Abstract
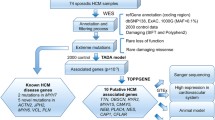
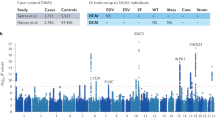
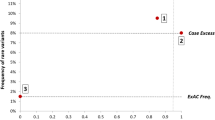
Despite the significant progress that has been made in identifying disease-associated mutations, the utility of the hypertrophic cardiomyopathy (HCM) genetic test is limited by a lack of understanding of the background genetic variation inherent to these sarcomeric genes in seemingly healthy subjects. This study represents the first comprehensive analysis of genetic variation in 427 ostensibly healthy individuals for the HCM genetic test using the “gold standard” Sanger sequencing method validating the background rate identified in the publically available exomes. While mutations are clearly overrepresented in disease, a background rate as high as ∼5 % among healthy individuals prevents diagnostic certainty. To this end, we have identified a number of estimated predictive value-based associations including gene-specific, topology, and conservation methods generating an algorithm aiding in the probabilistic interpretation of an HCM genetic test.





Similar content being viewed by others
Abbreviations
- 1kG:
-
1000 Genomes Sequencing Project
- ACTC:
-
Actin
- ARVC:
-
Arrhythmogenic right ventricular cardiomyopathy
- DHPLC:
-
Denaturing high-performance liquid chromatography heteroduplex analysis
- EPV:
-
Estimated predictive value
- ESP:
-
National Heart Lung and Blood Institute Exome Sequencing Project
- HCM:
-
Hypertrophic cardiomyopathy
- LQTS:
-
Long QT syndrome
- MYH7:
-
Beta myosin heavy chain
- MYL2:
-
Regulatory myosin light chain
- MYL3:
-
Essential myosin light chain
- MYPBC3:
-
Cardiac myosin binding protein C
- PCR:
-
Polymerase chain reaction
- SCD:
-
Sudden cardiac death
- TNNC1:
-
Cardiac troponin C
- TNNI3:
-
Cardiac troponin I
- TNNT2:
-
Cardiac troponin T
- TPM1:
-
Alpha-tropomyosin
- rNSV:
-
Rare non-synonymous variant
- VUS:
-
Variant of undetermined/uncertain significance
References
Maron, B. J., Epstein, S. E., & Roberts, W. C. (1986). Causes of sudden death in competitive athletes. JACC, 7(1), 204–214.
Maron, B. J., Doerer, J. J., Haas, T. S., Tierney, D. M., & Mueller, F. O. (2009). Sudden deaths in young competitive athletes: analysis of 1866 deaths in the United States, 1980–2006. Circulation, 119(8), 1085–1092. doi:10.1161/CIRCULATIONAHA.108.804617.
Maron, B. J. (2002). Hypertrophic cardiomyopathy: a systematic review. JAMA, 287(10), 1308–1320.
Bos, J. M., Ommen, S. R., & Ackerman, M. J. (2007). Genetics of hypertrophic cardiomyopathy: one, two, or more diseases? Current Opinion in Cardiology, 22(3), 193–199. doi:10.1097/HCO.0b013e3280e1cc7f.
Landstrom, A. P., & Ackerman, M. J. (2010). Mutation type is not clinically useful in predicting prognosis in hypertrophic cardiomyopathy. Circulation, 122(23), 2441–2449. doi:10.1161/CIRCULATIONAHA.110.954446. discussion 2450.
Ackerman, M. J., Priori, S. G., Willems, S., Berul, C., Brugada, R., Calkins, H., et al. (2011). HRS/EHRA expert consensus statement on the state of genetic testing for the channelopathies and cardiomyopathies: this document was developed as a partnership between the Heart Rhythm Society (HRS) and the European Heart Rhythm Association (EHRA). Europace, 13(8), 1077–1109. doi:10.1093/europace/eur245.
Van Driest, S. L., Ommen, S. R., Tajik, A. J., Gersh, B. J., & Ackerman, M. J. (2005). Yield of genetic testing in hypertrophic cardiomyopathy. Mayo Clinic Proceedings, 80(6), 739–744. doi:10.1016/S0025-6196(11)61527-9.
