Abstract
The development of engineered nucleases is the fruit of a new technological approach developed in the last two decades which has led to significant benefits on genome engineering, particularly on gene therapy. These applications enable efficient and specific genetic modifications via the induction of a double-strand break (DSB) in a specific genomic target sequence, followed by the homology-directed repair (HDR) or non-homologous end joining (NHEJ) pathways. In addition to the application on gene modification in cells and intact organisms, a number of recent papers have reported that this gene editing technology can be applied effectively to human diseases. With the promising data obtained using engineered endonucleases in gene therapy, it appears reasonable to expect that more diseases could be treated and even be cured in this new era of individualized medicine. This paper first brief introduces the development of engineered nucleases with a special emphasis on zinc-finger nucleases (ZFNs) and transcription activator-like effector (TALE) nucleases (TALENs), and then takes CCR5-based gene therapy as an example to discuss the therapeutic applications of engineered nucleases.
Similar content being viewed by others
Introduction
Gene therapy is a genetic intervention used for the prevention or treatment of diseases by targeting selected genes with specific nucleotides. It includes both traditional forward gene therapy and the recently developed reversed gene therapy. The major purpose of traditional gene therapy is to normalize an abnormal gene, or in other words it is to restore the function a dysfunctional gene. As an opposite strategy, reversed gene therapy [1] aims at suppressing or abolishing the function of a gene to balance a complicated regulatory system or at depleting cellular molecules essential for a disease mechanism as in pathogen infections.
In the past three decades, few diseases and few cases have been cured solely by traditional forward gene therapy and two major obstacles contributed to hampering its clinical output: the selection of gene targets and the transgene delivery system.
The technological development of several engineered nucleases may help to overcome the aforementioned difficulties and pave the way to gene therapy. Among the engineered nucleases, zinc-finger nucleases (ZFNs), transcription activator-like effector nucleases (TALENs), and meganucleases have been successfully employed for both in vitro and ex vivo gene modifications. Based on these milestone events and their great potential, gene editing technology with ZFNs and TALENs was recognized as the Technology of the Year by the journal “Nature-Methods.”
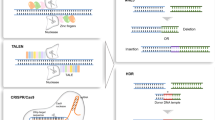
Zinc-finger nucleases (ZFNs) include a DNA binding domain of a serially repeated zinc fingers and a nuclease domain of the FokI restriction enzyme. Each zinc finger can precisely recognize and combine 3 nucleotides. ZFNs function as dimers, in which each monomer of ZFNs usually contains 3–4 or even more repeated elements to specifically digest one target per genome (Fig. 1). As the ZFNs, TALENs are also made of a number of DNA binding modules and the nuclease domain from FokI restriction enzyme. Each protein module typically contains 34 amino acids, and the 12th and 13th residues are the key site of the target identification, known as the repeated variable di-residues (RVDs) [2–5]. Currently, there are seven recognized units decoded and four commonly used identification units: NI identifies A, HD identifies C, NG identifies T, and NN identifies G or A (Fig. 2). According to the identity or similarity of the 12th amino acid among these units, one can easily know that the nucleotide identification is crucially performed by the 13th amino acid based on a common sense: the exclusive difference is located at the site of the 13th amino acid but it is almost the same as the one located at the 12th amino acid. Thus, the identification to different nucleotides must be attributed to the different composition of the amino acid rather than to the identical or largely overlapping amino acids. This logical inference has recently been confirmed by X-ray microfluorescence (XRF) data. When a ZFNs or TALENs have repeated DNA binding elements in its DNA binding domain, they can then efficiently and precisely cleave specific DNA sequences and induce a double-strand break (DSB), followed by the homology-directed repair (HDR) or non-homologous end joining (NHEJ) pathways [6, 7].
Structures and applications of ZFNs and TALENs. ZFNs and TALENs recognize DNA targets through their DNA binding domains. While each zinc-finger protein module identifies three nucleotides, a TALE module only recognizes one nucleotide through its 13th amino acid. The cleavage domains of homodimers or heterodimers form an endonuclease that digests the target sequence and creates a double-strand break. Following the creation of double-strand break, gene editing efficiencies increase by two log scales as compared to those formed through spontaneous gene recombination, which greatly facilitates gene disruption, gene correction, or gene insertion
TALE identification of specific nucleotide. Each TALE module typically contains 34 amino acids and the 12th and 13th residues are the key sites of the target identification known as the repeated variable di-residues (RVDs). There are seven recognized units decoded and four commonly used identification units: NI identifies A, HD identifies C, NG identifies T, and NN identifies G or A
Biomedical Applications of ZFNs and TALENs
ZFNs and TALENs have been successfully employed in genetic manipulations both in vitro and in vivo. Several endogenous genes modified by ZFNs and TALENs are listed in Table 1.
