Abstract
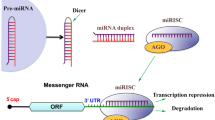
MicroRNAS (miRNAs) have been suggested to play important roles in the central nervous system during development as well as disease. miRNAs appear to be dysregulated in a number of neurodegenerative diseases, developmental disorders, and as a result of stroke. Each miRNA has the ability to regulate hundreds of messenger RNA transcripts, both by causing degradation of the mRNA and by inhibition of protein translation. Recent findings suggest that it may eventually be possible to treat some neurological disorders by restoring or inhibiting miRNAs altered by disease pathology. Both viral delivery and administration of modified oligonucleotides mimicking or inhibiting specific miRNAs have been effective in model systems. Artificial miRNAs have also been generated for the repression of specific transcripts. Alteration of miRNA expression by disease and insult also holds the potential for improved diagnostic tools. Finally, miRNAs have been shown to control cellular proliferation and specification, suggesting that manipulation of miRNAs in cultured cells could result in more convenient generation of pure cell populations for transplantation.

Similar content being viewed by others
References
Abelson, J. F., Kwan, K. Y., O’Roak, B. J., Baek, D. Y., Stillman, A. A., Morgan, T. M., et al. (2005). Sequence variants in SLITRK1 are associated with Tourette’s syndrome. Science, 310(5746), 317–320.
Baek, D., Villén, J., Shin, C., Camargo, F. D., Gygi, S. P., & Bartel, D. P. (2008). The impact of microRNAs on protein output. Nature, 455(7209), 64–71.
Beveridge, N. J., Tooney, P. A., Carroll, A. P., Gardiner, E., Bowden, N., Scott, R. J., et al. (2008). Dysregulation of miRNA 181b in the temporal cortex in schizophrenia. Human Molecular Genetics, 17(8), 1156–1168.
Boissonneault, V., Plante, I., Rivest, S., & Provost, P. (2009). MicroRNA-298 and microRNA-328 regulate expression of mouse beta-amyloid precursor protein-converting enzyme 1. Journal of Biological Chemistry, 284(4), 1971–1981.
Boudreau, R. L., Martins, I., & Davidson, B. L. (2009). Artificial microRNAs as siRNA shuttles: Improved safety as compared to shRNAs in vitro and in vivo. Molecular Therapy, 17(1), 169–175.
Burmistrova, O. A., Goltsov, A. Y., Abramova, L. I., Kaleda, V. G., Orlova, V. A., & Rogaev, E. I. (2007). MicroRNA in schizophrenia: Genetic and expression analysis of miR-130b (22q11). Biochemistry (Mosc), 72(5), 578–582.
Cao, X., Pfaff, S. L., & Gage, F. H. (2007). A functional study of miR-124 in the developing neural tube. Genes and Development, 21(5), 531–536.
Carthew, R. W., & Sontheimer, E. J. (2009). Origins and Mechanisms of miRNAs and siRNAs. Cell, 136, 642–655.
Castanotto, D., Sakurai, K., Lingeman, R., Li, H., Shively, L., Aagaard, L., et al. (2007). Combinatorial delivery of small interfering RNAs reduces RNAi efficacy by selective incorporation into RISC. Nucleic Acids Research, 35(15), 5154–5164.
Cataldo, A. M., Petanceska, S., Peterhoff, C. M., Terio, N. B., Epstein, C. J., Villar, A., et al. (2003). App gene dosage modulates endosomal abnormalities of Alzheimer’s disease in a segmental trisomy 16 mouse model of down syndrome. Journal of Neuroscience, 23, 6788–6792.
Cockrell, A. S., & Kafri, T. (2007). Gene delivery by lentivirus vectors. Molecular Biotechnology, 36, 184–204.
Cogswell, J. P., Ward, J., Taylor, I. A., Waters, M., Shi, Y., Cannon, B., et al. (2008). Identification of miRNA changes in Alzheimer’s disease brain and CSF yields putative biomarkers and insights into disease pathways. Journal of Alzheimer’s Disease, 14(1), 27–41.
Conaco, C., Otto, S., Han, J. J., & Mandel, G. (2006). Reciprocal actions of REST and a microRNA promote neuronal identity. Proceedings of the National Academy of Sciences of the United States of America, 103(7), 2422–2427.
Croce, C. M., & Calin, G. A. (2005). miRNAs, cancer, and stem cell division. Cell, 122(1), 6–7.
De Pietri Tonelli, D., Pulvers, J. N., Haffner, C., Murchison, E. P., Hannon, G. J., & Huttner, W. B. (2008). miRNAs are essential for survival and differentiation of newborn neurons but not for expansion of neural progenitors during early neurogenesis in the mouse embryonic neocortex. Development, 135, 3911–3921.
