Abstract
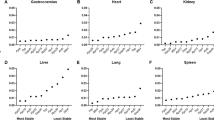
Liver regeneration (LR) is a process during which the liver recovers its mass and function after damage due to various causes such as partial hepatectomy (PH). It involves a sequence of well-orchestrated changes in physiological and biochemical activities, especially in the gene expression profile in a variety of liver cells. In order to produce reliable gene expression of target genes in eight kinds of rat hepatic cells during LR, the determination of internal control housekeeping genes (HKGs) is required. Eight kinds of hepatic cells were first isolated from liver tissue with high purity and activity. Then quantitative real-time reverse transcription (RT)-PCR was applied to detect expression changes of six commonly used HKGs (18SrRNA, B2M, ACTB, UBC, GAPDH, and HK1) in eight types of hepatic cells isolated from regenerating liver at 0, 2, 6, 12, 24, 30, 36, 72, 120, and 168 h after PH. The amplification of the HKGs was statistically analyzed by using geNorm algorithm. Using this method, 18SrRNA–UBC, ACTB–HK1, ACTB–GADPH, B2M–ACTB, 18SrRNA–UBC, B2M–UBC, B2M–ACTB, and B2M–UBC were found to be the two most stable reference genes for rat regenerating hepatocytes, hepatic stellate cells, Kupffer cells, biliary epithelial cells, sinusoidal endothelial cells, pit cells, dendritic cells, and oval cells, respectively, regardless of the stages of LR. In conclusion, this study has laid a good foundation for investigating gene expression of target genes in different types of hepatic cells during LR.



Similar content being viewed by others
References
Michalopoulos, G. K., & DeFrances, M. C. (1997). Liver regeneration. Science, 276, 60–66.
Taub, R. (2004). Liver regeneration: From myth to mechanism. Nature reviews. Molecular cell biology, 5, 836–847.
Bustin, S. A. (2002). Quantification of mRNA using real-time reverse transcription PCR (RT-PCR): Trends and problems. Journal of Molecular Endocrinology, 29, 23–39.
Robert, C., McGraw, S., Massicotte, L., Pravetoni, M., Gandolfi, F., & Sirard, M. A. (2002). Quantification of housekeeping transcript levels during the development of bovine preimplantation embryos. Biology of Reproduction, 67, 1465–1472.
Thellin, O., Zorzi, W., Lakaye, B., De Borman, B., Coumans, B., Hennen, G., et al. (1999). Housekeeping genes as internal standards: Use and limits. Journal of Biotechnology, 75, 291–295.
Bustin, S. A. (2000). Absolute quantification of mRNA using real-time reverse transcription polymerase chain reaction assays. Journal of Molecular Endocrinology, 25, 169–193.
Suzuki, T., Higgins, P. J., & Crawford, D. R. (2000). Control selection for RNA quantitation. Biotechniques, 29, 332–337.
Michalopoulos, G. K. (2007). Liver regeneration. Journal of Cellular Physiology, 213, 286–300.
Gao, Q., Wang, X. Y., Fan, J., Qiu, S. J., Zhou, J., Shi, Y. H., et al. (2008). Selection of reference genes for real-time PCR in human hepatocellular carcinoma tissues. Journal of Cancer Research and Clinical Oncology, 134, 979–986.
Vandesompele, J., De Preter, K., Pattyn, F., Poppe, B., Van Roy, N., De Paepe, A., et al. (2002). Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biology, 3, RESEARCH0034.
Higgins, G. M., & Anderson, R. M. (1931). Experimental pathology of the liver: Restoration of the liver of the white rat following partial surgical removal. Archives of Pathology & Laboratory Medicine, 186–202.
Kreamer, B. L., Staecker, J. L., Sawada, N., Sattler, G. L., Hsia, M. T., & Pitot, H. C. (1986). Use of a low-speed, iso-density percoll centrifugation method to increase the viability of isolated rat hepatocyte preparations. In Vitro Cellular & Developmental Biology, 22, 201–211.
Ferry, N., Duplessis, O., Houssin, D., Danos, O., & Heard, J. M. (1991). Retroviral-mediated gene transfer into hepatocytes in vivo. Proceedings of the National Academy of Sciences of the United States of America, 88, 8377–8381.