Gene Disruption
Gene disruption is the simplest means of gene editing. By causing a frameshift mutation, the disruption or abolishment of selected gene functions can easily be obtained. Engineered nucleases-driven gene disruption has already been used for model organisms and a variety of mammalian cells as listed in Table 1.
Injection of mRNA encoding ZFNs into the early Drosophila embryo has been used to disrupt a gene in D. Melanogaster [8]. As for ZFNs-mediated gene disruption in zebrafish [9–12], four separate studies illustrated that up to 50 % germline disruptions at the targeted genes were achieved. The first published example of the application of engineered ZFNs to disrupt an endogenous locus in a mammalian cell is a knockout of the dihydrofolate reductase (DHFR) gene in Chinese hamster ovary (CHO) cells, in which the disruption frequencies are up to 15 % of alleles in the cell population [13]. In addition, ZFNs have been used to make double [14] and triple [15] locus gene disruptions in CHO and K562 cells.
In a study related to a rat model of severe combined immune deficiency (SCID), ZFNs was used to target an integrated reporter and two endogenous rat genes of immunoglobulin M (IgM) and Rab38 [16]. From 25 to 100 % of the rats developed from these ZFNs-treated one-cell embryos showed genotypes carrying disrupted target loci. These mutations are faithfully and efficiently transmitted through the germlines. In human and other eukaryotic cells, specific ZFNs deleted predetermined genomic DNA segments in the range of 15 to several-hundred base pairs (bp) at frequencies of 10−3–10−1[17]. TALENs can also efficiently disrupt the zebrafish genes as well as the endogenous human genes NTF3 and CCR5 at efficiencies of up to 25 % [18, 19].
Gene addition
The great potential of genetically modified endonucleases in genetic medicine, biotechnology, and molecular biology made this genomic approach a hot topic in biomedical research. Currently, more and more cases in gene therapy seem to be focusing on targeted gene additions. This approach consists of introducing DNA sequences into a predetermined genomic location. In order to make this strategy work, two essential properties should be considered [20]. First, the inserted DNA sequences must functionally integrate within the modified cells. Second, the normal function and gene regulation in cells harboring the predetermined genomic location should not be blocked, or in other words, no off-target effect is required following the gene modification or gene addition.
ZFNs-driven gene addition was achieved in plants in two different ways. A Ti plasmid harboring both the ZFNs and a donor DNA construct including a disease resistance gene cassette was delivered in tobacco cells. This strategy achieved targeted, homology-directed transgene integration in approximately 10 % of the ZFN cleavage site [21]. Alternatively, simultaneous expression of ZFNs and delivery of a simple heterologous donor molecule also led to precise targeted addition of an herbicide-tolerance gene at the intended locus in plants of the crop species Zea mays [22].
In addition to plant cells, this approach has been applied to gene addition in cells from various species including human cells. In mouse embryo stem (ES) cells, ZFNs were used to generate an isogenic panel carrying a defined series of alleles for an endogenous gene [23]. Using ZFNs directed against IL2RG and an extrachromosomal DNA donor carrying homology arms of 750 bp, transgenes positioned precisely in the ZFNs recognition site. In this study, ZFNs could drive the addition of an as long as 8 kb sequence carrying three distinct promoter-transcription units into an endogenous locus at a frequency of 6 % [24].
Gene addition has also been achieved in human ES and induced pluripotent stem (iPS) cells. The first demonstration of ZFN-mediated gene addition was done with a ZFN expression cassette and a donor construct delivered by an integrase-defective lentiviral vectors (IDLV) in human ES cells [25]. While a high ratio of gene addition (up to 50 %) was obtained in a panel of human cell lines, a relatively low efficiency was observed in human embryonic stem cells (5 % with gene addition). In human mesenchymal stromal cells (MSC), zinc-finger nuclease (ZFN)-mediated targeted addition of the erythropoietin (Epo) gene into CCR5 gene locus [26] would result in 30–40 % targeted gene addition and stable transgene expression in vivo. With a similar approach for the addition of genes into hESCs [27] using ZFNs, TALEN expression constructs and the corresponding donor plasmids were introduced into human embryonic stem cell (ESC); the TALEN-mediated targeting of PPP1R12C [28] resulted in a robust transgene expression in pluripotent as well as in differentiated cells.
