Abstract
Purpose
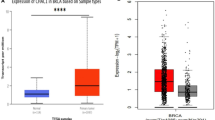
The aberrant mRNA expression of a UPR component Cation transport regulator homolog 1 (CHAC1) has been reported to be associated with poor survival in breast and ovarian cancer patients, however, the expression of CHAC1 at protein levels in malignant breast tissues is underreported. The following study aimed at analyzing CHAC1 protein expression in malignant breast cancer tissues.
Methods
Evaluation of CHAC1 expression in invasive ductal carcinomas (IDCs) with known ER, PR, and HER2 status was carried out using immunohistochemistry (IHC) with CHAC1 specific antibody. The Human breast cancer tissue microarray (TMA, cat# BR1503f, US Biomax, Inc., Rockville, MD) was used to determine CHAC1 expression. The analysis of CHAC1 IHC was done to determine its expression in terms of molecular subtypes of breast cancer, lymph node status, and proliferation index using Qu-Path software. Survival analysis was studied with a Kaplan–Meier plotter.
Results
Immunohistochemical analysis of CHAC1 in breast cancer tissues showed significant up-regulation of CHAC1 as compared to the adjacent normal and benign tissues. Interestingly, CHAC1 immunostaining revealed high expression in tumor tissues with high proliferation and positive lymph node metastasis suggesting that CHAC1 might have an important role to play in breast cancer progression. Furthermore, high CHAC1 expression is associated with poor overall survival (OS) in large breast cancer patient cohorts.
Conclusion
As a higher expression of CHAC1 was observed in tissue cores with high Ki67 index and positive lymph node metastasis it may be concluded that enhanced CHAC1 expression correlates with proliferation and metastasis. The further analysis of breast cancer patients’ survival data through KM plot indicated that high CHAC1 expression is associated with a bad prognosis hinting that CHAC1 may have a possible prognostic significance in breast cancer.





Similar content being viewed by others
References:
T Avril E Vauleon E Chevet 2017 Endoplasmic reticulum stress signaling and chemotherapy resistance in solid cancers Oncogenesis 6 8 e373 https://doi.org/10.1038/oncsis.2017.72
Clarke HJ, Chambers JE, Liniker E, Marciniak SJ. Endoplasmic reticulum stress in malignancy. Cancer Cell. 2014;25(5):563–73. https://doi.org/10.1016/j.ccr.2014.03.015.
Madden E, Logue SE, Healy SJ, Manie S, Samali A. The role of the unfolded protein response in cancer progression: from oncogenesis to chemoresistance. Biol Cell. 2019;111(1):1–17. https://doi.org/10.1111/boc.201800050.
Chevet E, Hetz C, Samali A. Endoplasmic reticulum stress–activated cell reprogramming in oncogenesis. Cancer Discov. 2015;5(6):586–97. https://doi.org/10.1158/2159-8290.CD-14-1490.
McGrath EP, Logue SE, Mnich K, Deegan S, Jäger R, Gorman AM, et al. The unfolded protein response in breast cancer. Cancers. 2018;10(10):344. https://doi.org/10.3390/cancers10100344.
Mungrue IN, Pagnon J, Kohannim O, Gargalovic PS, Lusis AJ. CHAC1/MGC4504 is a novel proapoptotic component of the unfolded protein response, downstream of the ATF4-ATF3-CHOP cascade. J Immunol. 2009;182(1):466–76. https://doi.org/10.4049/jimmunol.182.1.466.
Kumar A, Tikoo S, Maity S, Sengupta S, Sengupta S, Kaur A, et al. Mammalian proapoptotic factor ChaC1 and its homologues function as γ-glutamyl cyclotransferases acting specifically on glutathione. EMBO Rep. 2012;13(12):1095–101. https://doi.org/10.1038/embor.2012.156.