Dharap, A., Bowen, K., Place, R., Li, L. C., & Vemuganti, R. (2009). Transient focal ischemia induces extensive temporal changes in rat cerebral microRNAome. Journal of Cerebral Blood Flow and Metabolism, 29(4), 675–687.
Ebert, M. S., Neilson, J. R., & Sharp, P. A. (2007). MicroRNA sponges: Competitive inhibitors of small RNAs in mammalian cells. Nature Methods, 4(9), 721–726.
Elmén, J., Lindow, M., Schütz, S., Lawrence, M., Petri, A., Obad, S., et al. (2008a). LNA-mediated microRNA silencing in non-human primates. Nature, 452, 896–899.
Elmén, J., Lindow, M., Silahtaroglu, A., Bak, M., Christensen, M., Lind-Thomsen, A., et al. (2008b). Antagonism of microRNA-122 in mice by systemically administered LNA-antimiR leads to up-regulation of a large set of predicted target mRNAs in the liver. Nucleic Acids Research, 36, 1153–1162.
Faghihi, M. A., Modarresi, F., Khalil, A. M., Wood, D. E., Sahagan, B. G., Morgan, T. E., et al. (2008). Expression of a noncoding RNA is elevated in Alzheimer’s disease and drives rapid feed-forward regulation of beta-secretase. Nature Medicine, 14(7), 723–730.
Gonzalez-Alegre, P., Bode, N., Davidson, B. L., & Paulson, H. L. (2005). Silencing primary dystonia: Lentiviral-mediated RNA interference therapy for DYT1 dystonia. Journal of Neuroscience, 25, 10502–10509.
Grimm, D., Streetz, K. L., Jopling, C. L., Storm, T. A., Pandey, K., Davis, C. R., et al. (2006). Fatality in mice due to oversaturation of cellular microRNA/short hairpin RNA pathways. Nature, 441(7092), 537–541.
Harper, S. Q., & Gonzalez-Alegre, P. (2008). Lentivirus-mediated RNA interference in mammalian neurons. Methods in Molecular Biology, 442, 95–112.
Hébert, S. S., Horré, K., Nicolaï, L., Papadopoulou, A. S., Mandemakers, W., Silahtaroglu, A. N., et al. (2008). Loss of microRNA cluster miR-29a/b-1 in sporadic Alzheimer’s disease correlates with increased BACE1/beta-secretase expression. Proceedings of the National Academy of Sciences of the United States of America, 105(17), 6415–6420.
Hol, E. M., van Leeuwen, F. W., & Fischer, D. F. (2005). The proteasome in Alzheimer’s disease and Parkinson’s disease: Lessons from ubiquitin B+1. Trends in Molecular Medicine, 11, 488–495.
Jeyaseelan, K., Lim, K. Y., & Armugam, A. (2008). MicroRNA expression in the blood and brain of rats subjected to transient focal ischemia by middle cerebral artery occlusion. Stroke, 39(3), 959–966.
Johnson, R., Zuccato, C., Belyaev, N. D., Guest, D. J., Cattaneo, E., & Buckley, N. J. (2008). A microRNA-based gene dysregulation pathway in Huntington’s disease. Neurobiology of Diseases, 29(3), 438–445.
Kapsimali, M., Kloosterman, W. P., de Bruijn, E., Rosa, F., Plasterk, R. H., & Wilson, S. W. (2007). MicroRNAs show a wide diversity of expression profiles in the developing and mature central nervous system. Genome Biology, 8(8), R173.
Keller, J. N., Hanni, K. B., & Markesbery, W. R. (2000). Impaired proteasome function in Alzheimer’s disease. Journal of Neurochemistry, 75, 436–439.
Kim, J., Inoue, K., Ishii, J., Vanti, W. B., Voronov, S. V., Murchison, E., et al. (2007). A MicroRNA feedback circuit in midbrain dopamine neurons. Science, 317(5842), 1220–1224.
Krichevsky, A. M., Sonntag, K. C., Isacson, O., & Kosik, K. S. (2006). Specific microRNAs modulate embryonic stem cell-derived neurogenesis. Stem Cells, 24, 857–864.
Krützfeldt, J., Rajewsky, N., Braich, R., Rajeev, K. G., Tuschl, T., Manoharan, M., et al. (2005). Silencing of microRNAs in vivo with ‘antagomirs’. Nature, 438(7068), 685–689.
Kumar, P., Wu, H., McBride, J. L., Jung, K. E., Kim, M. H., Davidson, B. L., et al. (2007). Transvascular delivery of small interfering RNA to the central nervous system. Nature, 448(7149), 39–43.
