Abstract
The techniques of diffusion analysis based on optical microscopy approaches have revealed a great diversity of the dynamic organisation of cell membranes. For a long period, two frameworks have dominated the way of representing the membrane structure: the membrane skeleton fences and the lipid raft models. Progresses in the methods of data analysis have shed light on the features and consequently the possible origin of membrane domains: Inter-protein interactions play a role in confinement. Innovative developments pushing forward the spatiotemporal resolution limits are currently emerging, which are likely to provide in the future a detailed understanding of the intimate functional dynamic organisation of the cell membrane.

Similar content being viewed by others
References
Kahya N (2006) Chem Phys Lipids 141:158–168
Simons K, van Meer G (1988) Biochemistry 27:6197–6202
Jacobson K, Mouritsen OG, Anderson RG (2007) Nat Cell Biol 9:7–14
London E (2005) Biochim Biophys Acta 1746:203–220
Mishra S, Joshi PG (2007) J Neurochem 103(Suppl 1):135–142
Singer SJ, Nicolson GL (1972) Science 175:720–731
Yechiel E, Edidin M (1987) J Cell Biol 105:755–760
Edidin M, Stroynowski I (1991) J Cell Biol 112:1143–1150
van Meer G (2005) EMBO J 24:3159–3165
Sharma P, Varma R, Sarasij RC, Ira Gousset K, Krishnamoorthy G, Rao M, Mayor S (2004) Cell 116:577–589
Baumgart T, Hess ST, Webb WW (2003) Nature 425:821–824
Dumas F, Sperotto MM, Lebrun MC, Tocanne JF, Mouritsen OG (1997) Biophys J 73:1940–1953
Sprong H, van der Sluijs P, van Meer G (2001) Nat Rev Mol Cell Biol 2:504–513
Abney JR, Scalettar BA (1995) Biophys Chem 57:27–36
Daumas F, Destainville N, Millot C, Lopez A, Dean D, Salome L (2003) Biophys J 84:356–366
Kusumi A, Sako Y, Yamamoto M (1993) Biophys J 65:2021–2040
Fujiwara T, Ritchie K, Murakoshi H, Jacobson K, Kusumi A (2002) J Cell Biol 157:1071–1081
Sheetz MP (1983) Semin Hematol 20:175–188
Tsuji A, Kawasaki K, Ohnishi S, Merkle H, Kusumi A (1988) Biochemistry 27:7447–7452
Mouritsen OG (2005) Life as a matter or fat: the emerging science of lipidomic. Springer, Berlin
Brown DA, London E (1998) Annu Rev Cell Dev Biol 14:111–136
Sun X, Whittaker GR (2003) J Virol 77:12543–12551
Ono A, Freed EO (2001) Proc Natl Acad Sci U S A 98:13925–13930
Jolly C, Sattentau QJ (2005) J Virol 79:12088–12094
Moffett S, Brown DA, Linder ME (2000) J Biol Chem 275:2191–2198
Baumgart T, Hammond AT, Sengupta P, Hess ST, Holowka DA, Baird BA, Webb WW (2007) Proc Natl Acad Sci USA 104:3165–3170
Schroeder RJ, Ahmed SN, Zhu Y, London E, Brown DA (1998) J Biol Chem 273:1150–1157
Wolf AA, Fujinaga Y, Lencer WI (2002) J Biol Chem 277:16249–16256
Schuck S, Honsho M, Ekroos K, Shevchenko A, Simons K (2003) Proc Natl Acad Sci U S A 100:5795–5800
Chen Y, Yang B, Jacobson K (2004) Lipids 39:1115–1119
Heerklotz H (2002) Biophys J 83:2693–2701
Simons K, Toomre D (2000) Nat Rev Mol Cell Biol 1:31–39
Foret L (2005) Europhys Lett 71:508–514
Turner MS, Sens P, Socci ND (2005) Phys Rev Lett 95:168301
Yethiraj A, Weisshaar JC (2007) Biophys J 93:3113–3119
Dean DS, Manghi M (2006) Phys Rev E Stat Nonlin Soft Matter Phys 74:021916
Joly E (2004) BMC Cell Biol 5:3
Kenworthy AK, Nichols BJ, Remmert CL, Hendrix GM, Kumar M, Zimmerberg J, Lippincott-Schwartz J (2004) J Cell Biol 165:735–746