Gene correction
According to the Human Genome Mutation Database (Table 2), in more than a hundred thousand mutations causing human diseases, 80–90 % is caused by the substitution, deletion, and insertion of one nucleotide. Therefore, the application of engineered nucleases for gene correction at a specific locus has been considered as a crucial target for gene therapy. As evidenced by its widespread use in experimental model organisms, plant, and mammalian cells, engineered nucleases-mediated gene corrections appear to have high therapeutic potential, particularly to point mutations and several other types of mutations with short sequences involved.
The gene correction based on zinc-finger nucleases in human cells was first reported in a X-linked SCID mutation in the IL2Rγ gene in which more than 18 % of gene-modified human cells were observed [29]. It is noteworthy that about 7 % of the cells acquired the desired genetic modification on both X chromosomes. High frequencies were also observed at this locus in human CD4+ T cell. The same approach was applied with relatively high efficiencies (1–50 %) to three additional human genes (VEGF-A, HoxB13, and CFTR) in transformed cells [30].
In a mouse model of hemophilia B [31], ZFNs co-delivered with an appropriately designed gene-targeting vector stimulated gene correction in vivo. As a consequence, the gene therapy treatment was sufficient to correct the prolonged clotting times in this mouse model of hemophilia B, and remained a persistent effect after the induced liver regeneration. In addition, gene corrections using ZFNs and TALENs have also been achieved in tobacco, fruit flies, Arabidopsis thaliana, and mouse ES cells for a variety of gene abnormalities [32–34].
Comparison Between the Enzymatic Properties and Potential Application Values of ZFNs and TALENs
TALENs have shown some advantages over ZFNs in genetic modifications, gene therapy, and selected other uses [35]. The base identification mechanisms of these enzymes are different. Each zinc finger can recognize three nucleotides whereas each TALE only recognizes one nucleotide. Compared with ZFNs, the cloning process of TALENs is easier, the specificity of recognized target sequences is higher, and their off-target effects are lower. Additionally, there will be less possible intellectual property disputes in the application of TALENs as compared to ZFNs. In Table 3, the characteristic features of ZFNs and TALENs are summarized.
The proper targets of TALENs used in gene therapy are single base mutations, small deletions, or small insertions. As reported in the Human Genome Mutation Database, these three types of mutations cover over 80 % of known monogenic diseases. Additionally, TALENs can be used in certain polygenic diseases such as essential hypertension or infectious diseases such as HIV. Interestingly, a recent study by Dr. JS Kim demonstrated that ZFNs are able to edit DNA over a range of kilobase-pairs and cause mutations [36]. In addition to the possible application in the making of selected disease models, Dr. Kim’s study may suggest the therapeutic potential of engineered nucleases of both ZFNs and TALENs in complicated genetic diseases [17].
As mentioned previously, reversed gene therapy involves functional or structural normal gene knockout. Functional gene knockout usually refers to simple gene code frameshifting. However, structural gene knockout requires exogenous gene fragment. Specific DNA sequences are precisely cleaved to form a DSB, followed by the HDR or NHEJ pathways. Therefore, the engineered nucleases such as ZFNs and TALENs could achieve efficient and designed homologous recombinations with their specific cleavage of gene targets in the presence of exogenous homologous genes.
Therapeutic Application of ZFNs and TALENs in HIV/AIDS Treatment
Conventional strategies including chemotherapy and vaccination have limited efficacy in fighting HIV infection. Chemotherapy treatment is restricted due to drug resistant virus and debilitating side effects. HIV vaccine development has never succeeded to pass the clinical trial phase. As listed in Table 4, HIV infection is a candidate for nuclease-mediated gene therapy and a few projects are actually in phase II clinical trials.
HIV infection requires the presence of the co-receptors chemokine (C–C motif) receptor type 5 (CCR5) [37] or chemokine (C-X-C motif) receptor type 4 (CXCR4) [38]. Epidemiological data showed that individuals with homozygous Δ32 deletion are absolutely resistant to the infection by CCR5-tropic HIV-1. Individuals with the heterozygous CCR5-Δ32 genotype have a lower possibility of being infected with HIV and take a longer time to progress to AIDS after the infection. On the other hand, the protective effect of anti-CCR5 IgA against HIV infection has been confirmed [39]. The aforementioned pieces of evidence prompted us and other groups to target CCR5 as a novel gene therapy strategy. In a paper published in April 2005, we described these HIV gene therapy strategies as follows: “In a therapeutic point of view AIDS can be treated as aplastic anemia or leukemia and the patients transplanted with bone marrow carrying CCR5-Δ32. … However, allogenic bone marrow transplantation has two drawbacks: immune rejection and the limited number of available donors. … Thus, in order to be more beneficial to more HIV-infected people, it is technically easier to introduce the CCR5-Δ32 genotype through genetic engineering of hemopoietic stem cells isolated from the patients themselves.”
