Abstract
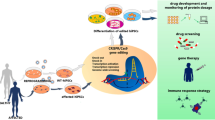
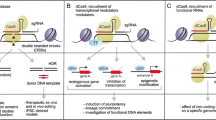
The newly developed TALENs and emerging CRISPR/Cas9 have spurred interests in the field of genome engineering because of their ease of customization and high-efficient site-specific cleavages. Although these novel technologies have been successfully used in many types of cells, it is of great importance to apply them in human-derived cells to further observe and evaluate their clinical potentials in gene therapy. Here, we review the working mechanism of TALEN and CRISPR/Cas9, their effectiveness and specificity in human cells, and current methods to enhance efficiency and reduce off-target effects. Besides, CCR5 gene was chosen as a target example to illustrate their clinical potentials. Finally, some questions are raised for future research and for researchers to consider when making a proper choice bases on different purposes.


Similar content being viewed by others
References
Verma, I. M., & Weitzman, M. D. (2005). Gene therapy: Twenty-first century medicine. Annual Review of Biochemistry, 74, 711–738.
Perez, E. E., Wang, J., Miller, J. C., Jouvenot, Y., Kim, K. A., Liu, O., et al. (2008). Establishment of HIV-1 resistance in CD4+T cells by genome editing using zinc-finger nucleases. Nature Biotechnology, 26, 808–816.
Li, L., Krymskaya, L., Wang, J., Henley, J., Rao, A., Cao, L.-F., et al. (2013). Genomic editing of the HIV-1 coreceptor CCR5 in adult hematopoietic stem and progenitor cells using zinc finger nucleases. Molecular Therapy, 21, 1259–1269.
Mussolino, C., Morbitzer, R., Lütge, F., Dannemann, N., Lahaye, T., & Cathomen, T. (2011). A novel TALE nuclease scaffold enables high genome editing activity in combination with low toxicity. Nucleic Acids Research, 39, 9283–9293.
Ding, Q., Lee, Y.-K., Schaefer, E. A., Peters, D. T., Veres, A., Kim, K., et al. (2012). A TALEN genome-editing system for generating human stem cell-based disease models. Cell Stem Cell, 12, 238–251.
Takata, M., Sasaki, M. S., Sonoda, E., Morrison, C., Hashimoto, M., Utsumi, H., et al. (1998). Homologous recombination and non-homologous end-joining pathways of DNA double-strand break repair have overlapping roles in the maintenance of chromosomal integrity in vertebrate cells. The EMBO Journal, 17, 5497–5508.
Christian, M., Cermak, T., Doyle, E. L., Schmidt, C., Zhang, F., Hummel, A., et al. (2010). Targeting DNA double-strand breaks with TAL effector nucleases. Genetics, 186, 757–761.
Miller, J. C., Tan, S., Qiao, G., Barlow, K. A., Wang, J., Xia, D. F., et al. (2010). A TALE nuclease architecture for efficient genome editing. Nature Biotechnology, 29, 143–148.
Hockemeyer, D., Wang, H., Kiani, S., Lai, C. S., Gao, Q., Cassady, J. P., et al. (2011). Genetic engineering of human pluripotent cells using TALE nucleases. Nature Biotechnology, 29, 731–734.
Li, T., Huang, S., Jiang, W. Z., Wright, D., Spalding, M. H., Weeks, D. P., et al. (2011). TAL nucleases (TALNs): Hybrid proteins composed of TAL effectors and FokI DNA-cleavage domain. Nucleic Acids Research, 39, 359–372.
Santiago, Y., Chan, E., Liu, P.-Q., Orlando, S., Zhang, L., Urnov, F. D., et al. (2008). Targeted gene knockout in mammalian cells by using engineered zinc-finger nucleases. Proceedings of the National Academy of Sciences, 105, 5809–5814.
Bogdanove, A. J., & Voytas, D. F. (2011). TAL effectors: Customizable proteins for DNA targeting. Science, 333, 1843–1846.