Dietrich C, Yang B, Fujiwara T, Kusumi A, Jacobson K (2002) Biophys J 82:274–284
Munro S (2003) Cell 115:377–388
Pike LJ (2006) J Lipid Res 47:1597–1598
Jin S, Verkman AS (2007) J Phys Chem B 111:3625–3632
Saffarian S, Li Y, Elson EL, Pike LJ (2007) Biophys J 93:1021–1031
Pink D (1985) Biochim Biophys Acta 818:200–204
Pink DA, Laidlaw DJ, Chisholm DM (1986) Biochim Biophys Acta 863:9–17
Abney JR, Scalettar BA, Owicki JC (1989) Biophys J 55:817–833
Zwanzig RW (2001) Nonequilibrium statistical mechanics. Oxford University Press, USA
Meilhac N, Le Guyader L, Salome L, Destainville N (2006) Phys Rev E Stat Nonlin Soft Matter Phys 73:011915
Cherry RJ, Godfrey RE, Peters R (1982) Biochem Soc Trans 10:342–343
Murase K, Fujiwara T, Umemura Y, Suzuki K, Iino R, Yamashita H, Saito M, Murakoshi H, Ritchie K, Kusumi A (2004) Biophys J 86:4075–4093
Jacquier V, Prummer M, Segura JM, Pick H, Vogel H (2006) Proc Natl Acad Sci U S A 103:14325–14330
Smith SM, Lei Y, Liu J, Cahill ME, Hagen GM, Barisas BG, Roess DA (2006) Endocrinology 147:1789–1795
Sauliere A (2007) Université Paul Sabatier, Toulouse, p 150
Frick M, Schmidt K, Nichols BJ (2007) Curr Biol 17:462–467
Destainville N (2008) Phys Rev E Stat Nonlin Soft Matter Phys 77:011913
Stradner A, Sedgwick H, Cardinaux F, Poon WC, Egelhaaf SU, Schurtenberger P (2004) Nature 432:492–495
Segrè PN, Prasad V, Schofield AB, DA Weitz DA (2001) Phys Rev Lett 86:6042–6045
Sear RP, Gelbart WM (1999) J Chem Phys 110:4582–4588
Groenewold J, Kegel WK (2001) J Phys Chem B 105:11702–11709
Destainville N, Salome L (2006) Biophys J 90:L17–L19
Reynwar BJ, Illya G, Harmandaris VA, Muller MM, Kremer K, Deserno M (2007) Nature 447:461–464
Periole X, Huber T, Marrink SJ, Sakmar TP (2007) J Am Chem Soc 129:10126–10132
Fournier JB, Dommersnes PG, Galatola P (2003) C R Biologies 326:467–476
Alberts B, Johnson A, Lewis J, Raff M, Roberts K, Walter P (2002) Molecular biology of the cell. Garland Science, New York
Romer W, Berland L, Chambon V, Gaus K, Windschiegl B, Tenza D, Aly MR, Fraisier V, Florent JC, Perrais D, Lamaze C, Raposo G, Steinem C, Sens P, Bassereau P, Johannes L (2007) Nature 450:670–675
Borodich A, Rojdestvenski I, Cottam M (2003) Biophys J 85:774–789
Sieber JJ, Willig KI, Kutzner C, Gerding-Reimers C, Harke B, Donnert G, Rammner B, Eggeling C, Hell SW, Grubmuller H, Lang T (2007) Science 317:1072–1076
Sieber JJ, Willig KI, Heintzmann R, Hell SW, Lang T (2006) Biophys J 90:2843–2851
Donnert G, Keller J, Medda R, Andrei MA, Rizzoli SO, Luhrmann R, Jahn R, Eggeling C, Hell SW (2006) Proc Natl Acad Sci U S A 103:11440–11445
Davare MA, Avdonin V, Hall DD, Peden EM, Burette A, Weinberg RJ, Horne MC, Hoshi T, Hell JW (2001) Science 293:98–101
Park PS, Palczewski K (2005) Nat Chem Biol 1:184–185
Zhang J, Campbell RE, Ting AY, Tsien RY (2002) Nat Rev Mol Cell Biol 3:906–918
Shaner NC, Steinbach PA, Tsien RY (2005) Nat Methods 2:905–909
Fu CC, Lee HY, Chen K, Lim TS, Wu HY, Lin PK, Wei PK, Tsao PH, Chang HC, Fann W (2007) Proc Natl Acad Sci U S A 104:727–732
Silvius JR, Nabi IR (2006) Mol Membr Biol 23:5–16
Boyer D, Tamarat P, Maali A, Lounis B, Orrit M (2002) Science 297:1160–1163
Cognet L, Tardin C, Boyer D, Choquet D, Tamarat P, Lounis B (2003) Proc Natl Acad Sci U S A 100:11350–11355
Fotiadis D, Liang Y, Filipek S, Saperstein DA, Engel A, Palczewski K (2003) Nature 421:127–128