Due to its efficacy and specificity, ZFN-mediated gene therapy has been used for the clinical treatment of HIV. Delivery of ZFNs targeting CCR5 using recombinant adenoviral vector was shown to disrupt >50 % of CCR5 alleles both in model cell lines and the primary human CD4+ T cells [40]. In the HIV-1-infected mice that received ZFN-modified CD4+ T cells, there was a substantial reduction in viral load and an increase in CD4+ T cell counts in peripheral tissues. Another study showed that ZFNs can disrupt CCR5 in human CD34+ hematopoietic stem/progenitor cells (HSPCs) at a mean frequency of 17 % of the total alleles in a population [41]. The results of this study suggested that ZFN-modified autologous hematopoietic stem cells could be considered a clinical approach to treating HIV-1.
Currently, two phase I clinical trials (NCT00842634 and NCT01044654) have been completed and one phase II trial investigating the clinical application of ZFNs-modified CD4+ T cells targeting CCR5 in patients with HIV/AIDS is in progress. These trials should bring support to the use of CCR5 ZFN-modified autologous CD4+ T cells in HIV patients as an attractive approach for the treatment of HIV-1 infection.
As one of the valuable engineered nucleases, TALENs were also designed to disrupt an EGFP marker gene and the human loci CCR5 [42]. Gene editing was achieved in up to 45 % of transfected cells. In comparison with ZFNs, TALENs showed similar gene disruption activities but significantly reduced nuclease-associated cytotoxicity and revealed only minimal off-target activity. Consequently, TALENs are considered as a key technology for targeted modifications of complex genomes.
Recently, the reported cure of an HIV-1 patient with acute myeloid leukemia appeared as an important milestone for the application of strategies targeting CCR5 [43, 44]. These results strongly supported the application of engineered nucleases in ex vivo and in vivo gene therapies to HIV infection.
Future Expectations
The fundamental barriers preventing the application of ZFNs and TALENs in patient treatment are their off-target effects and the immunological response of the organism to new genetically modified or corrected proteins. To some extent, reversed gene therapy may help to bypass these barriers [45, 46]. First, reversed gene therapy involves either local or temporal gene knock-out and may have a therapeutic effect or even cure selected diseases. It is less toxic compared to systematic or life time gene knock-out. Second, reversed gene therapy can be achieved simply by functional knock-out. Off-target effects can be better controlled in reversed gene therapy compared to gene correction as reversed gene therapy has a nearly whole-gene zone for the target of choice.
As far as the therapeutic outcomes of reversed gene therapy are concerned, the following points need to be understood. In genetic disorders such as enzyme deficiencies, a 5–10 % elevation of a missed enzyme may significantly relieve or even eliminate the clinical symptoms. Although a proportional disruption of an oncogene may have limited therapeutic effects on tumor, reversed gene therapy can be a component of a combined or synergistic cancer therapy. At this early age of reversed gene therapy, it would not be used to replace traditional chemotherapy. Instead, it will be mainly used for diseases lacking effective therapies, such as the aforementioned inherited genetic diseases and cancers. Infectious diseases are another type of candidates suitable for reversed gene therapy. Since most of the infections are strictly dependent on the pathogen–host interactions, genetic manipulations blocking these interactions either by targeting the pathogen or the host may successfully prevent or inhibit an infection. Just like chemotherapy treatments, an infection is not cured by direct pathogen killing by antibiotics, but by the immune system. For example, the transfusion of in vitro modified CD4 cells have no effect at all in wiping out existing HIV, but the increased level of CD4 cells helps to improve host’s immune system and to slow down the replication process of HIV, which makes a cure of HIV infection possible.
The newly developed reversed gene therapy appears promising for the treatment of hemophilia, thalassemia, DMD, neuronal deafness, Stargardt’s syndrome, essential hypertension, and TP53 gene mutation as well as lung, breast, cervix, prostate, colon, rectum cancers, and leukemia. This list will get longer with a better understanding of the pathophysiology of human diseases. TALENs are a new type of “drugable” macromolecules. As compared to traditional drugs, the effectiveness and toxicity of TALENs are easy to determine by screening their gene editing spectra and off-target effects. With the availability of the next generation sequencing technology, both gene editing spectra and off-target effects can be efficiently assayed using cultured human cells. Thus, engineered nucleases particularly the “drugable” TALENs are expected to be invaluable for future gene therapy. However, the gene editing technology is only one approach in gene therapy as the lack of gene delivery is still an obstacle to overcome. Fortunately, both lentiviral and adenoviral vectors are efficient enough to deliver ZFN and TALENs in vivo and ex vivo, particularly when used in reversed gene therapies that only requires a partial knockout of a selected gene to reach an observable clinical symptom improvement.