Jurima-Romet, M., Huang, H. S., Paul, C. J., & Thomas, B. H. (1991). Enalapril cytotoxicity in primary cultures of rat hepatocytes. II. Role of glutathione. Toxicology Letters, 58, 269–277.
Haber, B. A., Mohn, K. L., Diamond, R. H., & Taub, R. (1993). Induction patterns of 70 genes during nine days after hepatectomy define the temporal course of liver regeneration. Journal of Clinical Investigation, 91, 1319–1326.
Alpini, G., Phillips, J. O., Vroman, B., & LaRusso, N. F. (1994). Recent advances in the isolation of liver cells. Hepatology, 20, 494–514.
Jurima-Romet, M., Abbott, F. S., Tang, W., Huang, H. S., & Whitehouse, L. W. (1996). Cytotoxicity of unsaturated metabolites of valproic acid and protection by vitamins C and E in glutathione-depleted rat hepatocytes. Toxicology, 112, 69–85.
Blair, J. B., Ostrander, G. K., Miller, M. R., & Hinton, D. E. (1995). Isolation and characterization of biliary epithelial cells from rainbow trout liver. In Vitro Cellular & Developmental Biology. Animal, 31, 780–789.
Fiegel, H. C., Park, J. J., Lioznov, M. V., Martin, A., Jaeschke-Melli, S., Kaufmann, P. M., et al. (2003). Characterization of cell types during rat liver development. Hepatology, 37, 148–154.
Oren, A., Husebo, C., Iversen, A. C., & Austgulen, R. (2005). A comparative study of immunomagnetic methods used for separation of human natural killer cells from peripheral blood. Journal of Immunological Methods, 303, 1–10.
Yang, H., Qu, L., Wimbrow, A. N., Jiang, X., & Sun, Y. (2007). Rapid detection of Listeria monocytogenes by nanoparticle-based immunomagnetic separation and real-time PCR. International Journal of Food Microbiology, 118, 132–138.
Chen, Y., Dong, X. J., Zhang, G. R., Shao, J. Z., & Xiang, L. X. (2006). Transdifferentiation of mouse BM cells into hepatocyte-like cells. Cytotherapy, 8, 381–389.
Handharyani, E., Ochiai, K., Iwata, N., & Umemura, T. (2001). Immunohistochemical and ultrastructural study of ito cells (fat-storing cells) in response to extrahepatic bile duct ligation in broiler chickens. Journal of Veterinary Medical Science, 63, 547–552.
Dory, D., Echchannaoui, H., Letiembre, M., Ferracin, F., Pieters, J., Adachi, Y., et al. (2003). Generation and functional characterization of a clonal murine periportal Kupffer cell line from H-2Kb-tsA58 mice. Journal of Leukocyte Biology, 74, 49–59.
Tokiwa, T., Yamazaki, T., Ono, M., Enosawa, S., & Tsukiyama, T. (2008). Cloning and characterization of liver progenitor cells from the scattered cell clusters in primary culture of porcine livers. Cell Transplantation, 17, 179–186.
Scoazec, J. Y., & Feldmann, G. (1991). In situ immunophenotyping study of endothelial cells of the human hepatic sinusoid: Results and functional implications. Hepatology, 14, 789–797.
Mochizuki, K., Ohno, Y., Kanematsu, T., Sakurai-Yamashita, Y., Niwa, M., Hishikawa, Y., et al. (2007). Possible protection of sinusoidal endothelial cells by endothelin B receptor during hepatic warm ischemia-reperfusion. Surgery Today, 37, 460–467.
Liu, X. H., & Zhou, X. J. (1999). Ultrastructural observations and analysis of liver-associated lymphocytes in hepatitis B and C. Journal of Chinese Electron Microscopy Society, 18, 606–610.
Hagenaars, M., Ensink, N. G., Eggermont, A. M., van der Velde, E. A., van de Velde, C. J., Fleuren, G. J., et al. (2000). Endogenous natural killer cells do not play a role in antitumor effects induced by interleukin-2 in a syngeneic rat colon tumor model. Cancer Immunology, Immunotherapy, 48, 561–568.
