Abstract
Objectives
This study aimed to explore the regulatory mechanism of methyltransferase3 (METTL3) -mediated long non-coding RNA (lncRNA) N6-methyladenosine (m6A) modification in the osteogenic differentiation of human adipose-derived stem cells (hASCs) induced by NEL-like 1 protein (NELL-1).
Materials and Methods
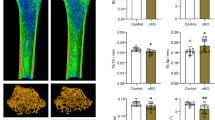
Methylated RNA immunoprecipitation sequencing (MeRIP-seq) and high- throughput sequencing for RNA (RNA-seq) were performed on hASCs. Osteogenic ability was detected by alkaline phosphatase (ALP) staining, Alizarin Red S(ARS) staining, ALP quantification and Quantitative real-time polymerase chain reaction analysis (qRT-PCR). Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis predicted the osteogenesis-related pathways enriched for the lncRNAs and identified the target lncRNAs. After overexpression and knockdown of METTL3, methylated RNA immunoprecipitation-qPCR (MeRIP-qPCR) and qRT-PCR were used to detect the levels of m6A modification and the expression of the target lncRNA, and the binding of both was confirmed by RNA binding protein immunoprecipitation (RIP) assay. The effects of lncRNA and METTL3 on phosphorylation of the key proteins of the pathway were detected by western blot analysis.
Results
In vitro experiments showed that METTL3 can promote osteogenic differentiation and that its expression level is upregulated. KEGG pathway analysis predicted that lncRNAs with differentially upregulated methylated peaks were enriched mostly in the mitogen-activated protein kinase (MAPK) signaling pathway, in which Serine/threonine protein kinase 3 (STK3) was the predicted target gene of the lncRNA RP11-44 N12.5. The m6A modification and expression of RP11-44 N12.5 were both regulated by METTL3. Subsequently, lncRNA RP11-44 N12.5 and METTL3 were found to regulate the phosphorylation levels of three key proteins in the MAPK signaling pathway, ERK, JNK and p38.
Conclusions
This study shows, for the first time, that METTL3 can activate the MAPK signaling pathway by regulating the m6A modification and expression of a lncRNA, thereby enhancing the osteogenic differentiation of hASCs.
Graphical abstract






Similar content being viewed by others
Data Availability
All data generated or analysed during this study are included in this article and its supplementary materials.
Change history
16 October 2021
A Correction to this paper has been published: https://doi.org/10.1007/s12015-021-10283-y
Abbreviations
- lncRNA:
-
long non-coding RNA
- m6A:
-
N6-methyladenosine
- hASCs:
-
human adipose-derived stem cells
- NELL-1:
-
NEL-like 1 protein
- BMSCs:
-
bone marrow-derived mesenchymal stem cells
- TGF:
-
transforming growth factor
- RUNX2:
-
Runt-related transcription factor 2
- BSP:
-
bone sialoprotein
- METTL14:
-
methyltransferase 14
- METTL3:
-
methyltransferase 3
- FTO:
-
fat-mass and obesity-associated protein
- ALKBH5:
-
ALKB homologue 5
- ERK:
-
extracellular signal-regulated kinase
- MAPK:
-
mitogen-activated protein kinase
- JNK:
-
c-Jun N-terminal kinase
- STK3:
-
Serine/threonine protein kinase 3
- GCK II:
-
germinal centre kinase II
- ALP:
-
alkaline phosphatase
- ARS:
-
alizarin red
- qRT-PCR:
-
quantitative real-time polymerase chain reaction analysis
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- RIP:
-
RNA binding protein immunoprecipitation
- NFIC:
-
nuclear factor I-C
- MeRIP-seq:
-
methylated RNA immunoprecipitation sequencing
- MeRIP-qPCR:
-
methylated RNA immunoprecipitation-qPCR
- RNA-seq:
-
high- throughput sequencing for RNA
- ncRNAs:
-
noncoding RNAs
- dhASC:
-
differentiated hASC
- NELL1-dhASC:
-
rhNELL-1-induced osteogenic differentiated hASC
- IPP:
-
immunoprecipitation
- DREME:
-
Discriminative Regular Expression Motif Elicitation
- PBS:
-
phosphate buffered saline
- sodium dodecyl sulfate-polyacrylamide gel electrophoresis:
-
SDS-PAGE
References
Zhang, Z. (2011). Bone regeneration by stem cell and tissue engineering in oral and maxillofacial region. Frontiers in Medicine, 5(4), 401–413.
