Abstract
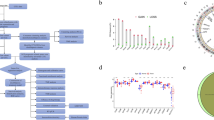
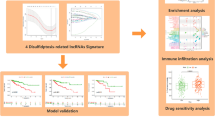
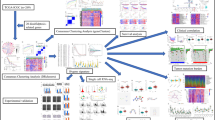
Pancreatic cancer is a lethal, extremely aggressive gastrointestinal tumor with a poor prognosis and limited treatment alternatives. Disulfidptosis is a newly defined type of cell death with potential influence on cancer. Research on the association between disulfidptosis and pancreatic cancer is scarce. The expression data of disulfidptosis-related genes were downloaded from The Cancer Genome Atlas-Pancreatic Adenocarcinoma (TCGA). Disulfidptosis-related lncRNA signature (DRLS) was developed through the Cox and the least absolute shrinkage and selection operator (LASSO) analysis. Differences in enrichment functions, mutational landscape, immune microenvironment, and predicted therapeutic efficacy between high- and low-risk groups were assessed. Consensus clustering analysis was applied to identify the DRLS-related subtypes. Among 98 disulfidptosis-related lncRNAs, 5 lncRNAs were screened thus constructing a prognostic DRLS. DRLS showed high predictive accuracy and was an independent prognostic factor for pancreatic cancer. According to the risk scores calculated from the signature, samples were categorized into high- and low- risk groups. Overall, low-risk patients had a better prognosis, lower mutational occurrences, higher immune cell infiltration and more sensitivity to anti-tumor agents. The DRLS performed well in predicting prognosis and revealed intimate correlation with biological function, mutation status and immune infiltration landscape of pancreatic cancer, providing some insights for future research on the relationship between disulfidptosis and pancreatic cancer.









Similar content being viewed by others
Data Availability
Publicly available datasets can be found here: (https://portal.gdc.cancer.gov/). The datasets generated and analyzed in this study are available on request from the corresponding authors.
References
Siegel, R. L., Miller, K. D., Wagle, N. S., & Jemal, A. (2023). Cancer statistics, 2023. C Ca: A Cancer Journal for Clinicians, 73(1), 17–48.
Diaz-Cano, S. J. (2012). Tumor heterogeneity: Mechanisms and bases for a reliable application of molecular marker design. International Journal of Molecular Sciences, 13(2), 1951–2011.
Wang, D., Liu, B., & Zhang, Z. (2023). Accelerating the understanding of cancer biology through the lens of genomics. Cell, 186(8), 1755–1771.
Vitale, I., Shema, E., Loi, S., & Galluzzi, L. (2021). Intratumoral heterogeneity in cancer progression and response to immunotherapy. Nature Medicine, 27(2), 212–224.
Muller, S., Raulefs, S., Bruns, P., Afonso-Grunz, F., Plotner, A., Thermann, R., Jager, C., Schlitter, A. M., Kong, B., Regel, I., Roth, W. K., Rotter, B., Hoffmeier, K., Kahl, G., Koch, I., Theis, F. J., Kleeff, J., Winter, P., & Michalski, C. W. (2015). Next-generation sequencing reveals novel differentially regulated mRNAs, lncRNAs, miRNAs, sdRNAs and a piRNA in pancreatic cancer. Molecular Cancer, 14, 94.
Kisling, S. G., Atri, P., Shah, A., Cox, J. L., Sharma, S., Smith, L. M., Ghersi, D., & Batra, S. K. (2023). A Novel HOXA10-Associated 5 gene-based prognostic signature for stratification of short-term survivors of pancreatic ductal adenocarcinoma. Clinical Cancer Research. https://doi.org/10.1158/1078-0432.CCR-23-0825
Chen, J., Liu, Z., Wu, Z., Li, W., & Tan, X. (2023). Identification of a chemoresistance-related prognostic gene signature by comprehensive analysis and experimental validation in pancreatic cancer. Frontiers in Oncology, 13, 1132424.
Chen, L., Zhang, L., He, H., Shao, F., Gao, Y., & He, J. (2023). Systemic analyses of cuproptosis-related lncRNAs in pancreatic adenocarcinoma, with a focus on the molecular mechanism of LINC00853. International Journal Of Molecular Sciences, 24(9), 7923.
Ren, X., Cui, H., Wu, J., Zhou, R., Wang, N., Liu, D., Xie, X., Zhang, H., Liu, D., Ma, X., Dang, C., Kang, H., & Lin, S. (2022). Identification of a combined apoptosis and hypoxia gene signature for predicting prognosis and immune infiltration in breast cancer. Cancer Medicine, 11(20), 3886–3901.
Tang, R., Wu, Z., Rong, Z., Xu, J., Wang, W., Zhang, B., Yu, X., & Shi, S. (2022). Ferroptosis-related lncRNA pairs to predict the clinical outcome and molecular characteristics of pancreatic ductal adenocarcinoma. Briefings In Bioinformatics, 23(1), bbab388.