Moehle, E. A., Rock, J. M., Lee, Y.-L., Jouvenot, Y., DeKelver, R. C., Gregory, P. D., et al. (2007). Targeted gene addition into a specified location in the human genome using designed zinc finger nucleases. Proceedings of the National Academy of Sciences, 104, 3055–3060.
Gaj, T., Gersbach, C. A., & Barbas III, C. F. (2013). ZFN, TALEN, and CRISPR/Cas-based methods for genome engineering. Trends in Biotechnology, 31, 397–405.
Boch, J., Scholze, H., Schornack, S., Landgraf, A., Hahn, S., Kay, S., et al. (2009). Breaking the code of DNA binding specificity of TAL-type III effectors. Science, 326, 1509–1512.
Bogdanove, A. J., Schornack, S., & Lahaye, T. (2010). TAL effectors: Finding plant genes for disease and defense. Current Opinion in Plant Biology, 13, 394–401.
Moscou, M. J., & Bogdanove, A. J. (2009). A simple cipher governs DNA recognition by TAL effectors. Science, 326, 1501.
Ishino, Y., Shinagawa, H., Makino, K., Amemura, M., & Nakata, A. (1987). Nucleotide sequence of the iap gene, responsible for alkaline phosphatase isozyme conversion in Escherichia coli, and identification of the gene product. Journal of Bacteriology, 169, 5429–5433.
Horvath, P., & Barrangou, R. (2010). CRISPR/Cas, the immune system of bacteria and archaea. Science, 327, 167–170.
Bhaya, D., Davison, M., & Barrangou, R. (2011). CRISPR-Cas systems in bacteria and archaea: Versatile small RNAs for adaptive defense and regulation. Annual Review of Genetics, 45, 273–297.
Hwang, W. Y., Fu, Y., Reyon, D., Maeder, M. L., Tsai, S. Q., Sander, J. D., et al. (2013). Efficient genome editing in zebrafish using a CRISPR-Cas system. Nature Biotechnology, 31, 227–229.
Jiang, W., Bikard, D., Cox, D., Zhang, F., & Marraffini, L. A. (2013). RNA-guided editing of bacterial genomes using CRISPR-Cas systems. Nature Biotechnology, 31, 233–239.
Mali, P., Yang, L., Esvelt, K. M., Aach, J., Guell, M., DiCarlo, J. E., et al. (2013). RNA-guided human genome engineering via Cas9. Science, 339, 823–826.
Jinek, M., Chylinski, K., Fonfara, I., Hauer, M., Doudna, J. A., & Charpentier, E. (2012). A programmable dual-RNA–guided DNA endonuclease in adaptive bacterial immunity. Science, 337, 816–821.
Qi, L. S., Larson, M. H., Gilbert, L. A., Doudna, J. A., Weissman, J. S., Arkin, A. P., et al. (2013). Repurposing CRISPR as an RNA-guided platform for sequence-specific control of gene expression. Cell, 152, 1173–1183.
Gilbert, L. A., Larson, M. H., Morsut, L., Liu, Z., Brar, G. A., Torres, S. E., et al. (2013). CRISPR-mediated modular RNA-guided regulation of transcription in eukaryotes. Cell, 154, 442–451.
Stroud, D. A., Formosa, L. E., Wijeyeratne, X. W., Nguyen, T. N., & Ryan, M. T. (2013). Gene knockout using transcription activator-like effector nucleases (TALENs) reveals that human NDUFA9 protein is essential for stabilizing the junction between membrane and matrix arms of complex I. Journal of Biological Chemistry, 288, 1685–1690.
Hu, R., Wallace, J., Dahlem, T. J., Grunwald, D. J., & O’Connell, R. M. (2013). Targeting human MicroRNA genes using engineered Tal-effector nucleases (TALENs). PLoS ONE, 8, e63074.
Jinek, M., East, A., Cheng, A., Lin, S., Ma, E., & Doudna, J. (2013). RNA-programmed genome editing in human cells. Elife, 2, e00471.
Cho, S. W., Kim, S., Kim, J. M., & Kim, J.-S. (2013). Targeted genome engineering in human cells with the Cas9 RNA-guided endonuclease. Nature Biotechnology, 31, 230–232.