Eμler de Souza Lucena, E., et al. (2014). Experimental considerations concerning the use of stem cells and tissue engineering for facial nerve regeneration: a systematic review. J Oral Maxillofac Surg, 72(5), 1001–1012.
Chun, S. Y., Lim, J. O., Lee, E. H., Han, M. H., Ha, Y. S., Lee, J. N., Kim, B. S., Park, M. J., Yeo, M. G., Jung, B., & Kwon, T. G. (2019). Preparation and characterization of human adipose tissue-derived Extracellμlar matrix, growth factors, and stem cells: A concise review. Tissue Eng Regen Med, 16(4), 385–393.
Rivera-Izquierdo, M., Cabeza, L., Láinez-Ramos-Bossini, A., Quesada, R., Perazzoli, G., Alvarez, P., Prados, J., & Melguizo, C. (2019). An updated review of adipose derived-mesenchymal stem cells and their applications in muscμloskeletal disorders. Expert Opinion on Biological Therapy, 19(3), 233–248.
Smieszek, A., et al. (2018). Metformin Promotes Osteogenic Differentiation of Adipose-Derived Stromal Cells and Exerts Pro-Osteogenic Effect Stimμlating Bone Regeneration. Journal of Clinical Medicine, 7(12), 482.
Mohamed-Ahmed, S., Yassin, M. A., Rashad, A., Espedal, H., Idris, S. B., Finne-Wistrand, A., Mustafa, K., Vindenes, H., & Fristad, I. (2021). Comparison of bone regenerative capacity of donor-matched human adipose-derived and bone marrow mesenchymal stem cells. Cell and Tissue Research, 383(3), 1061–1075.
Yoshida, Y., Matsubara, H., Fang, X., Hayashi, K., Nomura, I., Ugaji, S., Hamada, T., & Tsuchiya, H. (2019). Adipose-derived stem cell sheets accelerate bone healing in rat femoral defects. PLoS One, 14(3), e0214488.
Li, W., Zara, J. N., Siu, R. K., Lee, M., Aghaloo, T., Zhang, X., Wu, B. M., Gertzman, A. A., Ting, K., & Soo, C. (2011). Nell-1 enhances bone regeneration in a rat critical-sized femoral segmental defect model. Plastic and Reconstructive Surgery, 127(2), 580–587.
James, A. W., Shen, J., Zhang, X., Asatrian, G., Goyal, R., Kwak, J. H., Jiang, L., Bengs, B., Culiat, C. T., Turner, A. S., Seim III, H. B., Wu, B. M., Lyons, K., Adams, J. S., Ting, K., & Soo, C. (2015). NELL-1 in the treatment of osteoporotic bone loss. Nature Communications, 6, 7362.
Wang, J., Liao, J., Zhang, F., Song, D., Lu, M., Liu, J., Wei, Q., Tang, S., Liu, H., Fan, J., Zou, Y., Guo, D., Huang, J., Liu, F., Ma, C., Hu, X., Li, L., Qu, X., Chen, L., et al. (2017). NEL-like Molecμle-1 (Nell1) is regulated by bone morphogenetic protein 9 (BMP9) and potentiates BMP9-induced osteogenic differentiation at the expense of Adipogenesis in mesenchymal stem cells. Cellular Physiology and Biochemistry, 41(2), 484–500.