Gargalovic PS, Imura M, Zhang B, Gharavi NM, Clark MJ, Pagnon J, et al. Identification of inflammatory gene modules based on variations of human endothelial cell responses to oxidized lipids. Proc Natl Acad Sci. 2006;103(34):12741–6. https://doi.org/10.1073/pnas.0605457103.
Perra L, Balloy V, Foussignière T, Moissenet D, Petat H, Mungrue IN, et al. CHAC1 is differentially expressed in normal and cystic fibrosis bronchial epithelial cells and regulates the inflammatory response induced by Pseudomonas aeruginosa. Front Immunol. 2018;9:2823. https://doi.org/10.3389/fimmu.2018.02823.
Li S, Jia Y, Xue M, Hu F, Zheng Z, Zhang S, et al. Inhibiting Rab27a in renal tubular epithelial cells attenuates the inflammation of diabetic kidney disease through the miR-26a-5p/CHAC1/NF-kB pathway. Life Sci. 2020;261: 118347. https://doi.org/10.1016/j.lfs.2020.118347.
Chen M-S, Wang S-F, Hsu C-Y, Yin P-H, Yeh T-S, Lee H-C, et al. CHAC1 degradation of glutathione enhances cystine-starvation-induced necroptosis and ferroptosis in human triple negative breast cancer cells via the GCN2-eIF2α-ATF4 pathway. Oncotarget. 2017;8(70): 114588. https://doi.org/10.18632/oncotarget.23055.
Wang N, Zeng G-Z, Yin J-L, Bian Z-X. Artesunate activates the ATF4-CHOP-CHAC1 pathway and affects ferroptosis in Burkitt’s Lymphoma. Biochem Biophys Res Commun. 2019;519(3):533–9. https://doi.org/10.1016/j.bbrc.2019.09.023.
Ogawa T, Wada Y, Takemura K, Board PG, Uchida K, Kitagaki K, et al. CHAC1 overexpression in human gastric parietal cells with Helicobacter pylori infection in the secretory canaliculi. Helicobacter. 2019;24(4): e12598. https://doi.org/10.1111/hel.12598.
Wada Y, Takemura K, Tummala P, Uchida K, Kitagaki K, Furukawa A, et al. Helicobacter pylori induces somatic mutations in TP 53 via overexpression of CHAC 1 in infected gastric epithelial cells. FEBS Open Bio. 2018;8(4):671–9. https://doi.org/10.1002/2211-5463.12402.
Liu Y, Li M, Shi D, Zhu Y. Higher expression of cation transport regulator-like protein 1 (CHAC1) predicts of poor outcomes in uveal melanoma (UM) patients. Int Opthalmol. 2019;39(12):2825–32. https://doi.org/10.1007/s10792-019-01129-1.
Stelzer Y, Yanuka O, Benvenisty N. Global analysis of parental imprinting in human parthenogenetic induced pluripotent stem cells. Nat Struct Biol. 2011;18(6):735. https://doi.org/10.1038/nsmb.2050.
Chen X, Iliopoulos D, Zhang Q, Tang Q, Greenblatt MB, Hatziapostolou M, et al. XBP1 promotes triple-negative breast cancer by controlling the HIF1α pathway. Nature. 2014;508(7494):103–7. https://doi.org/10.1038/nature13119.
Cook KL, Clarke R. Role of GRP78 in promoting therapeutic-resistant breast cancer. Future Med Chem. 2015;7(12):1529–34. https://doi.org/10.4155/FMC.15.80.
Goebel G, Berger R, Strasak A, Egle D, Müller-Holzner E, Schmidt S, et al. Elevated mRNA expression of CHAC1 splicing variants is associated with poor outcome for breast and ovarian cancer patients. Br J Cancer. 2012;106(1):189–98. https://doi.org/10.1038/bjc.2011.510.
Jahn B, Arvandi M, Rochau U, Fiegl H, Goebel G, Marth C, et al. Development of a novel prognostic score for breast cancer patients using mRNA expression of CHAC1. J Comp Eff Res. 2017;6(7):563–74. https://doi.org/10.2217/cer-2017-0015.