Cong, L., Ran, F. A., Cox, D., Lin, S., Barretto, R., Habib, N., et al. (2013). Multiplex genome engineering using CRISPR/Cas systems. Science, 339, 819–823.
Fu, Y., Foden, J. A., Khayter, C., Maeder, M. L., Reyon, D., Joung, J. K., et al. (2013). High-frequency off-target mutagenesis induced by CRISPR-Cas nucleases in human cells. Nature Biotechnology, 31, 822–826.
Kim, Y., Kweon, J., Kim, A., Chon, J. K., Yoo, J. Y., Kim, H. J., et al. (2013). A library of TAL effector nucleases spanning the human genome. Nature Biotechnology, 31, 251–258.
Ding, Q., Regan, S. N., Xia, Y., Oostrom, L. A., Cowan, C. A., & Musunuru, K. (2013). Enhanced efficiency of human pluripotent stem cell genome editing through replacing TALENs with CRISPRs. Cell Stem Cell, 12, 393–394.
Wei, C., Liu, J., Yu, Z., Zhang, B., Gao, G., & Jiao, R. (2013). TALEN or Cas9–rapid, efficient and specific choices for genome modifications. Journal of Genetics and Genomics, 40, 281–289.
Yang, L., Guell, M., Byrne, S., Yang, J. L., De Los Angeles, A., Mali, P., et al. (2013). Optimization of scarless human stem cell genome editing. Nucleic Acids Research, 41, 9049–9061.
Streubel, J., Blücher, C., Landgraf, A., & Boch, J. (2012). TAL effector RVD specificities and efficiencies. Nature Biotechnology, 30, 593–595.
Sakuma, T., Ochiai, H., Kaneko, T., Mashimo, T., Tokumasu, D., Sakane, Y., et al. (2013). Repeating pattern of non-RVD variations in DNA-binding modules enhances TALEN activity. Scientific Reports, 3, 3379.
Wang, T., Wei, J. J., Sabatini, D. M., & Lander, E. S. (2014). Genetic screens in human cells using the CRISPR-Cas9 system. Science, 343, 80–84.
Kim, E., Kim, S., Kim, D. H., Choi, B.-S., Choi, I.-Y., & Kim, J.-S. (2012). Precision genome engineering with programmable DNA-nicking enzymes. Genome Research, 22, 1327–1333.
Hsu, P. D., Scott, D. A., Weinstein, J. A., Ran, F. A., Konermann, S., Agarwala, V., et al. (2013). DNA targeting specificity of RNA-guided Cas9 nucleases. Nature Biotechnology, 31, 827–832.
Ran, F. A., Hsu, P. D., Wright, J., Agarwala, V., Scott, D. A., & Zhang, F. (2013). Genome engineering using the CRISPR-Cas9 system. Nature Protocols, 8, 2281–2308.
Cho, S. W., Kim, S., Kim, Y., Kweon, J., Kim, H. S., Bae, S., et al. (2013). Analysis of off-target effects of CRISPR/Cas-derived RNA-guided endonucleases and nickases. Genome Research, 24, 132–141.
Ran, F., Hsu, P. D., Lin, C.-Y., Gootenberg, J. S., Konermann, S., Trevino, A. E., et al. (2013). Double nicking by RNA-guided CRISPR Cas9 for enhanced genome editing specificity. Cell, 154, 1380–1389.
Pattanayak, V., Lin, S., Guilinger, J. P., Ma, E., Doudna, J. A., & Liu, D. R. (2013). High-throughput profiling of off-target DNA cleavage reveals RNA-programmed Cas9 nuclease specificity. Nature Biotechnology, 31, 839–843.
Fu, Y., Sander, J. D., Reyon, D., Cascio, V. M., & Joung, J. K. (2014). Improving CRISPR-Cas nuclease specificity using truncated guide RNAs. Nature Biotechnology, 32, 279–284.
Smith, A. M., Takeuchi, R., Pellenz, S., Davis, L., Maizels, N., Monnat, R. J., et al. (2009). Generation of a nicking enzyme that stimulates site-specific gene conversion from the I-AniI LAGLIDADG homing endonuclease. Proceedings of the National Academy of Sciences, 106, 5099–5104.