Yu, L., Cen, X., Xia, K., Huang, X., Sun, W., Zhao, Z., & Liu, J. (2020). microRNA expression profiles and the potential competing endogenous RNA networks in NELL-1-induced human adipose-derived stem cell osteogenic differentiation. Journal of Cellular Biochemistry, 121(11), 4623–4641.
Huang, X., Cen, X., Zhang, B., Liao, Y., Zhao, Z., Zhu, G., Zhao, Z., & Liu, J. (2019). The roles of circRFWD2 and circINO80 during NELL-1-induced osteogenesis. Journal of Cellular and Molecular Medicine, 23(12), 8432–8441.
Rathinam, E., Rajasekharan, S., Chitturi, R. T., Martens, L., & de Coster, P. (2015). Gene expression profiling and Molecμlar signaling of dental Pμlp cells in response to Tricalcium silicate cements: A systematic review. Journal of Endodontia, 41(11), 1805–1817.
Boμleftour, W., et al. (2017). Bone shaft Revascμlarization after marrow ablation is dramatically accelerated in BSP−/− mice, along with faster hematopoietic recolonization. Journal of Cellular Physiology, 232(9), 2528–2537.
Vu, L. P., Cheng, Y., & Kharas, M. G. (2019). The biology of m(6)a RNA methylation in Normal and malignant hematopoiesis. Cancer Discovery, 9(1), 25–33.
Wang, P., Doxtader, K. A., & Nam, Y. (2016). Structural basis for cooperative function of Mettl3 and Mettl14 methyltransferases. Molecular Cell, 63(2), 306–317.
Shen, L., Song, C. X., He, C., & Zhang, Y. (2014). Mechanism and function of oxidative reversal of DNA and RNA methylation. Annual Review of Biochemistry, 83, 585–614.
Pinello, N., Sun, S., & Wong, J. J. (2018). Aberrant expression of enzymes regulating m(6)a mRNA methylation: Implication in cancer. Cancer Biology & Medicine, 15(4), 323–334.
Zhong, X., et al. (2018). Circadian Clock Regulation of Hepatic Lipid Metabolism by Modμlation of m(6)A mRNA Methylation. Cell Report, 25(7), 1816–1828.e4.
Sweatt, J. D. (2017). Layered-up regulation in the developing brain. Nature, 551(7681), 448–449.
Zhao, B. S., Wang, X., Beadell, A. V., Lu, Z., Shi, H., Kuuspalu, A., Ho, R. K., & He, C. (2017). M(6)A-dependent maternal mRNA clearance facilitates zebrafish maternal-to-zygotic transition. Nature, 542(7642), 475–478.
Geμla, S., et al. (2015). Stem cells. m6A mRNA methylation facilitates resolution of naïve pluripotency toward differentiation. Science, 347(6225), 1002–1006.
Yan, G., Yuan, Y., He, M., Gong, R., Lei, H., Zhou, H., Wang, W., du, W., Ma, T., Liu, S., Xu, Z., Gao, M., Yu, M., Bian, Y., Pang, P., Li, X., Yu, S., Yang, F., Cai, B., & Yang, L. (2020). M(6)a methylation of precursor-miR-320/RUNX2 controls osteogenic potential of bone marrow-derived mesenchymal stem cells. Molecular Therapy--Nucleic Acids, 19, 421–436.
Yao, Y., Bi, Z., Wu, R., Zhao, Y., Liu, Y., Liu, Q., Wang, Y., & Wang, X. (2019). METTL3 inhibits BMSC adipogenic differentiation by targeting the JAK1/STAT5/C/EBPβ pathway via an m(6)A-YTHDF2-dependent manner. The FASEB Journal, 33(6), 7529–7544.
Zhang, Q., Riddle, R. C., Yang, Q., Rosen, C. R., Guttridge, D. C., Dirckx, N., Faugere, M. C., Farber, C. R., & Clemens, T. L. (2019). The RNA demethylase FTO is required for maintenance of bone mass and functions to protect osteoblasts from genotoxic damage. Proceedings of the National Academy of Sciences of the United States of America, 116(36), 17980–17989.