Chander H, Truesdell P, Meens J, Craig AW. Transducer of Cdc42-dependent actin assembly promotes breast cancer invasion and metastasis. Oncogene. 2013;32(25):3080–90. https://doi.org/10.1038/onc.2012.317.
Suman P, Mehta V, Craig AW, Chander H. Wild-type p53 suppresses formin-binding protein-17 (FBP17) to reduce invasion. Carcinogenesis. 2022;43(5):494–503. https://doi.org/10.1093/carcin/bgac015.
Bankhead P, Loughrey MB, Fernandez JA, Dombrowski Y, McArt DG, Dunne PD, et al. QuPath: open source software for digital pathology image analysis. Sci Rep. 2017;7(1):16878. https://doi.org/10.1038/s41598-017-17204-5.
Ramachandran P, Dobie R, Wilson-Kanamori J, Dora E, Henderson B, Luu N, et al. Resolving the fibrotic niche of human liver cirrhosis at single-cell level. Nature. 2019;575(7783):512–8. https://doi.org/10.1038/s41586-019-1631-3.
Györffy B, Lanczky A, Eklund AC, Denkert C, Budczies J, Li Q, et al. An online survival analysis tool to rapidly assess the effect of 22,277 genes on breast cancer prognosis using microarray data of 1,809 patients. Breast Cancer Res Treat. 2010;123(3):725–31. https://doi.org/10.1007/s10549-009-0674-9.
Rivenbark AG, O’Connor SM, Coleman WB. Molecular and cellular heterogeneity in breast cancer: challenges for personalized medicine. Am J Pathol. 2013;183(4):1113–24. https://doi.org/10.1016/j.ajpath.2013.08.002.
Butti R, Das S, Gunasekaran VP, Yadav AS, Kumar D, Kundu GC. Receptor tyrosine kinases (RTKs) in breast cancer: signaling, therapeutic implications and challenges. Mol Cancer. 2018;17(1):34. https://doi.org/10.1186/s12943-018-0797-x.
Du Z, Lovly CM. Mechanisms of receptor tyrosine kinase activation in cancer. Mol Cancer. 2018;17(1):58. https://doi.org/10.1186/s12943-018-0782-4.
Weigelt B, Ng CK, Shen R, Popova T, Schizas M, Natrajan R, et al. Metastatic breast carcinomas display genomic and transcriptomic heterogeneity. Mod Pathol. 2015;28(3):340–51. https://doi.org/10.1038/modpathol.2014.163.
Mehta V, Chander H, Munshi A. Complex roles of discoidin domain receptor tyrosine kinases in cancer. Clin TranslOncol. 2021;23(8):1497–1510. https://doi.org/10.1007/s12094-021-02552-6.
Prat A, Pineda E, Adamo B, Galván P, Fernández A, Gaba L, et al. Clinical implications of the intrinsic molecular subtypes of breast cancer. The Breast. 2015;24:S26–35. https://doi.org/10.1016/j.breast.2015.07.008.
Cheang MC, Chia SK, Voduc D, Gao D, Leung S, Snider J, et al. Ki67 index, HER2 status, and prognosis of patients with luminal B breast cancer. J Natl Cancer Inst. 2009;101(10):736–50. https://doi.org/10.1093/jnci/djp082.
Feeley LP, Mulligan AM, Pinnaduwage D, Bull SB, Andrulis IL. Distinguishing luminal breast cancer subtypes by Ki67, progesterone receptor or TP53 status provides prognostic information. Mod Pathol. 2014;27(4):554–61. https://doi.org/10.1038/modpathol.2013.153.
De Azambuja E, Cardoso F, de Castro G, Colozza M, Mano MS, Durbecq V, et al. Ki-67 as prognostic marker in early breast cancer: a meta-analysis of published studies involving 12 155 patients. Br J Cancer. 2007;96(10):1504–13. https://doi.org/10.1038/sj.bjc.6603756.