Yao, H. F., Xu, D. P., Zheng, J. H., Xu, Y., Jia, Q. Y., Zhu, Y. H., Yang, J., He, R. Z., Ma, D., Yang, M. W., Fu, X. L., Liu, D. J., Huo, Y. M., Yang, J. Y., & Zhang, J. F. (2023). Analysis of cuproptosis-related lncRNA signature for predicting prognosis and tumor immune microenvironment in pancreatic cancer. Apoptosis, 28(7–8), 1090–1112.
Liu, X., Nie, L., Zhang, Y., Yan, Y., Wang, C., Colic, M., Olszewski, K., Horbath, A., Chen, X., Lei, G., Mao, C., Wu, S., Zhuang, L., Poyurovsky, M. V., James You, M., Hart, T., Billadeau, D. D., Chen, J., & Gan, B. (2023). Actin cytoskeleton vulnerability to disulfide stress mediates disulfidptosis. Nature Cell Biology, 25(3), 404–414.
Zhao, S., Wang, L., Ding, W., Ye, B., Cheng, C., Shao, J., Liu, J., & Zhou, H. (2023). Crosstalk of disulfidptosis-related subtypes, establishment of a prognostic signature and immune infiltration characteristics in bladder cancer based on a machine learning survival framework. Frontiers in Endocrinology (Lausanne), 14, 1180404.
Zhang, D., Zhang, X., Liu, Z., Han, T., Zhao, K., Xu, X., Zhang, X., Ren, X., & Qin, C. (2023). An integrative multi-omics analysis based on disulfidptosis-related prognostic signature and distinct subtypes of clear cell renal cell carcinoma. Frontiers in Oncology, 13, 1207068.
Yang, L., Zhang, W., & Yan, Y. (2023). Identification and characterization of a novel molecular classification based on disulfidptosis-related genes to predict prognosis and immunotherapy efficacy in hepatocellular carcinoma. Aging (Albany NY), 15(13), 6135.
Sha, D., Jin, Z., Budczies, J., Kluck, K., Stenzinger, A., & Sinicrope, F. A. (2020). Tumor Mutational Burden as a predictive biomarker in solid tumors. Cancer Discovery, 10(12), 1808–1825.
Barthel, S., Falcomata, C., Rad, R., Theis, F. J., & Saur, D. (2023). Single-cell profiling to explore pancreatic cancer heterogeneity, plasticity and response to therapy. Nature Cancer, 4(4), 454–467.
Zhang, A., Miao, K., Sun, H., & Deng, C. X. (2022). Tumor heterogeneity reshapes the tumor microenvironment to influence drug resistance. International Journal of Biological Sciences, 18(7), 3019–3033.
Zhao, K., Li, X., Shi, Y., Lu, Y., Qiu, P., Deng, Z., Yao, W., & Wang, J. (2022). A comprehensive analysis of pyroptosis-related lncRNAs signature Associated with Prognosis and Tumor Immune Microenvironment of pancreatic adenocarcinoma. Frontiers in Genetics, 13, 899496.
Chi, H., Peng, G., Wang, R., Yang, F., Xie, X., Zhang, J., Xu, K., Gu, T., Yang, X., & Tian, G. (2022). Cuprotosis programmed-cell-death-related lncrna signature predicts prognosis and immune landscape in PAAD patients. Cells, 11(21), 3436.
Liu, D., Zhao, J., Wang, H., Li, H., Li, Y., & Qin, W. (2020). Long intergenic non-protein coding RNA 519 promotes the biological activities of tongue squamous cell carcinoma by sponging microRNA-876-3p and consequently upregulating MACC1. Onco Targets and Therapy, 13, 11975–11990.
Ye, P., Lv, X., Aizemaiti, R., Cheng, J., Xia, P., & Di, M. (2020). H3K27ac-activated LINC00519 promotes lung squamous cell carcinoma progression by targeting miR-450b-5p/miR-515-5p/YAP1 axis. Cell Proliferation, 53(5), e12797.
Zhou, X. Y., Dai, H. Y., Zhang, H., Zhu, J. L., & Hu, H. (2022). Ferroptosis-related lncRNA for the establishment of Novel Prognostic signature and therapeutic response prediction to endometrial carcinoma. Biomed Research International, 2022, 2056913.
Jiang, K., Wu, L., Yin, X., Tang, Q., Yin, J., Zhou, Z., Yu, H., & Yan, S. (2022). Prognostic implications of necroptosis-related long noncoding RNA signatures in muscle-invasive bladder cancer. Frontiers in Genetics, 13, 1036098.
Priyanka, P., Sharma, M., Das, S., & Saxena, S. (2022). E2F1-induced lncRNA, EMSLR regulates lncRNA LncPRESS1. Scientific Reports, 12(1), 2548.
Propper, D. J., & Balkwill, F. R. (2022). Harnessing cytokines and chemokines for cancer therapy. Nature Reviews. Clinical Oncology, 19(4), 237–253.
