Abstract
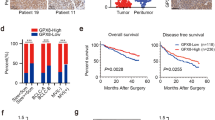
Tripartite motif-containing 3 (TRIM3) is a member of the tripartite motif (TRIM) protein family and is reported to be involved in the pathogenesis of various cancers. The role of TRIM3 in hepatocellular carcinoma (HCC) is unknown; thus, the goal of this study was to explore the expression level and prognostic value of TRIM3 in HCC. The expression level of TRIM3 in HCC surgically resected tumors and corresponding nontumorous samples was detected by real-time quantitative RT-PCR, Western blotting, and immunohistochemistry. The correlation between TRIM3 expression level and the clinicopathological features and prognosis of HCC patients was also analyzed. We observed that TRIM3 expression was remarkably decreased in tumor tissue samples from HCC patients, relative to matched nontumorous tissue samples, at the mRNA (p = 0.018) and protein level (p = 0.02). Similarly, immunohistochemical analysis showed that 53.4 % of samples had low TRIM3 protein expression. Clinicopathological analysis revealed that low TRIM3 expression was significantly correlated with tumor size (p = 0.034), histological grade (p < 0.001), serum AFP (p = 0.025), and TNM stage (p = 0.021). Furthermore, Kaplan–Meier survival analysis revealed that low TRIM3 expression was associated with poor survival in HCC patients. Finally, our multivariate Cox regression analysis showed that TRIM3 expression was an independent prognostic factor for overall survival of HCC patients. In conclusion, this study suggests that TRIM3 may play a significant role in HCC progression and acts as a valuable prognostic marker and potential therapeutic target for HCC.




Similar content being viewed by others
References
Shariff MI, Cox IJ, Gomaa AI, Khan SA, Gedroyc W, Taylor-Robinson SD. Hepatocellular carcinoma: current trends in worldwide epidemiology, risk factors, diagnosis and therapeutics. Expert Rev Gastroenterol Hepatol. 2009;3:353–67.
Yang S, Yan HL, Tao QF, Yuan SX, Tang GN, Yang Y, et al. Low CADM2 expression predicts high recurrence risk of hepatocellular carcinoma patients after hepatectomy. J Cancer Res Clin Oncol. 2014;140:109–16.
Wu J, Du C, Lv Z, Ding C, Cheng J, Xie H, et al. The up-regulation of histone deacetylase 8 promotes proliferation and inhibits apoptosis in hepatocellular carcinoma. Dig Dis Sci. 2013;58:3545–53.
Pan QZ, Pan K, Zhao JJ, Chen JG, Li JJ, Lv L, et al. Decreased expression of interleukin-36alpha correlates with poor prognosis in hepatocellular carcinoma. Cancer Immunol Immunother. 2013;62:1675–85.
Chen S, Chen J, Xi W, Xu W, Yin G. Clinical therapeutic effect and biological monitoring of p53 gene in advanced hepatocellular carcinoma. Am J Clin Oncol. 2014;37:24–9.
Wang KF, Pan W, Wang F, Wang GF, Madhava P, Pan HM, et al. Geometric optimization of a mathematical model of radiofrequency ablation in hepatic carcinoma. Asian Pac J Cancer Prev. 2013;14:6151–8.
Zhao JJ, Pan K, Li JJ, Chen YB, Chen JG, Lv L, et al. Identification of LZAP as a new candidate tumor suppressor in hepatocellular carcinoma. PLoS ONE. 2011;6:e26608.
Zabaleta J, Aguinagalde B, Izquierdo JM, Bazterargui N, Laguna SM, Martin-Arruti M, et al. The presence of mutations in the K-RAS gene does not affect survival after resection of pulmonary metastases from colorectal cancer. ISRN Surg. 2014;2014:157586.
Hu S, Wu X, Zhou B, Xu Z, Qin J, Lu H, et al. IMP3 combined with CD44s, a novel predictor for prognosis of patients with hepatocellular carcinoma. J Cancer Res Clin Oncol. 2014;140:883–93.
Yang Y, Fan YC, Gao S, Dou CY, Zhang JJ, Sun FK, et al. Methylated cysteine dioxygenase-1 gene promoter in the serum is a potential biomarker for hepatitis B virus-related hepatocellular carcinoma. Tohoku J Exp Med. 2014;232:187–94.
