climate Change.
Kimura, M. 1980. A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J. Mol. Evol. 16, 111–120.
Lin, C., L. Raskin, and D.A. Stahl. 1997. Microbial community structure in gastrointestinal tracts of domestic animals: comparative analyses using rRNA-targeted oligonucleotide probes. FEMS Microbiol. Ecol. 22, 281–294.
Makkar, H.P.S., M. Blümmel, and K. Becker. 1995. Formation of complexes between polyvinyl pyrrolidones or polyethylene glycols and tannins, and their implication in gas production and true digestibility in in vitro techniques. Br. J. Nutr. 73, 897–913.
Menke, K.H. and H. Steingass. 1988. Estimation of the energetic feed value obtained by chemical analysis and in vitro gas production using rumen fluid. Anim. Res. Dev. 28, 7–55.
Pei, C.X., S.Y. Mao, Y.F. Cheng, and W.Y. Zhu. 2009. Diversity, abundance and novel 16S rRNA gene sequences of methanogens in rumen liquid, solid and epithelium fractions of Jinnan cattle. Animal 4, 20–29.
Regensbogenova, M., P. Pristas, P. Javorsky, S.Y. Moon-Van Der Staay, G.W.M. Van Der Staay, J.H.P. Hackstein, C.J. Newbold, and N.R. McEwan. 2004. Assessment of ciliates in the sheep rumen by DGGE. Lett. Appl. Microbiol. 39, 144–147.
Ruepp, A., W. Graml, M.L. Santos-Martinez, K.K. Koretke, C. Volker, H.W. Mewes, D. Frishman, S. Stocker, A.N. Lupas, and W. Baumeister. 2000. The genome sequence of the thermoacidophilic scavenger Thermoplasma acidophilum. Nature 407, 508–513.
Saitou, N. and M. Nei. 1987. The neighbor-joining method: A new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 4, 406–425.
Shannon, C.E. and W. Weaver. 1949. The mathematical theory of communication. University of Illinois Press, Urbana, USA.
Sharp, R., C.J. Ziemer, M.D. Stern, and D.A. Stahl. 1998. Taxon-specific associations between protozoal and methanogen populations in the rumen and a model rumen system. FEMS Microbiol. Ecol. 26, 71–78.
Shin, E.C., B.R. Choi, W.J. Lim, S.Y. Hong, C.L. An, K.M. Cho, Y.K. Kim, and H.D. Yun. 2004. Phylogenetic analysis of archaea in three fractions of cow rumen based on the 16S rDNA sequence. Anaerobe 10, 313–319.
Shinzato, N., T. Matsumoto, I. Yamaoka, T. Oshima, and A. Yamagishi. 1999. Phylogenetic diversity of symbiotic methanogens living in the hindgut of the lower termite Reticulitermes speratus analyzed by PCR and in situ hybridization. Appl. Environ. Microbiol. 65, 837–840.
Skillman, L.C., P.N. Evans, G.E. Naylor, B. Morvan, G.N. Jarvis, and K.N. Joblin. 2004. 16S ribosomal DNA-directed PCR primers for ruminal methanogens and identification of methanogens colonising young lambs. Anaerobe 10, 277–285.
Tajima, K., T. Nagamine, H. Matsui, M. Nakamura, and R.I. Aminov. 2001. Phylogenetic analysis of archaeal 16S rRNA libraries from the rumen suggests the existence of a novel group of archaea not associated with known methanogens. FEMS Microbiol. Lett. 200, 67–72.
Tamura, K., J. Dudley, M. Nei, and S. Kumar. 2007. MEGA4: molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol. Biol. Evol. 24, 1596–1599.
Tavendale, M.H., L.P. Meagher, D. Pacheco, N. Walker, G.T. Attwood, and S. Sivakumaran. 2005. Methane production from in vitro rumen incubations with Lotus pedunculatus and Medicago sativa, and effects of extractable condensed tannin fractions on methanogenesis. Anim. Feed Sci. Technol. 123, 403–419.
Terrill, T.H., A.M. Rowan, G.B. Douglas, and T.N. Barry. 1992. Determination of extractable and bound condensed tannin concentrations in forage plants, protein concentrate meals and cereal grains. J. Sci. Food Agric. 58, 321–329.
Terrill, T.H., W.R. Windham, J.J. Evans, and C.S. Hoveland. 1990. Condensed tannin concentration in Sericea lespedeza as influenced by preservation method. Crop Sci. 30, 219–224.
Thompson, J.D., D.G. Higgins, and T.J. Gibson. 1994. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 22, 4673–4680.
US-EPA. 2006. Washington, D.C.: United States Environmental Protection Agency EPA 430-R-06-003.
Waghorn, G.C., M.H. Tavendale, and D.R. Woodfield. 2002. Methanogenesis from forages fed to sheep. Proc. N.Z. Grasslands Assoc. 64, 167–171.
Whitford, M.F., R.M. Teather, and R.J. Forster. 2001. Phylogenetic analysis of methanogens from the bovine rumen. BMC Microbiol. 1, 5.
Woodward, S.I., G.C. Waghorn, M.J. Ulyatt, and K.R. Lassey. 2001. Early indications that methane emissions from ruminants. Proc. N.Z. Soc. Anim. Prod. 61, 23–26.
Wright, A.D.G., C.H. Auckland, and D.H. Lynn. 2007. Molecular diversity of methanogens in feedlot cattle from Ontario and Prince Edward Island, Canada. Appl. Environ. Microbiol. 73, 4206–4210.
Wright, A.D.G., X. Ma, and N.E. Obispo. 2008. Methanobrevibacter phylotypes are the dominant methanogens in sheep from Venezuela. Microb. Ecol. 56, 390–394.
Wright, A.D.G. and C.L. Pimm. 2003. Improved strategy for presumptive identification of methanogens using 16S riboprinting. J. Microbiol. Methods 55, 337–349.
Wright, A.D.G., A.F. Toovey, and C.L. Pimm. 2006. Molecular identification of methanogenic archaea from sheep in Queensland, Australia reveal more uncultured novel archaea. Anaerobe 12, 134–139.
Wright, A.D.G., A.J. Williams, B. Winder, C. Christophersen, S. Rodgers, and K. Smith. 2004. Molecular diversity of rumen methanogens from sheep in Western Australia. Appl. Environ. Microbiol. 70, 1263–1270.
Yanagita, K., Y. Kamagata, M. Kawaharasaki, T. Suzuki, Y. Nakamura, and H. Minato. 2000. Phylogenetic analysis of methanogens in sheep rumen ecosystem and detection of Methanomicrobium mobile by fluorescence in situ hybridization. Biosci. Biotechnol. Biochem. 64, 1737–1742.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Tan, H.Y., Sieo, C.C., Lee, C.M. et al. Diversity of bovine rumen methanogens In vitro in the presence of condensed tannins, as determined by sequence analysis of 16S rRNA gene library. J Microbiol. 49, 492–498 (2011). https://doi.org/10.1007/s12275-011-0319-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12275-011-0319-7