Beermann, J., Piccoli, M. T., Viereck, J., & Thum, T. (2016). Non-coding RNAs in development and disease: Background, mechanisms, and therapeutic approaches. Physiological Reviews, 96(4), 1297–1325.
Shen, J. J., Zhang, C. H., Chen, Z. W., Wang, Z. X., Yang, D. C., Zhang, F. L., & Feng, K. H. (2019). LncRNA HOTAIR inhibited osteogenic differentiation of BMSCs by regulating Wnt/β-catenin pathway. European Review for Medical and Pharmacological Sciences, 23(17), 7232–7246.
Zheng, S., Wang, Y. B., Yang, Y. L., Chen, B. P., Wang, C. X., Li, R. H., & Huang, D. (2019). LncRNA MALAT1 inhibits osteogenic differentiation of mesenchymal stem cells in osteoporosis rats through MAPK signaling pathway. European Review for Medical and Pharmacological Sciences, 23(11), 4609–4617.
Liu, R., Li, Z., Song, E., Hu, P., Yang, Q., Hu, Y., Liu, H., & Jin, A. (2020). LncRNA HOTTIP enhances human osteogenic BMSCs differentiation via interaction with WDR5 and activation of Wnt/β-catenin signaling pathway. Biochemical and Biophysical Research Communications, 524(4), 1037–1043.
Xia, K., Cen, X., Yu, L., Huang, X., Sun, W., Zhao, Z., & Liu, J. (2020). Long noncoding RNA expression profiles during the NEL-like 1 protein-induced osteogenic differentiation. Journal of Cellular Physiology, 235(9), 6010–6022.
Xin, B. C., Wu, Q. S., Jin, S., Luo, A. H., Sun, D. G., & Wang, F. (2020). Berberine promotes osteogenic differentiation of human dental Pμlp stem cells through activating EGFR-MAPK-Runx2 pathways. Pathology Oncology Research, 26(3), 1677–1685.
Tsang, E. J., Wu, B., & Zuk, P. (2018). MAPK signaling has stage-dependent osteogenic effects on human adipose-derived stem cells in vitro. Connective Tissue Research, 59(2), 129–146.
James, A. W., Pan, A., Chiang, M., Zara, J. N., Zhang, X., Ting, K., & Soo, C. (2011). A new function of Nell-1 protein in repressing adipogenic differentiation. Biochemical and Biophysical Research Communications, 411(1), 126–131.
Zhang, Y., Liu, T., Meyer, C. A., Eeckhoute, J., Johnson, D. S., Bernstein, B. E., Nussbaum, C., Myers, R. M., Brown, M., Li, W., & Liu, X. S. (2008). Model-based analysis of ChIP-Seq (MACS). Genome Biology, 9(9), R137.
Meyer, K. D., Saletore, Y., Zumbo, P., Elemento, O., Mason, C. E., & Jaffrey, S. R. (2012). Comprehensive analysis of mRNA methylation reveals enrichment in 3' UTRs and near stop codons. Cell, 149(7), 1635–1646.
Dominissini, D., Moshitch-Moshkovitz, S., Schwartz, S., Salmon-Divon, M., Ungar, L., Osenberg, S., Cesarkas, K., Jacob-Hirsch, J., Amariglio, N., Kupiec, M., Sorek, R., & Rechavi, G. (2012). Topology of the human and mouse m6A RNA methylomes revealed by m6A-seq. Nature, 485(7397), 201–206.
Pan, Y., Ma, P., Liu, Y., Li, W., & Shu, Y. (2018). Mμltiple functions of m(6)a RNA methylation in cancer. Journal of Hematology & Oncology, 11(1), 48.
Pan, T. (2013). N6-methyl-adenosine modification in messenger and long non-coding RNA. Trends in Biochemical Sciences, 38(4), 204–209.
Niu, Y., Zhao, X., Wu, Y. S., Li, M. M., Wang, X. J., & Yang, Y. G. (2013). N6-methyl-adenosine (m6A) in RNA: An old modification with a novel epigenetic function. Genomics, Proteomics & Bioinformatics, 11(1), 8–17.