Porembka, M. R., Mitchem, J. B., Belt, B. A., Hsieh, C. S., Lee, H. M., Herndon, J., Gillanders, W. E., Linehan, D. C., & Goedegebuure, P. (2012). Pancreatic adenocarcinoma induces bone marrow mobilization of myeloid-derived suppressor cells which promote primary tumor growth. Cancer Immunology, Immunotherapy, 61(9), 1373–1385.
Hamarsheh, S., Gross, O., Brummer, T., & Zeiser, R. (2020). Immune modulatory effects of oncogenic KRAS in cancer. Nature Communications, 11(1), 5439.
Casey, S. C., Tong, L., Li, Y., Do, R., Walz, S., Fitzgerald, K. N., Gouw, A. M., Baylot, V., Gutgemann, I., Eilers, M., & Felsher, D. W. (2016). MYC regulates the antitumor immune response through CD47 and PD-L1. Science, 352(6282), 227–231.
Nia, H. T., Munn, L. L., & Jain, R. K. (2020). Physical traits of cancer. Science, 370(6516), eaaz0868.
Hartmann, N., Giese, N. A., Giese, T., Poschke, I., Offringa, R., Werner, J., & Ryschich, E. (2014). Prevailing role of contact guidance in intrastromal T-cell trapping in human pancreatic cancer. Clinical Cancer Research, 20(13), 3422–3433.
Rothlin, C. V., Hille, T. D., & Ghosh, S. (2021). Determining the effector response to cell death. Nature Reviews Immunology, 21(5), 292–304.
Tang, R., Xu, J., Zhang, B., Liu, J., Liang, C., Hua, J., Meng, Q., Yu, X., & Shi, S. (2020). Ferroptosis, necroptosis, and pyroptosis in anticancer immunity. Journal of Hematology & Oncology, 13(1), 110.
Kindler, H. L., Yoo, H. K., Hettle, R., Cui, K. Y., Joo, S., Locker, G. Y., & Golan, T. (2023). Patient-centered outcomes in the POLO study of active maintenance olaparib for germline BRCA-mutated metastatic pancreatic cancer. Cancer, 129(9), 1411–1418.
Strickler, J. H., Satake, H., George, T. J., Yaeger, R., Hollebecque, A., Garrido-Laguna, I., Schuler, M., Burns, T. F., Coveler, A. L., Falchook, G. S., Vincent, M., Sunakawa, Y., Dahan, L., Bajor, D., Rha, S. Y., Lemech, C., Juric, D., Rehn, M., Ngarmchamnanrith, G., Jafarinasabian, P., Tran, Q., & Hong, D. S. (2023). Sotorasib in KRAS p.G12C-Mutated Advanced Pancreatic Cancer. New England Journal of Medicine, 388(1), 33–43.
Grierson, P. M., Tan, B., Pedersen, K. S., Park, H., Suresh, R., Amin, M. A., Trikalinos, N. A., Knoerzer, D., Kreider, B., Reddy, A., Liu, J., Der, C. J., Wang-Gillam, A., & Lim, K. H. (2023). Phase ib study of Ulixertinib Plus Gemcitabine and Nab-Paclitaxel in patients with metastatic pancreatic adenocarcinoma. The Oncologist, 28(2), e115–e123.
Goodwin, C. M., Waters, A. M., Klomp, J. E., Javaid, S., Bryant, K. L., Stalnecker, C. A., Drizyte-Miller, K., Papke, B., Yang, R., Amparo, A. M., Ozkan-Dagliyan, I., Baldelli, E., Calvert, V., Pierobon, M., Sorrentino, J. A., Beelen, A. P., Bublitz, N., Luthen, M., Wood, K. C., Petricoin, E. F., Sers, C., McRee, A. J., Cox, A. D., & Der, C. J. (2023). Combination therapies with CDK4/6 inhibitors to treat KRAS-Mutant Pancreatic Cancer. Cancer Research, 83(1), 141–157.
Javle, M., Shacham-Shmueli, E., Xiao, L., Varadhachary, G., Halpern, N., Fogelman, D., Boursi, B., Uruba, S., Margalit, O., Wolff, R. A., & Golan, T. (2021). Olaparib monotherapy for previously treated pancreatic cancer with DNA damage repair genetic alterations other than germline BRCA variants: Findings from 2 phase 2 Nonrandomized clinical trials. JAMA Oncology, 7(5), 693–699.
Acknowledgements
We appreciate all the participants and researchers for their contributions and commitment to this study.
Funding
This work was jointly supported by the National Natural Science Foundation of China (No. 82173178, U21A20374), Shanghai Municipal Science and Technology Major Project (21JC1401500), Scientific Innovation Project of Shanghai Education Committee (2019-01-07-00-07-E00057), Clinical Research Plan of Shanghai Hospital Development Center (SHDC2020CR1006A), and Xuhui District Artificial Intelligence Medical Hospital Cooperation Project (2021-011).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare that they have no competing interests.
Ethical Approval
Not applicable.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Xing, F., Qin, Y., Xu, J. et al. Construction of a Novel Disulfidptosis-Related lncRNA Prognostic Signature in Pancreatic Cancer. Mol Biotechnol (2023). https://doi.org/10.1007/s12033-023-00875-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12033-023-00875-z