On the basis of the current state of knowledge, it is reasonable to suggest that the development of engineered nucleases described in this review will hold much promise for their use in safe and effective gene therapy approaches. We also have reasons to believe that gene therapy including reversed gene therapy with these nucleases and advances in stem cell technologies will provide effective treatments to lower the clinical severities or even cure a number of currently untreated or poorly treated diseases.
References
He, N., Zeng, X., Wang, W., Deng, K., Pan, Y., Xiao, L., et al. (2011). Challenges and future expectations of reversed gene therapy. Nanoscience and Nanotechnology, 11, 8634–8638.
Moscou, M. J., & Bogdanove, A. J. (2009). A simple cipher governs DNA recognition by TAL effectors. Science, 326, 1501.
Boch, J., Scholze, H., Schornack, S., Landgraf, A., Hahn, S., Kay, S., et al. (2009). Breaking the code of DNA binding specificity of TAL-type III effectors. Science, 326, 1509–1512.
Morbitzer, R., Römer, P., Boch, J., & Lahaye, T. (2011). Regulation of selected genome loci using de novo-engineered transcription activator-like effector (TALE)-type transcription factor. Proceedings of the National Academy of Sciences of the United States of America, 107, 21617–21622.
Geissler, R., Scholze, H., Hahn, S., Streubel, J., Bonas, U., Behrens, S. E., et al. (2011). Transcriptional activators of human genes with programmable DNA-specificity. PLoS ONE, 6, e19509.
Christian, M., Cermak, T., Doyle, E. L., Schmidt, C., Zhang, F., Hummel, A., et al. (2010). TAL effector nucleases create targeted DNA double-strand breaks. Genetics, 186, 757–761.
Mahfouz, M. M., Li, L., Shamimuzzaman, M., Wibowo, A., Fang, X., & Zhu, J. K. (2011). De novo-engineered transcription activator-like effector (TALE) hybrid nuclease with novel DNA binding specificity creates double-strand breaks. Proceedings of the National Academy of Sciences of the United States of America, 108, 2623–2628.
Beumer, K. J., Trautman, J. K., Bozas, A., Liu, J. L., Rutter, J., Gall, J. G., et al. (2008). Efficient gene targeting in drosophila by direct embryo injection with zinc-finger nucleases. Proceedings of the National Academy of Sciences of the United States of America, 105, 19821–19826.
Doyon, Y., McCammon, J. M., Miller, J. C., Faraji, F., Ngo, C., Katibah, G. E., et al. (2008). Heritable targeted gene disruption in zebrafish using designed zinc-finger nucleases. Nature Biotech, 26, 702–708.
Meng, X., Noyes, M. B., Zhu, L. J., Lawson, N. D., & Wolfe, S. A. (2008). Targeted gene inactivation in zebrafish using engineered zinc-finger nucleases. Nature Biotech, 26, 695–701.
Foley, J. E., Yeh, J. R., Maeder, M. L., Reyon, D., Sander, J. D., Peterson, R. T., et al. (2009). Rapid mutation of endogenous zebrafish genes using zinc finger nucleases made by Oligomerized Pool ENgineering (OPEN). PLoS ONE, 4, e4348.
Siekmann, A. F., Standley, C., Fogarty, K. E., Wolfe, S. A., & Lawson, N. D. (2009). Chemokine signaling guides regional patterning of the first embryonic artery. Genes & Development, 23, 2272–2277.
Santiago, Y., Chan, E., Liu, P. Q., Orlando, S., Zhang, L., Urnov, F. D., et al. (2008). Targeted gene knockout in mammalian cells using engineered zinc finger nucleases. Proceedings of the National Academy of Sciences of the United States of America, 105, 5809–5814.
Cost, G. J., Freyvert, Y., Vafiadis, A., Santiago, Y., Miller, J. C., Rebar, E., et al. (2009). BAK and BAX deletion using zinc-finger nucleases yields apoptosis-resistant CHO cells. Biotechnology and Bioengineering, 105, 330–340.
Liu, P. Q., Chan, E. M., Cost, G. J., Zhang, L., Wang, J., Miller, J. C., et al. (2010). Generation of a triple-gene knockout mammalian cell line using engineered zinc-finger nucleases. Biotechnology and Bioengineering, 106, 97–105.