Liu C, Xia Y, Jiang W, Liu Y, Yu L. Low expression of GABARAPL1 is associated with a poor outcome for patients with hepatocellular carcinoma. Oncol Rep. 2014;31:2043–8.
Xu X, Guo HJ, Xie HY, Li J, Zhuang RZ, Ling Q, et al. ZIP4, a novel determinant of tumor invasion in hepatocellular carcinoma, contributes to tumor recurrence after liver transplantation. Int J Biol Sci. 2014;10:245–56.
Mann DA. Epigenetics in liver disease. Hepatology. 2014. doi:10.1002/hep.27131.
Napolitano LM, Meroni G. TRIM family: pleiotropy and diversification through homomultimer and heteromultimer formation. IUBMB Life. 2012;64:64–71.
Nisole S, Stoye JP, Saib A. TRIM family proteins: retroviral restriction and antiviral defence. Nat Rev Microbiol. 2005;3:799–808.
El-Husseini AE, Vincent SR. Cloning and characterization of a novel RING finger protein that interacts with class V myosins. J Biol Chem. 1999;274:19771–7.
El-Husseini AE, Kwasnicka D, Yamada T, Hirohashi S, Vincent SR. BERP, a novel ring finger protein, binds to alpha-actinin-4. Biochem Biophys Res Commun. 2000;267:906–11.
Zoumpoulidou G, Broceno C, Li H, Bird D, Thomas G, Mittnacht S. Role of the tripartite motif protein 27 in cancer development. J Natl Cancer Inst. 2012;104:941–52.
Zhou ZY, Yang GY, Zhou J, Yu MH. Significance of TRIM29 and beta-catenin expression in non-small-cell lung cancer. J Chin Med Assoc. 2012;75:269–74.
Herquel B, Ouararhni K, Khetchoumian K, Ignat M, Teletin M, Mark M, et al. Transcription cofactors TRIM24, TRIM28, and TRIM33 associate to form regulatory complexes that suppress murine hepatocellular carcinoma. Proc Natl Acad Sci USA. 2011;108:8212–7.
Marshall GM, Bell JL, Koach J, Tan O, Kim P, Malyukova A, et al. TRIM16 acts as a tumour suppressor by inhibitory effects on cytoplasmic vimentin and nuclear E2F1 in neuroblastoma cells. Oncogene. 2010;29:6172–83.
Liu J, Welm B, Boucher KM, Ebbert MT, Bernard PS. TRIM29 functions as a tumor suppressor in nontumorigenic breast cells and invasive ER+breast cancer. Am J Pathol. 2012;180:839–47.
Liu Y, Raheja R, Yeh N, Ciznadija D, Pedraza AM, Ozawa T, et al. TRIM3, a tumor suppressor linked to regulation of p21(Waf1/Cip1). Oncogene. 2014;33:308–15.
Raheja R, Liu Y, Hukkelhoven E, Yeh N, Koff A. The ability of TRIM3 to induce growth arrest depends on RING-dependent E3 ligase activity. Biochem J. 2014;458:537–45.
Lim L, Balakrishnan A, Huskey N, Jones KD, Jodari M, Ng R, et al. MicroRNA-494 within an oncogenic microRNA megacluster regulates G1/S transition in liver tumorigenesis through suppression of mutated in colorectal cancer. Hepatology. 2014;59:202–15.
Dong YD, Cui L, Peng CH, Cheng DF, Han BS, Huang F. Expression and clinical significance of HMGB1 in human liver cancer: knockdown inhibits tumor growth and metastasis in vitro and in vivo. Oncol Rep. 2013;29:87–94.
Kashimoto K, Komatsu S, Ichikawa D, Arita T, Konishi H, Nagata H, et al. Overexpression of TRIM44 contributes to malignant outcome in gastric carcinoma. Cancer Sci. 2012;103:2021–6.
Acknowledgments
This study was supported by the Guangdong Province Science and Technology Plan project (2011A030400004).
Conflict of interest
The authors declare no conflict of interest.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Chao, J., Zhang, XF., Pan, QZ. et al. Decreased expression of TRIM3 is associated with poor prognosis in patients with primary hepatocellular carcinoma. Med Oncol 31, 102 (2014). https://doi.org/10.1007/s12032-014-0102-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12032-014-0102-9