Cade, L., Reyon, D., Hwang, W. Y., Tsai, S. Q., Patel, S., Khayter, C., et al. (2012). Highly efficient generation of heritable zebrafish gene mutations using homo-and heterodimeric TALENs. Nucleic Acids Research, 40, 8001–8010.
Mali, P., Aach, J., Stranges, P. B., Esvelt, K. M., Moosburner, M., Kosuri, S., et al. (2013). CAS9 transcriptional activators for target specificity screening and paired nickases for cooperative genome engineering. Nature Biotechnology, 31, 833–838.
Pan, Y., Xiao, L., Li, A. S., Zhang, X., Sirois, P., Zhang, J., et al. (2013). Biological and biomedical applications of engineered nucleases. Molecular Biotechnology, 55, 54–62.
Bloom, K., Ely, A., Mussolino, C., Cathomen, T., & Arbuthnot, P. (2013). Inactivation of hepatitis B virus replication in cultured cells and in vivo with engineered transcription activator-like effector nucleases. Molecular Therapy, 21, 1889–1897.
Ousterout, D. G., Perez-Pinera, P., Thakore, P. I., Kabadi, A. M., Brown, M. T., Qin, X., et al. (2013). Reading frame correction by targeted genome editing restores dystrophin expression in cells from duchenne muscular dystrophy patients. Molecular Therapy, 21, 1718–1726.
Osborn, M. J., Starker, C. G., McElroy, A. N., Webber, B. R., Riddle, M. J., Xia, L., et al. (2013). TALEN-based gene correction for epidermolysis bullosa. Molecular Therapy, 21, 1151–1159.
Xu, L., Zhao, P., Mariano, A., Han, R. (2013). Targeted myostatin gene editing in multiple mammalian species directed by a single pair of TALE nucleases. Molecular Therapy—Nucleic Acids, 2, 112.
Bacman, S. R., Williams, S. L., Pinto, M., Peralta, S., & Moraes, C. T. (2013). Specific elimination of mutant mitochondrial genomes in patient-derived cells by mitoTALENs. Nature Medicine, 19, 1111–1113.
Schwank, G., Koo, B.-K., Sasselli, V., Dekkers, J. F., Heo, I., Demircan, T., et al. (2013). Functional repair of CFTR by CRISPR/Cas9 in intestinal stem cell organoids of cystic fibrosis patients. Cell Stem Cell, 13, 653–658.
Choe, H., Farzan, M., Sun, Y., Sullivan, N., Rollins, B., Ponath, P. D., et al. (1996). The β-chemokine receptors CCR3 and CCR5 facilitate infection by primary HIV-1 isolates. Cell, 85, 1135–1148.
Liu, R., Paxton, W. A., Choe, S., Ceradini, D., Martin, S. R., Horuk, R., et al. (1996). Homozygous defect in HIV-1 coreceptor accounts for resistance of some multiply-exposed individuals to HIV-1 infection. Cell, 86, 367–377.
Kim, H. J., Lee, H. J., Kim, H., Cho, S. W., & Kim, J.-S. (2009). Targeted genome editing in human cells with zinc finger nucleases constructed via modular assembly. Genome Research, 19, 1279–1288.
Sakuma, T., Hosoi, S., Woltjen, K., Suzuki, K. I., Kashiwagi, K., Wada, H., et al. (2013). Efficient TALEN construction and evaluation methods for human cell and animal applications. Genes to Cells, 18, 315–326.
Doench, G., & Zhang, F. (2014). Genome-scale CRISPR-Cas9 knockout screening in human cells. Science, 343, 84–87.
Acknowledgments
This study is supported by The “863 Projects” of Ministry of Science and Technology of PR China (No. 2013AA020103); National Science and Technology Major Project (No. 2013ZX1001003).
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Niu, J., Zhang, B. & Chen, H. Applications of TALENs and CRISPR/Cas9 in Human Cells and Their Potentials for Gene Therapy. Mol Biotechnol 56, 681–688 (2014). https://doi.org/10.1007/s12033-014-9771-z
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12033-014-9771-z