Czajkowsky DM, Hotze EM, Shao Z, Tweten RK (2004) EMBO J 23:3206–3215
Scheuring S, Sturgis JN (2005) Science 309:484–487
Bodnar A, Bacso Z, Jenei A, Jovin TM, Edidin M, Damjanovich S, Matko J (2003) Int Immunol 15:331–339
Ianoul A, Grant DD, Rouleau Y, Bani-Yaghoub M, Johnston LJ, Pezacki JP (2005) Nat Chem Biol 1:196–202
de Bakker BI, de Lange F, Cambi A, Korterik JP, van Dijk EM, van Hulst NF, Figdor CG, Garcia-Parajo MF (2007) Chemphyschem 8:1473–1480
Chen Y, Shao L, Ali Z, Cai J, Chen ZW (2008) Blood (in press)
Chen Y, Lagerholm BC, Yang B, Jacobson K (2006) Methods 39:147–153
Soumpasis DM (1983) Biophys J 41:95–97
Salome L, Cazeils JL, Lopez A, Tocanne JF (1998) Eur Biophys J 27:391–402
Cezanne L, Lecat S, Lagane B, Millot C, Vollmer JY, Matthes H, Galzi JL, Lopez A (2004) J Biol Chem 279:45057–45067
Baker A, Sauliere A, Gaibelet G, Lagane B, Mazeres S, Fourage M, Bachelerie F, Salome L, Lopez A, Dumas F (2008) J Biol Chem (in press)
Rigler R, Elson ES (2001) Fluorescence correlation spectroscopy: theory and applications. Springer, New York
Wawrezinieck L, Rigneault H, Marguet D, Lenne PF (2005) Biophys J 89:4029–4042
Wenger J, Conchonaud F, Dintinger J, Wawrezinieck L, Ebbesen TW, Rigneault H, Marguet D, Lenne PF (2007) Biophys J 92:913–919
Haustein E, Schwille P (2007) Annu Rev Biophys Biomol Struct 36:151–169
Bouzigues C, Morel M, Triller A, Dahan M (2007) Proc Natl Acad Sci U S A 104:11251–11256
Groc L, Heine M, Cousins SL, Stephenson FA, Lounis B, Cognet L, Choquet D (2006) Proc Natl Acad Sci U S A 103:18769–18774
Cheezum MK, Walker WF, Guilford WH (2001) Biophys J 81:2378–2388
Saxton MJ, Jacobson K (1997) Annu Rev Biophys Biomol Struct 26:373–399
Bouzigues C, Dahan M (2007) Biophys J 92:654–660
Huet S, Karatekin E, Tran VS, Fanget I, Cribier S, Henry JP (2006) Biophys J 91:3542–3559
Bates IR, Hebert B, Luo Y, Liao J, Bachir AI, Kolin DL, Wiseman PW, Hanrahan JW (2006) Biophys J 91:1046–1058
Killian JA (1998) Biochim Biophys Acta 1376:401–415
Salamon Z, Cowell S, Varga E, Yamamura HI, Hruby VJ, Tollin G (2000) Biophys J 79:2463–2474
Fournier JB (1998) Eur Phys J B 11:261–272
Gil T, Ipsen GH, Mouritsen OG, Sabra MC, Sperotto MM, Zuckermann MJ (1998) Biochim Biophys Acta 1376:245–266
Sabra MC, Mouritsen OG (1998) Biophys J 74:745–752
Likos CN (2001) Phys Rep 348:267–439
Goulian M, Bruinsma R, Pincus P (1993) Europhys Lett 22:145–150
Park JM, Lubensky TC (1996) J Phys I 6:1217–1235
Fournier JB, Dommersnes PG (1997) Europhys Lett 39:681–682
Chou T, Kim KS, Oster G (2001) Biophys J 80:1075–1087
Deisenhofer J, Epp O, Sinning I, Michel H (1995) J Mol Biol 246:429–457
Dommersnes PG, Fournier JB (1999) Europhys Lett 46:256–261
Naji A, Brown FL (2007) J Chem Phys 126:235103
Kim KS, Neu JC, Oster GF (1999) Europhys Lett 48:99–105
Dommersnes PG, Fournier JB (1999) Eur Phys J B 12:9–12
Destainville N (2008) Soft Matter. DOI 10.1039/b718583a 4:1288–1301
Acknowledgements
We thank B. Larijani for her positive criticism and help to improve this manuscript. We also are grateful to the reviewers for their suggestions.
Author information
Authors and Affiliations
Corresponding authors
Additional information
N. Destainville, F. Dumas and L. Salomé contributed equally to this work.
Appendices
Appendix 1. Methods for the analysis of diffusion