Perez, L., Shurin, M. R., Collins, B., Kogan, D., Tourkova, I. L., & Shurin, G. V. (2005). Comparative analysis of CD1a, S-100, CD83, and CD11c human dendritic cells in normal, premalignant, and malignant tissues. Histology and Histopathology, 20, 1165–1172.
Shan, Y. F., Zhou, W. P., Fu, X. Y., Yan, H. X., Yang, W., Liu, S. Q., et al. (2006). The role of p28GANK in rat oval cells activation and proliferation. Liver International, 26, 240–247.
You, W. J., Xiao, M. D., & Yuan, Z. X. (2004). Significance of changes in transforming growth factor-beta mRNA levels in autogenous vein grafts. Chinese Medical Journal, 117, 1060–1065.
Knepp, J. H., Geahr, M. A., Forman, M. S., & Valsamakis, A. (2003). Comparison of automated and manual nucleic acid extraction methods for detection of enterovirus RNA. Journal of Clinical Microbiology, 41, 3532–3536.
Arkin, A., Ross, J., & McAdams, H. H. (1998). Stochastic kinetic analysis of developmental pathway bifurcation in phage lambda-infected Escherichia coli cells. Genetics, 149, 1633–1648.
Joplin, R., Hishida, T., Tsubouchi, H., Daikuhara, Y., Ayres, R., Neuberger, J. M., et al. (1992). Human intrahepatic biliary epithelial cells proliferate in vitro in response to human hepatocyte growth factor. Journal of Clinical Investigation, 90, 1284–1289.
Clavien, P. A. (2008). Liver regeneration: A spotlight on the novel role of platelets and serotonin. Swiss Medical Weekly, 138, 361–370.
Arosh, J. A., Banu, S. K., Chapdelaine, P., Emond, V., Kim, J. J., MacLaren, L. A., et al. (2003). Molecular cloning and characterization of bovine prostaglandin E2 receptors EP2 and EP4: Expression and regulation in endometrium and myometrium during the estrous cycle and early pregnancy. Endocrinology, 144, 3076–3091.
Makihira, S., Yan, W., Murakami, H., Furukawa, M., Kawai, T., Nikawa, H., et al. (2003). Thyroid hormone enhances aggrecanase-2/ADAM-TS5 expression and proteoglycan degradation in growth plate cartilage. Endocrinology, 144, 2480–2488.
Fausto, N., Campbell, J. S., & Riehle, K. J. (2006). Liver regeneration. Hepatology, 43, S45–S53.
Tatsumi, K., Ohashi, K., Taminishi, S., Okano, T., Yoshioka, A., & Shima, M. (2008). Reference gene selection for real-time RT-PCR in regenerating mouse livers. Biochemical and Biophysical Research Communications, 374, 106–110.
Schmittgen, T. D., & Zakrajsek, B. A. (2000). Effect of experimental treatment on housekeeping gene expression: validation by real-time, quantitative RT-PCR. Journal of Biochemical and Biophysical Methods, 46, 69–81.
Goidin, D., Mamessier, A., Staquet, M. J., Schmitt, D., & Berthier-Vergnes, O. (2001). Ribosomal 18S RNA prevails over glyceraldehyde-3-phosphate dehydrogenase and beta-actin genes as internal standard for quantitative comparison of mRNA levels in invasive and noninvasive human melanoma cell subpopulations. Analytical Biochemistry, 295, 17–21.
Conaway, R. C., Brower, C. S., & Conaway, J. W. (2002). Emerging roles of ubiquitin in transcription regulation. Science, 296, 1254–1258.
Glare, E. M., Divjak, M., Bailey, M. J., & Walters, E. H. (2002). beta-Actin and GAPDH housekeeping gene expression in asthmatic airways is variable and not suitable for normalising mRNA levels. Thorax, 57, 765–770.
Dheda, K., Huggett, J. F., Bustin, S. A., Johnson, M. A., Rook, G., & Zumla, A. (2004). Validation of housekeeping genes for normalizing RNA expression in real-time PCR. Biotechniques, 37, 112–114, 6, 8–9.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wang, GP., Xu, CS. Reference Gene Selection for Real-Time RT-PCR in Eight Kinds of Rat Regenerating Hepatic Cells. Mol Biotechnol 46, 49–57 (2010). https://doi.org/10.1007/s12033-010-9274-5
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12033-010-9274-5