Lee, R. C., Feinbaum, R. L., & Ambros, V. (1993). The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell, 75, 843–854.
Lim, L. P., Lau, N. C., Garrett-Engele, P., Grimson, A., Schelter, J. M., Castle, J., et al. (2005). Microarray analysis shows that some microRNAs downregulate large numbers of target mRNAs. Nature, 433(7027), 769–773.
Lukiw, W. J., Zhao, Y., & Cui, J. G. (2008). An NF-kappaB-sensitive micro RNA-146a-mediated inflammatory circuit in Alzheimer disease and in stressed human brain cells. Journal of Biological Chemistry, 283(46), 31315–31322.
Makeyev, E. V., et al. (2007). The microRNA miR-124 promotes neuronal differentiation by triggering brain-specific alternative pre-mRNA splicing. Molecular Cell, 27, 435–448.
Mattson, M. P. (2004). Pathways towards and away from Alzheimer’s disease. Nature, 430(7000), 631–639.
Mattson, M. P., Maudsley, S., & Martin, B. (2004). BDNF and 5-HT: A dynamic duo in age-related neuronal plasticity and neurodegenerative disorders. Trends in Neurosciences, 27(10), 589–594.
McBride, J. L., Boudreau, R. L., Harper, S. Q., Staber, P. D., Monteys, A. M., Martins, I., et al. (2008). Artificial miRNAs mitigate shRNA-mediated toxicity in the brain: Implications for the therapeutic development of RNAi. Proceedings of the National Academy of Sciences of the United States of America, 105(15), 5868–5873.
Mellios, N., Huang, H. S., Grigorenko, A., Rogaev, E., & Akbarian, S. (2008). A set of differentially expressed miRNAs, including miR-30a–5p, act as post-transcriptional inhibitors of BDNF in prefrontal cortex. Human Molecular Genetics, 17(19), 3030–3042.
Miska, E. A., Alvarez-Saavedra, E., Townsend, M., Yoshii, A., Sestan, N., Rakic, P., et al. (2004). Microarray analysis of microRNA expression in the developing mammalian brain. Genome Biology, 5(9):R68.
Packer, A. N., Xing, Y., Harper, S. Q., Jones, L., & Davidson, B. L. (2008). The bifunctional microRNA miR-9/miR-9* regulates REST and CoREST and is downregulated in Huntington’s disease. Journal of Neuroscience, 28(53), 14341–14346.
Perkins, D. O., Jeffries, C. D., Jarskog, L. F., Thomson, J. M., Woods, K., Newman, M. A., et al. (2007). microRNA expression in the prefrontal cortex of individuals with schizophrenia and schizoaffective disorder. Genome Biology, 8(2), R27.
Redell, J. B., Liu, Y., & Dash, P. K. (2009). Traumatic brain injury alters expression of hippocampal microRNAs: Potential regulators of multiple pathophysiological processes. Journal of Neuroscience Research, 87(6), 1435–1448.
Ren, G., Li, T., Lan, J. Q., Wilz, A., Simon, R. P., & Boison, D. (2007). Lentiviral RNAi-induced downregulation of adenosine kinase in human mesenchymal stem cell grafts: A novel perspective for seizure control. Experimental Neurology, 208(1), 26–37.
Saba, R., Goodman, C. D., Huzarewich, R. L., Robertson, C., & Booth, S. A. (2008). A miRNA signature of prion induced neurodegeneration. PLoS ONE, 3(11), e3652.
Salehi, A., Delcroix, J. D., Belichenko, P. V., Zhan, K., Wu, C., Valletta, J. S., et al. (2006). Increased App expression in a mouse model of Down’s syndrome disrupts NGF transport and causes cholinergic neuron degeneration. Neuron, 51, 29–42.
Sapru, M. K., Yates, J. W., Hogan, S., Jiang, L., Halter, J., & Bohn, M. C. (2006). Silencing of human alpha-synuclein in vitro and in rat brain using lentiviral-mediated RNAi. Experimental Neurology, 198, 382–390.
Schwamborn, J. C., Berezikov, E., & Knoblich, J. A. (2009). The TRIM-NHL protein TRIM32 activates microRNAs and prevents self-renewal in mouse neural progenitors. Cell, 136(5), 913–925.
Selbach, M., Schwanhäusser, B., Thierfelder, N., Fang, Z., Khanin, R., & Rajewsky, N. (2008). Widespread changes in protein synthesis induced by microRNAs. Nature, 455(7209), 58–63.
Sempere, L. F., Freemantle, S., Pitha-Rowe, I., Moss, E., Dmitrovsky, E., & Ambros, V. (2004). Expression profiling of mammalian microRNAs uncovers a subset of brain-expressed microRNAs with possible roles in murine and human neuronal differentiation. Genome Biology, 5(3):R13.