The analysis of the diffusion of molecules in cells is usually performed by optical microscopy. FRAP and FCS provide average measurements, while single-particle or single-molecule tracking methods (SPT or SMT) address the individual behaviours of the molecules. Although single-molecule approaches directly detect and analyse sub-populations, they present drawbacks discussed in the following section that make the global measurement methods FRAP and FCS still of valuable interest.
For global measurement of molecule dynamics within the membrane, methods such as variable radius FRAP or variable waist FCS are advantageous over the single-molecule methods. For a detailed description of these methods, see [85]. For the scope of this review, we discuss how these methods have been exploited to characterise the compartmentalisation of lipids and proteins in the membrane.
Fluorescence recovery after photobleaching
The fluorescence intensity is monitored in a geometrically defined zone after having photobleached the probes by the application of an intense and extremely brief light pulse (Fig. 2). The analysis of the fluorescence recovery curve due to the diffusion of the intact fluorescent molecules derived from outside of this zone determines the diffusion coefficient D of the molecules (the accessible range is between 10−3 and 10μm2/s) and the fraction of mobile molecules M. These parameters are deduced from the fit of the normalised fluorescence intensity recovery curve by the expected solution of the diffusion equations that are dependent on the intensity profile of the bleaching beam. This has been established, in the case of a uniform illumination disk of radius R by Soumpasis [86].
Principle of a fluorescence recovery after photobleaching experiment. a The initial fluorescence intensity I o of the labelled molecules (green “sparks”) is measured in a defined illuminated region of the membrane (blue zone). b A bleaching pulse is applied leading to a reduced fluorescence intensity I b. c Due to the diffusion of the molecules, a fluorescence recovery is observed. d The recording of the fluorescence intensity as a function of time is then used to determine both the diffusion coefficient and the mobile fraction of the labelled molecules
where R is the radius of illumination; τ D = R 2/4D and I 0 and I 1 are the modified Bessel functions of the first order.
Microdomains that are smaller than 1μm down to about 200 nm can be characterised by varying the illumination radius from 1 to 5μm. As seen in Eq. 2, M and D are then expected to vary as a function of R [87].
where M p is the permanent mobile fraction. Depending on the M p value, two cases can be considered:
-
If M p ≈ 0, there is a single population of tracers confined in closed domains. The measured diffusion coefficient is an apparent one (D app) and the real diffusion coefficient inside the domains D conf can be evaluated from Eq. 3.
$$D_{{\text{conf}}} \;{\text{ = }}\;{{\text{1}} \mathord{\left/{\vphantom {{\text{1}} {\text{2}}}} \right.\kern-\nulldelimiterspace} {\text{2}}}\;D_{{\text{app}}} \left( {{L \mathord{\left/{\vphantom {L R}} \right.\kern-\nulldelimiterspace} R}} \right)^{\text{2}} {\text{.}}$$(3)
The validation of the analysis can then be obtained by verifying the invariance of D conf with R.