Geurts, A. M., Cost, G. J., Freyvert, Y., Zeitler, B., Miller, J. C., Choi, V. M., et al. (2009). Knockout rats via embryo microinjection of zinc-finger nucleases. Science, 325, 433.
Lee, H. J., Kim, E., & Kim, J. S. (2010). Targeted chromosomal deletions in human cells using zinc finger nucleases. Genome Research, 20, 81–89.
Miller, J. C., Tan, S., Qiao, G., Barlow, K. A., Wang, J., Xia, D. F., et al. (2011). A TALE nuclease architecture for efficient genome editing. Nature Biotechnology, 29, 143–148.
Huang, P., Xiao, A., Zhou, M., Zhu, Z., Lin, S., & Zhang, B. (2011). Heritable gene targeting in zebrafish using customized TALENs. Nature Biotechnology, 29, 699–700.
Silva, G., Poirot, L., Galetto, R., Smith, J., Montoya, G., Duchateau, P., et al. (2011). Meganucleases and other tools for targeted genome engineering: perspectives and challenges for gene therapy. Current Gene Therapy, 11, 11–27.
Cai, C. Q., Doyon, Y., Ainley, W. M., Miller, J. C., Dekelver, R. C., Moehle, E. A., et al. (2009). Targeted transgene integration in plant cells using designed zinc finger nucleases. Plant Molecular Biology, 69, 699–709.
Shukla, V. K., Doyon, Y., Miller, J. C., DeKelver, R. C., Moehle, E. A., Worden, S. E., et al. (2009). Precise genome modification in the crop species Zea mays using zinc-finger nucleases. Nature, 459, 437–441.
Goldberg, A. D., Banaszynski, L. A., Noh, K. M., Lewis, P. W., Elsaesser, S. J., Stadler, S., et al. (2010). Distinct factors control histone variant H3.3 localization at specific genomic regions. Cell, 140, 678–691.
Moehle, E. A., Rock, J. M., Lee, Y. L., Jouvenot, Y., DeKelver, R. C., Gregory, P. D., et al. (2007). Targeted gene addition into a specified location in the human genome using designed zinc finger nucleases. Proceedings of the National Academy of Sciences of the United States of America, 104, 3055–3060.
Lombardo, A., Genovese, P., Beausejour, C. M., Colleoni, S., Lee, Y. L., Kim, K. A., et al. (2007). Gene editing in human stem cells using zinc finger nucleases and integrase-defective lentiviral vector delivery. Nature Biotech, 25, 1298–1306.
Benabdallah, B. F., Allard, E., Yao, S., Friedman, G., Gregory, P. D., Eliopoulos, N., et al. (2010). Targeted gene addition to human mesenchymal stromal cells as a cell-based plasma-soluble protein delivery platform. Cytotherapy, 12, 394–399.
Hockemeyer, D., Soldner, F., Beard, C., Gao, Q., Mitalipova, M., DeKelver, R. C., et al. (2009). Efficient targeting of expressed and silent genes in human ESCs and iPSCs using zinc-finger nucleases. Nature Biotechnology, 27, 851–857.
Hockemeyer, D., Wang, H., Kiani, S., Lai, C. S., Gao, Q., Cassady, J. P., et al. (2011). Genetic engineering of human pluripotent cells using TALE nucleases. Nature Biotechnology, 29, 731–734.
Urnov, F. D., Miller, J. C., Lee, Y. L., Beausejour, C. M., Rock, J. M., Augustus, S., et al. (2005). Highly efficient endogenous human gene correction using designed zinc-finger nucleases. Nature, 435, 646–651.
Maeder, M. L., Thibodeau-Beganny, S., Osiak, A., Wright, D. A., Anthony, R. M., Eichtinger, M., et al. (2008). Rapid ‘open-source’ engineering of customized zinc-finger nucleases for highly efficient gene modification. Molecular Cell, 31, 294–301.
Li, H., Haurigot, V., Doyon, Y., Li, T., Wong, S. Y., Bhagwat, A. S., et al. (2011). In vivo genome editing restores haemostasis in a mouse model of haemophilia. Nature, 475, 217–221.
Townsend, J. A., Wright, D. A., Winfrey, R. J., Fu, F., Maeder, M. L., Joung, J. K., et al. (2009). High-frequency modification of plant genes using engineered zinc-finger nucleases. Nature, 459, 442–445.
Osakabe, K., Osakabe, Y., & Toki, S. (2010). Site-directed mutagenesis in Arabidopsis using custom-designed zinc finger nucleases. Proceedings of the National Academy of Sciences of the United States of America, 107, 12034–12039.