Hashmi AA, Hashmi KA, Irfan M, Khan SM, Edhi MM, Ali JP, et al. Ki67 index in intrinsic breast cancer subtypes and its association with prognostic parameters. BMC Res Notes. 2019;12(1):1–5. https://doi.org/10.1186/s13104-019-4653-x.
Yerushalmi R, Woods R, Ravdin PM, Hayes MM, Gelmon KA. Ki67 in breast cancer: prognostic and predictive potential. Lancet Oncol. 2010;11(2):174–83. https://doi.org/10.1016/S1470-2045(09)70262-1.
Gao J, Aksoy BA, Dogrusoz U, Dresdner G, Gross B, Sumer SO et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal. 2013;6(269):pl1-pl1. https://doi.org/10.1126/scisignal.2004088.
Cerami E, Gao J, Dogrusoz U, Gross BE, Sumer SO, Aksoy BA et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. AACR; 2012.
Ji R-C. Lymph nodes and cancer metastasis: new perspectives on the role of intranodal lymphatic sinuses. Int J Mol Sci. 2017;18(1):51. https://doi.org/10.3390/ijms18010051.
Stacker SA, Achen MG, Jussila L, Baldwin ME, Alitalo K. Lymphangiogenesis and cancer metastasis. Nat Rev Cancer. 2002;2(8):573–83. https://doi.org/10.1038/nrc863.
Mehta V, Meena J, Kasana H, Munshi A, Chander H. Prognostic significance of CHAC1 expression in breast cancer. Mol Biol Rep. 2022:1–10. doi:https://doi.org/10.1007/s11033-022-07673-x
Russnes HG, Navin N, Hicks J, Borresen-Dale A-L. Insight into the heterogeneity of breast cancer through next-generation sequencing. J Clin Invest. 2011;121(10):3810–8. https://doi.org/10.1172/JCI57088.
Kalinowski L, Saunus JM, Reed AEM, Lakhani SR. Breast Cancer Heterogeneity in Primary and Metastatic Disease. Breast Cancer Metastasis and Drug Resistance. Springer; 2019. p. 75–104.
Riggio AI, Varley KE, Welm AL. The lingering mysteries of metastatic recurrence in breast cancer. Br J Cancer. 2020;124(1):1–14. https://doi.org/10.1038/s41416-020-01161-4.
Turashvili G, Brogi E. Tumor heterogeneity in breast cancer. Front Med. 2017;4:227. https://doi.org/10.3389/fmed.2017.00227.
Pece S, Tosoni D, Confalonieri S, Mazzarol G, Vecchi M, Ronzoni S, et al. Biological and molecular heterogeneity of breast cancers correlates with their cancer stem cell content. Cell. 2010;140(1):62–73. https://doi.org/10.1016/j.cell.2009.12.007.
Kamdje AHN, Etet PFS, Vecchio L, Muller JM, Krampera M, Lukong KE. Signaling pathways in breast cancer: therapeutic targeting of the microenvironment. Cell Signal. 2014;26(12):2843–56. https://doi.org/10.1016/j.cellsig.2014.07.034.
Eroles P, Bosch A, Pérez-Fidalgo JA, Lluch A. Molecular biology in breast cancer: intrinsic subtypes and signaling pathways. Cancer Treat Rev. 2012;38(6):698–707. https://doi.org/10.1016/j.ctrv.2011.11.005.
Scriven P, Coulson S, Haines R, Balasubramanian S, Cross S, Wyld L. Activation and clinical significance of the unfolded protein response in breast cancer. Br J Cancer. 2009;101(10):1692–8. https://doi.org/10.1038/sj.bjc.6605365.
Suman P, Mishra S, Chander H. High formin binding protein 17 (FBP17) expression indicates poor differentiation and invasiveness of ductal carcinomas. Sci Rep. 2020;10(1):1–10. https://doi.org/10.1038/s41598-020-68454-9.