-
If M p > 0, the tracers are either inside juxtaposed but in open domains or distributed both inside and outside isolated closed domains.
An analysis of the recovery curves assuming two sub-populations of tracers is then necessary, leading to the determination of weight and diffusion coefficient of each population. Confined tracers present a diffusion coefficient, D conf, varying according to Eq. 3, while non-confined tracers present a diffusion coefficient, D free, invariant with R. This last population corresponds to tracers with a long-range free diffusion.
If D free > D conf, tracers are distributed both inside and outside isolated closed domains. Note that the relevance of the two diffusion coefficients analysis requires that the weight of the free diffusion population should be in the same range as M p.
Such an approach was first developed by Yechiel and Edidin [7] and Edidin and Stroynowski [8] and has been applied in the analysis of dynamic compartmentalisation of the NK2, hMOR, CCR5 and CD4 receptors [88, 89].
Fluorescence correlation spectroscopy
The fluorescence intensity is measured inside a small volume defined by a focalised laser beam (Fig. 3). The fluctuations of the fluorescence intensity result from the molecules that enter and escape the confocal volume. These fluctuations are recorded with a temporal resolution ranging typically from 1 ns to 1 min.
Principle of a fluorescence correlation spectroscopy measurement. a Labelled molecules (green “sparks”) diffusing in the membrane enter and leave a small observation zone defined by a focused laser beam. b The autocorrelation function of the fluorescence intensity is calculated from the fluctuations of fluorescence recorded over time
For a free Brownian diffusion, the diffusion time, τ d, is obtained from the autocorrelation function in Eq. 4
where N is the average number of molecules in the observation volume. The diffusion coefficient D of the molecule can then be deduced according to Eq. 5
where w corresponds to the transversal waist of the focused laser beam.
One of the principal advantages of FCS is its ability to measure diffusion constants lower than 0.001μm2/s and its accuracy for analysing fast diffusing molecules [90].
Currently, FCS is exploited at variable waist [91, 116]; therefore, the variation of D with w allows us to easily discriminate between two different diffusing populations.
Pure Brownian diffusion obeys the linear diffusion law τ d = f (w 2) equation. A negative value of the intercept at w = 0 corresponds to a compartmentalisation of proteins inside domains delineated by a meshwork. A positive intercept at w = 0 corresponds to a dynamic partition of molecules between isolated domains.
Typical FCS instruments have a spatial resolution of approximately 300nm that is determined by optical diffraction limit. However, by using metallic nano-apertures, 30-nm domains can be characterised [92]. Other variations of FCS instruments with multiple-colour cross-correlation have been developed [93].
Single-particle or single-molecule tracking
This methodology allows the direct observation of individual molecules at the surface of living cells with a nanometre scale precision in the position measurement by video-microscopy coupled to image analysis (Fig. 4).
Principle of the single-particle tracking and single-molecule tracking methodologies. a The molecule of interest is specifically labelled by a fluorophore (green “spark”) or a nanoparticle (usually attached via an antibody). b The trajectory of the labelled molecule is determined by video-microscopy coupled to image analysis
Probes used in SPT are nanometre particles such as latex spheres, gold colloids or nanocrystals. Generally, these particles are attached to the molecule of interest by means of antibodies or fragments of antibodies. This permits one to observe endogenously expressed proteins or receptors [94, 95]. Particular care must be taken to avoid non-specific binding and multiple attachments of multiple target molecules. This is circumvented in SMT, where the molecules of interest are fused to naturally fluorescent proteins. However, the use of Halo tag® or Snap tag® labelling, which restrict the labelling to proteins expressed at the membrane, ought to be preferred.
Both for SPT and SMT, the acquisition frequencies are comparable and can be decreased to a few milliseconds. Nevertheless, due to photobleaching, the duration of the acquisition with SMT cannot exceed more than 10s, while in the case of SPT it can vary between 1 and 30min.