Goldberg, A. D., Banaszynski, L. A., Noh, K. M., Lewis, P. W., Elsaesser, S. J., Stadler, S., et al. (2011). Distinct factors control histone variant H3.3 localization at specific genomic regions. Cell, 140, 678–691.
Caroll, D., & Zhang, B. (2011). Primer and interviews: Advances in targeted gene modification. Developmental Dynamics, 240, 2688–2696.
Lee, H. J., Kweon, J., Kim, E., Kim, S., & Kim, J. S. (2012). Targeted chromosomal duplications and inversions in the human genome using zinc finger nucleases. Genome Research, 22, 539–548.
Chen, L. L., Zhang, J., Gao, H. L., Liao, D. F., Li, K., (2005) The relationship between CCR5 and HIV-1 disease. Journal of University of South China (Medical Edition) 33, 166–170 (In Chinese).
Zaitseva, M., Blauvelt, A., Lee, S., Lapham, C. K., Klaus-Kovtun, V., Mostowski, H., et al. (1997). Expression and function of CCR5 and CXCR4 on human Langerhans cells and macrophages: Implications for HIV primary infection. Nature Medicine, 3, 1369–1375.
Lopalco, L., Barassi, C., Pastori, C., Longhi, R., Burastero, S. E., Tambussi, G., et al. (2000). CCR5-reactive antibodies in seronegative partners of HIV-seropositive individuals down-modulate surface CCR5 in vivo and neutralize the infectivity of R5 strains of HIV-1 in vitro. Journal of Immunology, 164, 3426–3433.
Perez, E. E., Wang, J., Miller, J. C., Jouvenot, Y., Kim, K. A., Liu, O., et al. (2008). Establishment of HIV-1 resistance in CD4+ T cells by genome editing using zinc-finger nucleases. Nature Biotechnology, 26, 808–816.
Holt, N., Wang, J., Kim, K., Friedman, G., Wang, X., Taupin, V., et al. (2010). Human hematopoietic stem/progenitor cells modified by zinc-finger nuclease targeted to CCR5 control HIV-1 in vivo. Nature Biotechnology, 28, 839–847.
Mussolino, C., Morbitzer, R., Lütge, F., Dannemann, N., Lahaye, T., Cathomen, T., et al. (2011). A novel TALE nuclease scaffold enables high genome editing activity in combination with low toxicity. Nucleic Acids Research, 39, 9283–9293.
Allers, K., Hütter, G., Hofmann, J., Loddenkemper, C., Rieger, K., Thiel, E., et al. (2011). Evidence for the cure of HIV infection by CCR5Δ32/Δ32 stem cell transplantation. Blood, 117, 2791–2799.
Li, K., Zhang, J., Xiao, L., & Sirois, Pierre. (2011). Gene therapy: Hope for HIV infection cure. Blood, e-letter, 01, 26.
He, N., Zeng, X., Wang, W., Deng, K. L., Pan, Y., Xiao, L., et al. (2011). Challenges and future expectations of reversed gene therapy. Journal of Nanoscience and Nanotechnology, 11, 8634–8638.
Zhang, J., Li, A. S. S., Kou, X., Xiao, L., Li, P., He, N., et al. (2012). Why CCR5 is chosen as the target for stem cell gene therapy for HIV infection? Journal of Nanoscience and Nanotechnology, 12, 2045–2048.
Bibikova, M., Beumer, K., Trautman, J. K., & Carroll, D. (2003). Enhancing gene targeting with designed zinc finger nucleases. Science, 300, 764.
Beumer, K., Bhattacharyya, G., Bibikova, M., Trautman, J. K., & Carroll, D. (2006). Efficient gene targeting in Drosophila with zinc-finger nucleases. Genetics, 172, 2391–2403.
Malphettes, L., Freyvert, Y., Chang, J., Liu, P. Q., Chan, E., Miller, J. C., et al. (2010). Highly efficient deletion of FUT8 in CHO cell lines using zinc-finger nucleases yields cells that produce completely nonfucosylated antibodies. Biotechnology and Bioengineering, 106, 774–783.
Mashimo, T., Takizawa, A., Voigt, B., Yoshimi, K., Hiai, H., Kuramoto, T., et al. (2010). Generation of knockout rats with X-linked severe combined immunodeficiency (X-SCID) using zinc-finger nucleases. PLoS ONE, 5, e8870.
Del Prete, G. Q., Haggarty, B., Leslie, G. J., Jordan, A. P., Romano, J., Wang, N., et al. (2009). Derivation and characterization of a simian immunodeficiency virus SIVmac239 variant with tropism for CXCR4. Journal of Virology, 83, 9911–9922.