Giudici F, Petracci E, Nanni O, Bottin C, Pinamonti M, Zanconati F, et al. Elevated levels of eEF1A2 protein expression in triple negative breast cancer relate with poor prognosis. PLoS ONE. 2019;14(6): e0218030. https://doi.org/10.1371/journal.pone.0218030.
Cerqueira OL, Truesdell P, Baldassarre T, Vilella-Arias SA, Watt K, Meens J, et al. CIP4 promotes metastasis in triple-negative breast cancer and is associated with poor patient prognosis. Oncotarget. 2015;6(11):9397. https://doi.org/10.18632/oncotarget.3351.
van de Vijver MJ, Peterse JL, Mooi WJ, Wisman P, Lomans J, Dalesio O, et al. Neu-protein overexpression in breast cancer. N Engl J Med. 1988;319(19):1239–45. https://doi.org/10.1056/NEJM198811103191902.
Zhang K, Liu H, Song Z, Jiang Y, Kim H, Samavati L, et al. The UPR Transducer IRE1 promotes breast cancer malignancy by degrading tumor suppressor microRNAs. Iscience. 2020;23(9): 101503. https://doi.org/10.1016/j.isci.2020.101503.
Li H, Chen X, Gao Y, Wu J, Zeng F, Song F. XBP1 induces snail expression to promote epithelial-to-mesenchymal transition and invasion of breast cancer cells. Cell Signal. 2015;27(1):82–9. https://doi.org/10.1016/j.cellsig.2014.09.018.
Pathmanathan N, Balleine RL. Ki67 and proliferation in breast cancer. J Clin Pathol. 2013;66(6):512–6. https://doi.org/10.1136/jclinpath-2012-201085.
van Diest PJ, van der Wall E, Baak JP. Prognostic value of proliferation in invasive breast cancer: a review. J Clin Pathol. 2004;57(7):675–81. https://doi.org/10.1136/jcp.2003.010777.
Gerashchenko TS, Novikov NM, Krakhmal NV, Zolotaryova SY, Zavyalova MV, Cherdyntseva NV, et al. Markers of cancer cell invasion: are they good enough? J Clin Med. 2019;8(8):1092. https://doi.org/10.3390/jcm8081092.
Wu J, Wang X, Wang N, Ma L, Xie X, Zhang H, et al. Identification of novel antioxidant gene signature to predict the prognosis of patients with gastric cancer. World J Surg Oncol. 2021;19(1):1–11. https://doi.org/10.1186/s12957-021-02328-w.
Xiao R, Wang S, Guo J, Liu S, Ding A, Wang G, et al. Ferroptosis-related gene NOX4, CHAC1 and HIF1A are valid biomarkers for stomach adenocarcinoma. J Cell Mol Med. 2022. https://doi.org/10.1111/jcmm.17171.
Hong Y, Lin M, Ou D, Huang Z, Shen P. A novel ferroptosis-related 12-gene signature predicts clinical prognosis and reveals immune relevancy in clear cell renal cell carcinoma. BMC Cancer. 2021;21(1):1–14. https://doi.org/10.1186/s12885-021-08559-0.
Tang W, Xu F, Zhao M, Zhang S. Ferroptosis regulators, especially SQLE, play an important role in prognosis, progression and immune environment of breast cancer. BMC Cancer. 2021;21(1):1–19. https://doi.org/10.1186/s12885-021-08892-4.
Liu C, Liu Y, Yu Y, Zhao Y, Yu A. Comprehensive analysis of ferroptosis-related genes and prognosis of cutaneous melanoma. BMC Med Genomics. 2022;15(1):1–17. https://doi.org/10.1186/s12920-022-01194-z.