After the establishment of the trajectories [96], the mean square displacement is calculated according Eq. 6.
where r is the vector between two positions separated by a time interval t and brackets denote a discrete average over images.
Diffusion modes are determined by the analysis of the MSD(t) best fits [97]:
-
A linear MSD(t) is characteristic from a random-free diffusion
$${\text{MSD}}\left( t \right)\;{\text{ = }}\;{\text{4}}Dt$$(7)where D is the diffusion coefficient of the protein.
-
A MSD(t) presenting a saturation at long term is characteristic of a pure confined diffusion
$${\text{MSD}}{\left( t \right)} = \frac{{L^{2} }}{3}{\left( {1 - \exp {\left( { - \frac{t}{\tau }} \right)}} \right)}$$(8)where τ = L 2/12D, L is the size of the domain and D the diffusion coefficient of the protein. For fast diffusing molecules, the detector time-averaging effect can be corrected by the use of a mathematical formula detailed in [60].
-
More complex behaviours such as transient confinement or other combinations of simple modes, that require a thorough statistical analysis can be evidenced. Algorithms have been developed for detecting confinement and jumps between adjacent domains [48, 60], directed motion [98] and multi-type motion [99].
Single molecule versus collective measurements
An interesting comparison between the collective and individual approaches has been recently performed by Bates et al. [100] who studied, under identical conditions, the dynamics of the transmembrane protein CFTR using FRAP, SPT and image correlation spectroscopy (ICS, the imaging analogue of FCS). They first observed differences between FRAP and ICS measurements: The diffusion coefficient obtained with FRAP was fourfold larger than that obtained by ICS. SPT experiments revealed complex movements of the proteins with a rapid diffusion of CFTR interspersed by intermittent trapping. Such behaviour remained unravelled by FRAP or ICS that are addressing populations of molecules instead of single molecules.
Owing to the extensive duration of the observations accessible to the single-particle tracking approach compared to single-fluorophore tracking, it is likely to be an unavoidable tool to address the membrane organisation. Nevertheless, efforts must be concentrated in the understanding of the effects of nanoparticles on the labelled molecules. The role of their nature, of their size and their interaction with the underlying membrane and the glycocalix will have to be rigorously quantified.
Appendix 2. Nature of protein–protein interactions
There is a rich variety of non-specific interactions between membrane proteins. Their nature is either electrostatic or thermodynamic. Thermodynamic interactions are mediated by the membrane. Note that the following discussion does not only concern transmembrane proteins but also peripheral proteins.
-
(a)
Short-range interactions
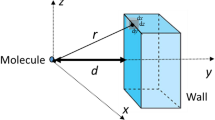
Short-range interactions are typically in a range of 1nm and involve energies of the order of magnitude of the thermal energy k B T. Dipole–dipole, Van der Waals forces always contribute to the short-range attraction between proteins. They decay rapidly at large distances (as 1/r 6) and add a binding energy of a fraction of k B T at short distance [66].
If the thickness of the hydrophobic core of a transmembrane protein is significantly different from the thickness of the membrane, the membrane will adjust so that the acyl chains of the lipids are not exposed to the aqueous medium. When the protein core is thicker than the membrane, the mismatch is said to be positive; conversely, it is negative. Membrane deformation costs a certain amount of elastic energy. Therefore, when two proteins inserted in the membrane [101] have the same mismatch sign, they attract. Conversely, they repulse if they present a different mismatch sign [101]. For instance, the activation of a receptor after ligand binding can change its hydrophobic thickness [102] and consequently the interaction potential with neighbouring proteins.
Furthermore, the shape of the protein can modulate hydrophobic interactions by deforming differently the lipid bilayer of the membrane [103]. Cell membranes are a mixture of many different lipids, with various fatty acid chain lengths. Therefore, lipids can be sorted by the protein to match its hydrophobic core. Mouritsen [20] and Gil et al. [104] have shown that an additional attraction between membrane proteins can occur to minimise the lipid/protein interface. This attraction leads to the formation of protein-rich domains. However, it is unclear why these domains are limited in size, i.e. why a single macro-domain does not recruit the majority of proteins. As in the case of the lipid rafts, this limitation in size may be due to the out-of-equilibrium thermodynamics of the cell [105].