Sheedy, F. J., & O’Neill, L. A. (2008). Adding fuel to fire: microRNAs as a new class of mediators of inflammation. Annals of the Rheumatic Diseases, 67(Suppl 3), iii50–iii55.
Silahtaroglu, A. N., Nolting, D., Dyrskjøt, L., Berezikov, E., Møller, M., Tommerup, N., et al. (2007). Detection of microRNAs in frozen tissue sections by fluorescence in situ hybridization using locked nucleic acid probes and tyramide signal amplification. Nature Protocols, 2(10), 2520–2528.
Singer, O., Marr, R. A., Rockenstein, E., Crews, L., Coufal, N. G., Gage, F. H., et al. (2005). Targeting BACE1 with siRNAs ameliorates Alzheimer disease neuropathology in a transgenic model. Nature Neuroscience, 8(10), 1343–1349.
Singleton, A. B., Farrer, M., Johnson, J., Singleton, A., Hague, S., Kachergus, J., et al. (2003). alpha-Synuclein locus triplication causes Parkinson’s disease. Science, 302, 841.
Singleton, A., Myers, A., & Hardy, J. (2004). The law of mass action applied to neurodegenerative disease: A hypothesis concerning the etiology and pathogenesis of complex diseases. Human and Molecular Genetics, 13(Spec No 1), R123–R126.
Smirnova, L., Gräfe, A., Seiler, A., Schumacher, S., Nitsch, R., & Wulczyn, F. G. (2005). Regulation of miRNA expression during neural cell specification. European Journal of Neuroscience, 21(6), 1469–1477.
Tsang, J., Zhu, J., & van Oudenaarden, A. (2007). MicroRNA-mediated feedback and feedforward loops are recurrent network motifs in mammals. Molecular Cell, 26(5), 753–767.
Urbich, C., Kuehbacher, A., & Dimmeler, S. (2008). Role of microRNAs in vascular diseases, inflammation, and angiogenesis. Cardiovascular Research, 79(4), 581–588.
van Solingen, C., Seghers, L., Bijkerk, R., Duijs, J. M., Roeten, M. K., van Oeveren-Rietdijk, A. M., et al. (2008). Antagomir-mediated silencing of endothelial cell specific microRNA-126 impairs ischemia-induced angiogenesis. Journal of Cellular and Molecular Medicine. 2008 Dec 16. [Epub ahead of print].
Várallyay, E., Burgyán, J., & Havelda, Z. (2008). MicroRNA detection by northern blotting using locked nucleic acid probes. Nature Protocols, 3(2), 190–196.
Wang, W. X., Rajeev, B. W., Stromberg, A. J., Ren, N., Tang, G., Huang, Q., et al. (2008a). The expression of microRNA miR-107 decreases early in Alzheimer’s disease and may accelerate disease progression through regulation of beta-site amyloid precursor protein-cleaving enzyme 1. Journal of Neuroscience, 28, 1213–1223.
Wang, G., van der Walt, J. M., Mayhew, G., Li, Y. J., Züchner, S., Scott, W. K., et al. (2008b). Variation in the miRNA-433 binding site of FGF20 confers risk for Parkinson disease by overexpression of alpha-synuclein. American Journal of Human Genetics, 82, 283–289.
Zhao, C., Deng, W., & Gage, F. H. (2008). Mechanisms and functional implications of adult neurogenesis. Cell, 132(4), 645–660.
Zhao, C., Sun, G., Li, S., & Shi, Y. (2009). A feedback regulatory loop involving microRNA-9 and nuclear receptor TLX in neural stem cell fate determination. Nature Structural & Molecular Biology, 16(4), 365–371.
Zhou, R., Yuan. P., Wang, Y., Hunsberger, J. G., Elkahloun, A., Wei, Y., et al. (2009). Evidence for selective microRNAs and their effectors as common long-term targets for the actions of mood stabilizers. Neuropsychopharmacology, 34, 1395–1405.
Zuccato, C., Belyaev, N., Conforti, P., Ooi, L., Tartari, M., Papadimou, E., et al. (2007). Widespread disruption of repressor element-1 silencing transcription factor/neuron-restrictive silencer factor occupancy at its target genes in Huntington's disease. Journal of Neuroscience, 27, 6972–6983.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Hutchison, E.R., Okun, E. & Mattson, M.P. The Therapeutic Potential of microRNAs in Nervous System Damage, Degeneration, and Repair. Neuromol Med 11, 153–161 (2009). https://doi.org/10.1007/s12017-009-8086-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12017-009-8086-x