Doyon, Y., Choi, V. M., Xia, D. F., Vo, T. D., Gregory, P. D., & Holmes, M. C. (2010). A transient cold shock enhances zinc-finger nuclease-mediated gene disruption. Nature Methods, 7, 459–460.
Zhang, F., Maeder, M. L., Unger-Wallace, E., Hoshaw, J. P., Reyon, D., Christian, M., et al. (2010). High frequency targeted mutagenesis in Arabidopsis thaliana using zinc finger nuclease. Proc. Proc Natl Acad Sci USA, 107, 12028–12033.
Zou, J., Maeder, M. L., Mali, P., Pruett-Miller, S. M., Thibodeau-Beganny, S., Chou, B. K., et al. (2009). Gene targeting of a disease-related gene in human induced pluripotent stem and embryonic stem cells. Cell Stem Cell, 5, 97–110.
Zhang, F., Cong, L., Lodato, S., Kosuri, S., Church, G. M., & Arlotta, P. (2011). Efficient construction of sequence-specific TAL effectors for modulating mammalian transcription. Nature Biotechnology, 29, 149–153.
Wood, A. J., Lo, T. W., Zeitler, B., Pickle, C. S., Ralston, E. J., Lee, A. H., et al. (2011). Targeted genome editing across species using ZFNs and TALENs. Science, 333, 307.
Tesson, L., Usal, C., Ménoret, S., Leung, E., Niles, B. J., Remy, S., et al. (2011). Knockout rats generated by embryo microinjection of TALENs. Nature Biotechnology, 29, 695–696.
Thermes, V., Grabher, C., Ristoratore, F., Bourrat, F., Choulika, A., Wittbrodt, J., et al. (2002). I-SceI meganuclease mediates highly efficient transgenesis in fish. Mechanisms of Development, 118, 91–98.
Grabher, C., & Wittbrodt, J. (2007). Meganuclease and transposon mediated transgenesis in medaka. Genome Biology, 8, S10.
Grosse, S., Huot, N., Mahiet, C., Arnould, S., Barradeau, S., Clerre, D. L., et al. (2011). Meganuclease-mediated inhibition of HSV1 infection in cultured cells. Molecular Therapy, 19, 694–702.
Kim, H. J., Lee, H. J., Kim, H., Cho, S. W., & Kim, J. S. (2009). Targeted genome editing in human cells with zinc finger nucleases constructed via modular assembly. Genome Research, 19, 1279–1288.
Kim, H., Um, E., Cho, S. R., Jung, C., Kim, H., & Kim, J. S. (2011). Surrogate reporters for enrichment of cells with nuclease-induced mutations. Nature Methods, 8, 941–943.
Kim, S., Lee, M. J., Kim, H., Kang, M., & Kim, J. S. (2011). Preassembled zinc-finger arrays for rapid construction of ZFNs. Nature Methods, 8, 7.
Sun, N., Liang, J., Abil, Z., & Zhao, H. (2012). Optimized TAL effector nucleases (TALENs) for use in treatment of sickle cell disease. Molecular BioSystems, 8, 1255–1263.
Bonas, U., Stall, R. E., & Stskawicz, B. J. (1989). Genetic and structural characterization of the avirulence gene avrBs3 from Xanthomonas campestris pv. campesris. Molecular and General Genetics, 218(1), 127–136.
Miller, J., McLachlan, A. D., & Klug, A. (1985). Repetitive zinc-binding domains in the protein transcription factor IIIA from Xenopus oocytes. EMBO Journal, 4, 1609–1614.
Kim, Y. G., Cha, J., & Chandrasegaran, S. (1996). Hybrid restriction enzymes: Zinc finger fusions to FokI cleavage domain. Proceedings of the National Academy of Sciences of the USA, 93, 1156–1160.
Acknowledgments
This study is partially supported by National Natural Science Foundation of China (No. 30970877), Chinese National 863 Major Grant (No. 2012AA020905), the Priority Academic Program Development of Jiangsu Higher Education Institutions and Innovation Project of Jiangsu Graduate Education (No. CXZZ11_0117).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Yunzhi Pan and Li Xiao have contributed equally to this paper.
Rights and permissions
About this article
Cite this article
Pan, Y., Xiao, L., Li, A.S.S. et al. Biological and Biomedical Applications of Engineered Nucleases. Mol Biotechnol 55, 54–62 (2013). https://doi.org/10.1007/s12033-012-9613-9
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12033-012-9613-9