Another short-range interaction known as the “depletion interaction” [106] occurs because the diameters of membrane proteins in the plane of the membrane are greater than the lipids [106]. The lipids surrounding an isolated protein exert an osmotic pressure that is parallel to the plane of the membrane. Due to the symmetry, the total force vanishes. However, if the distance between two proteins is smaller than the lipid diameter, the interval between the proteins will be devoid of lipids. Under these circumstances, the resulting force brings both proteins close to one another. The amplitude and range of this interaction are of the orders of one k B T and one nanometre [66].
-
(b)
Long-range interactions
The long-range interactions extend beyond ten nanometres. These interactions occur because embedded membrane proteins impose constraints on the membrane. Variations in entropy and elastic energy contribute to long-range interactions [107, 108].
Effects on long-range interactions of purely entropic origin are known as Casimir interactions. These Casimir attractive forces are related to the thermal fluctuations of the membrane [107]. Moreover, the influence of elastic energy on long-range interactions is due to the conical shape of the transmembrane proteins (with an aperture angle θ). This shape imposes a curvature on the membrane, inducing a repulsive force [107, 109]. Forces resulting from entropic and elastic energies decay slowly at the same rate, and their sum can be attractive or repulsive. This resulting force is repulsive if the protein curvature is superior to 5°. Given that the hydrophobic core of most transmembrane proteins, in particular G-protein coupled receptors (GPCRs), consists of several α-helices, it is unlikely that their shape can be approximated by a cylinder. For example, the average contact angle estimated for rhodopsin is greater than 10° [110, 111]. Peripheral proteins are also affected by both Casimir attractive force and elastic repulsion, since they impose an intrinsic local curvature when they bind to the membrane. The curvature anticipated in [63] is large enough to override the entropic attraction. Thus, it can be considered, in a first approximation, that all membrane proteins experience repulsion at long distances.
A more refined analysis of these longer range forces should also take into account that proteins structure is anisotropic (e.g. their cross-section in the plane of the membrane is not circular). This is achieved by modelling proteins by ellipses. It has been shown [108] that this can decrease the long-distance repulsion and even lead to attraction [110]. Finally, it has been argued that a torque applied to the proteins is also susceptible to affect the long-range pair potentials and even to enhance them [112]. This torque is, for example, the result of the electric field due to the difference of ion concentrations on both sides of the membrane.
To sum up, the coupling between the membrane curvature and the protein intrinsic curvature generically leads to a weak repulsive force. It is worth noting that this coupling enhances the protein rate of diffusion [113].
-
(c)
Effects of short- and long-range interactions
Short- and long-range interactions result in attractive or repulsive forces between two proteins. Figure 5 illustrates the energetic profile of protein–protein interactions. Hard-core repulsion occurs at distances shorter than a few nanometres and, at intermediate distances, a short-range binding potential well, of a few k B T, forms. At longer distances, repulsive forces dominate [55]. This model only takes into account pairwise interaction potentials. A more refined analysis should include many-body interactions [106, 108]. For instance, it has been shown that, under certain circumstances, repulsive forces between pairs of proteins can be counter-balanced by the attractive forces due to many-body attractions [114, 115].
Typical shape of the interaction potential between membrane proteins. F(r) is the free energy, in units of the thermal energy k B T; r is the distance between the proteins in nanometres. A hard-core repulsion occurs for distances inferior or equal to the protein diameter. Then, a narrow attraction, with a binding energy of a few k B T, occurs due to different short-range interactions discussed in “Appendix 2”. Beyond an energy barrier of a fraction of k B T, the potential decreases. The values chosen in this example are those used in [55]
Rights and permissions
About this article
Cite this article
Destainville, N., Dumas, F. & Salomé, L. What do diffusion measurements tell us about membrane compartmentalisation? Emergence of the role of interprotein interactions. J Chem Biol 1, 37–48 (2008). https://doi.org/10.1007/s12154-008-0005-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12154-008-0005-3