Landstrom, A. P., & Ackerman, M. J. (2009). GWAS or Gee Whiz, PSAS or Pshaw: elucidating the biologic and clinical significance of genetic variation in cardiovascular disease. Heart Rhythm, 6(12), 1751–1753. doi:10.1016/j.hrthm.2009.09.016.
Andreasen, C., Nielsen, J. B., Refsgaard, L., Holst, A. G., Christensen, A. H., Andreasen, L., et al. (2013). New population-based exome data are questioning the pathogenicity of previously cardiomyopathy-associated genetic variants. European Journal of Human Genetics. doi:10.1038/ejhg.2012.283.
Golbus, J. R., Puckelwartz, M. J., Fahrenbach, J. P., Dellefave-Castillo, L. M., Wolfgeher, D., & McNally, E. M. (2012). Population-based variation in cardiomyopathy genes. Circulation. Cardiovascular Genetics, 5(4), 391–399. doi:10.1161/CIRCGENETICS.112.962928.
Pan, S., Caleshu, C. A., Dunn, K. E., Foti, M. J., Moran, M. K., Soyinka, O., et al. (2012). Cardiac structural and sarcomere genes associated with cardiomyopathy exhibit marked intolerance of genetic variation. Circulation. Cardiovascular Genetics, 5(6), 602–610. doi:10.1161/CIRCGENETICS.112.963421.
Lopes, L. R., Zekavati, A., Syrris, P., Hubank, M., Giambartolomei, C., Dalageorgou, C., et al. (2013). Genetic complexity in hypertrophic cardiomyopathy revealed by high-throughput sequencing. Journal of Medical Genetics, 50(4), 228–239. doi:10.1136/jmedgenet-2012-101270.
Norton, N., Robertson, P. D., Rieder, M. J., Zuchner, S., Rampersaud, E., Martin, E., et al. (2012). Evaluating pathogenicity of rare variants from dilated cardiomyopathy in the exome era. Circulation. Cardiovascular Genetics, 5(2), 167–174. doi:10.1161/CIRCGENETICS.111.961805.
Bick, A. G., Flannick, J., Ito, K., Cheng, S., Vasan, R. S., Parfenov, M. G., et al. (2012). Burden of rare sarcomere gene variants in the Framingham and Jackson Heart Study cohorts. The American Journal of Human Genetics, 91(3), 513–519. doi:10.1016/j.ajhg.2012.07.017.
Abecasis, G. R., Auton, A., Brooks, L. D., DePristo, M. A., Durbin, R. M., Handsaker, R. E., et al. (2012). An integrated map of genetic variation from 1,092 human genomes. Nature, 491(7422), 56–65. doi:10.1038/nature11632.
Fu, W., O’Connor, T. D., Jun, G., Kang, H. M., Abecasis, G., Leal, S. M., et al. (2013). Analysis of 6,515 exomes reveals the recent origin of most human protein-coding variants. Nature, 493(7431), 216–220. doi:10.1038/nature11690.
Van Driest, S. L., Will, M. L., Atkins, D. L., & Ackerman, M. J. (2002). A novel TPM1 mutation in a family with hypertrophic cardiomyopathy and sudden cardiac death in childhood. The American Journal of Cardiology, 90(10), 1123–1127.
Kent, W. J., Sugnet, C. W., Furey, T. S., Roskin, K. M., Pringle, T. H., Zahler, A. M., et al. (2002). The human genome browser at UCSC. Genome Res, 12(6), 996–1006. doi:10.1101/gr.229102. Article published online before print in May 2002.
Kulikovskaya, I., McClellan, G. B., Levine, R., & Winegrad, S. (2007). Multiple forms of cardiac myosin-binding protein C exist and can regulate thick filament stability. The Journal of General Physiology, 129(5), 419–428. doi:10.1085/jgp.200609714.
Gautel, M., Zuffardi, O., Freiburg, A., & Labeit, S. (1995). Phosphorylation switches specific for the cardiac isoform of myosin binding protein-C: a modulator of cardiac contraction? Embo J, 14(9), 1952–1960.
Kapa, S., Tester, D. J., Salisbury, B. A., Harris-Kerr, C., Pungliya, M. S., Alders, M., et al. (2009). Genetic testing for long-QT syndrome: distinguishing pathogenic mutations from benign variants. Circulation, 120(18), 1752–1760. doi:10.1161/circulationaha.109.863076.