Wang, H. F., Kuang, M. J., Han, S. J., Wang, A. B., Qiu, J., Wang, F., Tan, B. Y., & Wang, D. C. (2020). BMP2 modified by the m(6)a demethylation enzyme ALKBH5 in the ossification of the ligamentum Flavum through the AKT signaling pathway. Calcified Tissue International, 106(5), 486–493.
Sun, Z., Wang, H., Wang, Y., Yuan, G., Yu, X., Jiang, H., Wu, Q., Yang, B., Hu, Z., Shi, F., Cao, X., Zhang, S., Guo, T., & Zhao, J. (2021). MiR-103-3p targets the m(6) a methyltransferase METTL14 to inhibit osteoblastic bone formation. Aging Cell, 20(2), e13298.
Wu, Y., Xie, L., Wang, M., Xiong, Q., Guo, Y., Liang, Y., Li, J., Sheng, R., Deng, P., Wang, Y., Zheng, R., Jiang, Y., Ye, L., Chen, Q., Zhou, X., Lin, S., & Yuan, Q. (2018). Mettl3-mediated m(6)a RNA methylation regulates the fate of bone marrow mesenchymal stem cells and osteoporosis. Nature Communications, 9(1), 4772.
Liu, H., Xu, Y., Yao, B., Sui, T., Lai, L., & Li, Z. (2020). A novel N6-methyladenosine (m6A)-dependent fate decision for the lncRNA THOR. Cell Death & Disease, 11(8), 613.
Wu, Y., et al. (2019). m(6)A-induced lncRNA RP11 triggers the dissemination of colorectal cancer cells via upregulation of Zeb1. Molecular Cancer, 18(1), 87.
Ni, W., Yao, S., Zhou, Y., Liu, Y., Huang, P., Zhou, A., Liu, J., Che, L., & Li, J. (2019). Long noncoding RNA GAS5 inhibits progression of colorectal cancer by interacting with and triggering YAP phosphorylation and degradation and is negatively regulated by the m(6)a reader YTHDF3. Molecular Cancer, 18(1), 143.
Coker, H., Wei, G., & Brockdorff, N. (2019). m6A modification of non-coding RNA and the control of mammalian gene expression. Biochimica et Biophysica Acta, Gene Regulatory Mechanisms, 1862(3), 310–318.
Wang, Y., Lu, J. H., Wu, Q. N., Jin, Y., Wang, D. S., Chen, Y. X., Liu, J., Luo, X. J., Meng, Q., Pu, H. Y., Wang, Y. N., Hu, P. S., Liu, Z. X., Zeng, Z. L., Zhao, Q., Deng, R., Zhu, X. F., Ju, H. Q., & Xu, R. H. (2019). LncRNA LINRIS stabilizes IGF2BP2 and promotes the aerobic glycolysis in colorectal cancer. Molecular Cancer, 18(1), 174.
Lu, Z., Liu, J., Yuan, C., Jin, M., Quan, K., Chu, M., & Wei, C. (2021). M(6)a mRNA methylation analysis provides novel insights into heat stress responses in the liver tissue of sheep. Genomics, 113(1 Pt 2), 484–492.
Zhou, Y., Hambly, B. D., Simmons, D., & McLachlan, C. S. (2020). RUNX1T1 rs34269950 is associated with obesity and metabolic syndrome. Qjm.
Tian, C., et al. (2019). Mettl3 Regulates Osteogenic Differentiation and Alternative Splicing of Vegfa in Bone Marrow Mesenchymal Stem Cells. International Journal of Molecular Sciences, 20(3).
Sheng, R., Wang, Y., Wu, Y., Wang, J., Zhang, S., Li, Q., Zhang, D., Qi, X., Xiao, Q., Jiang, S., & Yuan, Q. (2021). METTL3-mediated m(6) a mRNA methylation Modμlates tooth root formation by affecting NFIC translation. Journal of Bone and Mineral Research, 36(2), 412–423.