Xu Z, Xie Y, Mao Y, Huang J, Mei X, Song J et al. Ferroptosis-related gene signature predicts the prognosis of skin cutaneous melanoma and response to immunotherapy. Front Genet. 2021;12. doi:https://doi.org/10.3389/fgene.2021.758981.
Nguyen J, Tirla A, Rivera-Fuentes P. Disruption of mitochondrial redox homeostasis by enzymatic activation of a trialkylphosphine probe. Org Biomol Chem. 2021;19(12):2681–7. https://doi.org/10.1039/d0ob02259d.
Wang Z, Li M, Liu Y, Qiao Z, Bai T, Yang L, et al. Dihydroartemisinin triggers ferroptosis in primary liver cancer cells by promoting and unfolded protein response-induced upregulation of CHAC1 expression. Oncol Rep. 2021;46(5):1–14. https://doi.org/10.3892/or.2021.8191.
Li D, Liu S, Xu J, Chen L, Xu C, Chen F, et al. Ferroptosis-related gene CHAC1 is a valid indicator for the poor prognosis of kidney renal clear cell carcinoma. J Cell Mol Med. 2021;25(7):3610–21.
Acknowledgements
The authors are thankful for the help from the faculty and technical staff of the Department of Histopathology, PGIMER Chandigarh, India. We would like to acknowledge ICMR, New Delhi for providing a Senior research fellowship (SRF) to V.M. We thank Prof. Anjana Munshi for her administrative help to V.M.
Funding
The study received funding from the Department of Science and Technology-Science and Engineering Research Board (DST-SERB), India (Grant No. EMR/2015/000761 & EEQ/2017/000794).
Author information
Authors and Affiliations
Contributions
V.M and H.C designed the study. V.M and H.C. performed the experimental work. P.S contributed to experimental work in part. V.M and H.C drafted the figures and prepared the manuscript. All the authors analyzed the data and reviewed the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there are no conflicts of interest.
Availability of Data and materials
The public database used in this study includes Kaplan–Meier Plotter for CHAC1 transcripts (https://kmplot.com/analysis/index.php?p=service), cBioportal for correlation analysis between Ki67 and CHAC1 (https://www.cbioportal.org/).
Ethical Approval and Consent to participate.
No formal ethics approval was required for the study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Fig. S1
Specificity and validation of CHAC1 antibody. (a) Breast Cancer tissue stained with CHAC1 antibody, where CHAC1 antibody showed immunoreactivity. (b) Breast Cancer tissue showed no immunoreactivity towards CHAC1 by the antibody that was incubated with immunoprecipitated CHAC1 from the cell lysate. (c) CHAC1 Immunoblot after siRNA transfection at indicated doses in MCF7 cells. (d) Densitometric analysis of CHAC1 expression in MCF7 cells. (e) CHAC1 Immunoblot after siRNA transfection at indicated doses in SKBR3 cells. (f) Densitometric analysis of CHAC1 expression in SKBR3 cells. Fig. S2a Representative photomicrographs of the malignant cores of low (Ki67<25%) and high (Ki67>25%) index. Fig. S2b Representative photomicrographs of the malignant cores of negative and positive lymph node metastasis. Fig. S3 Correlation analysis of CHAC1 expression shows a positive correlation with Ki67 from cBioportal using (a) microarray and (b) RSEM data (TCGA, Firehose legacy dataset; n=1108). Fig. S4. Analysis of CHAC1 expression with tumor stage. Fig. S5. Distant metastasis-free survival plot as per CHAC1 expression in (a) metastatic (n=889, HR=1.62, logrank p=0.00016) vs (b) non-metastatic patients (n=1309, HR=1.5, logrank p=0.0013). (PDF 205 KB).
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Mehta, V., Suman, P. & Chander, H. High levels of unfolded protein response component CHAC1 associates with cancer progression signatures in malignant breast cancer tissues. Clin Transl Oncol 24, 2351–2365 (2022). https://doi.org/10.1007/s12094-022-02889-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12094-022-02889-6