Kapplinger, J. D., Landstrom, A. P., Salisbury, B. A., Callis, T. E., Pollevick, G. D., Tester, D. J., et al. (2011). Distinguishing arrhythmogenic right ventricular cardiomyopathy/dysplasia-associated mutations from background genetic noise. JACC, 57(23), 2317–2327. doi:10.1016/j.jacc.2010.12.036.
Van Driest, S. L., Vasile, V. C., Ommen, S. R., Will, M. L., Tajik, A. J., Gersh, B. J., et al. (2004). Myosin binding protein C mutations and compound heterozygosity in hypertrophic cardiomyopathy. JACC, 44(9), 1903–1910. doi:10.1016/j.jacc.2004.07.045.
Ng, D., Johnston, J. J., Teer, J. K., Singh, L. N., Peller, L. C., Wynter, J. S., et al. (2013). Interpreting secondary cardiac disease variants in an exome cohort. Circulation. Cardiovascular Genetics. doi:10.1161/CIRCGENETICS.113.000039.
Gersh, B. J., Maron, B. J., Bonow, R. O., Dearani, J. A., Fifer, M. A., Link, M. S., et al. (2011). 2011 ACCF/AHA guideline for the diagnosis and treatment of hypertrophic cardiomyopathy: executive summary: a report of the American College of Cardiology Foundation/American Heart Association Task Force on Practice Guidelines. Circulation, 124(24), 2761–2796. doi:10.1161/CIR.0b013e318223e230.
Bos, J. M., Towbin, J. A., & Ackerman, M. J. (2009). Diagnostic, prognostic, and therapeutic implications of genetic testing for hypertrophic cardiomyopathy. JACC, 54(3), 201–211. doi:10.1016/j.jacc.2009.02.075.
Olivotto, I., Girolami, F., Ackerman, M. J., Nistri, S., Bos, J. M., Zachara, E., et al. (2008). Myofilament protein gene mutation screening and outcome of patients with hypertrophic cardiomyopathy. Mayo Clinic Proceedings, 83(6), 630–638. doi:10.4065/83.6.630.
Petrovski, S., Wang, Q., Heinzen, E. L., Allen, A. S., & Goldstein, D. B. (2013). Genic intolerance to functional variation and the interpretation of personal genomes. PLoS Genetics, 9(8), e1003709. doi:10.1371/journal.pgen.1003709.
MacArthur, D. G., Balasubramanian, S., Frankish, A., Huang, N., Morris, J., Walter, K., et al. (2012). A systematic survey of loss-of-function variants in human protein-coding genes. Science, 335(6070), 823–828. doi:10.1126/science.1215040.
Acknowledgments
J.D.K. is supported by the NIH grant GM72474-08 and thanks the Mayo Clinic MSTP for fostering an outstanding environment for physician-scientist training. This project was supported by the Mayo Clinic Windland Smith Rice Comprehensive Sudden Cardiac Death Program (MJA).
Conflict of Interest
B.A.S is an employee of Knome, Inc. T.E.C. is an employee of Transgenomic Inc. MJA is a consultant for Boston Scientific, Medtronic, St. Jude Medical, Inc., and Transgenomic. Intellectual property derived from MJA’s research program resulted in license agreements in 2004 between Mayo Clinic Health Solutions (formerly Mayo Medical Ventures) and PGxHealth (formerly Genaissance Pharmaceuticals, now assigned to Transgenomic) with respect to their FAMILION-LQTS and FAMILION-CPVT genetic tests but not their FAMILION-HCM genetic test. The other authors have no conflicts of interest to disclose. None of the disclosures pertain to this study and none of the companies provided financial support for this study.
Author information
Authors and Affiliations
Corresponding author
Additional information
Associate Editor Daniel P. Judge oversaw the review of this article
Jamie D. Kapplinger and Andrew P. Landstrom contributed equally and are co-equal first authors.
Rights and permissions
About this article
Cite this article
Kapplinger, J.D., Landstrom, A.P., Bos, J.M. et al. Distinguishing Hypertrophic Cardiomyopathy-Associated Mutations from Background Genetic Noise. J. of Cardiovasc. Trans. Res. 7, 347–361 (2014). https://doi.org/10.1007/s12265-014-9542-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12265-014-9542-z