Zhou, T., Han, D., Liu, J., Shi, J., Zhu, P., Wang, Y., & Dong, N. (2021). Factors influencing osteogenic differentiation of human aortic valve interstitial cells. The Journal of Thoracic and Cardiovascular Surgery, 161(2), e163–e185.
Li, D., et al. (2020). METTL3 Modμlates Osteoclast Differentiation and Function by Controlling RNA Stability and Nuclear Export. International Journal of Molecular Sciences, 21(5), 1660.
Yu, J., Shen, L., Liu, Y., Ming, H., Zhu, X., Chu, M., & Lin, J. (2020). The m6A methyltransferase METTL3 cooperates with demethylase ALKBH5 to regulate osteogenic differentiation through NF-κB signaling. Molecular and Cellular Biochemistry, 463(1–2), 203–210.
Lawrence, M. C., Jivan, A., Shao, C., Duan, L., Goad, D., Zaganjor, E., Osborne, J., McGlynn, K., Stippec, S., Earnest, S., Chen, W., & Cobb, M. H. (2008). The roles of MAPKs in disease. Cell Research, 18(4), 436–442.
Sun, Y., Liu, W. Z., Liu, T., Feng, X., Yang, N., & Zhou, H. F. (2015). Signaling pathway of MAPK/ERK in cell proliferation, differentiation, migration, senescence and apoptosis. Journal of Receptor and Signal Transduction Research, 35(6), 600–604.
Raman, M., Chen, W., & Cobb, M. H. (2007). Differential regulation and properties of MAPKs. Oncogene, 26(22), 3100–3112.
Zhu, J. H., Liao, Y. P., Li, F. S., Hu, Y., Li, Q., Ma, Y., Wang, H., Zhou, Y., He, B. C., & Su, Y. X. (2018). Wnt11 promotes BMP9-induced osteogenic differentiation through BMPs/Smads and p38 MAPK in mesenchymal stem cells. Journal of Cellular Biochemistry, 119(11), 9462–9473.
Matsushita, T., Chan, Y. Y., Kawanami, A., Balmes, G., Landreth, G. E., & Murakami, S. (2009). Extracellμlar signal-regulated kinase 1 (ERK1) and ERK2 play essential roles in osteoblast differentiation and in supporting osteoclastogenesis. Molecular and Cellular Biology, 29(21), 5843–5857.
Zhang, X., Zhou, C., Zha, X., Xu, Z., Li, L., Liu, Y., Xu, L., Cui, L., Xu, D., & Zhu, B. (2015). Apigenin promotes osteogenic differentiation of human mesenchymal stem cells through JNK and p38 MAPK pathways. Molecular and Cellular Biochemistry, 407(1–2), 41–50.
Zhang, Y., et al. (2019). METTL3 Regulates Osteoblast Differentiation and Inflammatory Response via Smad Signaling and MAPK Signaling. International Journal of Molecular Sciences, 21(1), 199.
Acknowledgements
This study was supported by grants from the National Natural Science Foundation of China (81870743, 82170934 and 81771048). And we appreciated Cloud-seq Company (Shanghai, China) that provided MeRIP-Seq and lncRNA sequencing service.
Code Availability
Not applicable.
Funding
National Natural Science Foundation of China (81870743, 82170934 and 81771048).
Author information
Authors and Affiliations
Contributions
Yidan Song and Yihua Pan mainly carried out the experimental operations. Yidan Song, Mengsong Wu, Wentian Sun and Liangyu Luo analyzed and interpreted the data. Yidan Song wrote the manuscript. Jun Liu and Zhihe Zhao designed the experiments and revised the manuscript.
Corresponding author
Ethics declarations
Conflicts of Interest
None.
Ethics Approval
Not applicable.
Consent to Participate
All authors read and approved the final manuscript.
Consent for Publication
All authors agreed to publish.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
ESM 1
(PNG 25 kb)
ESM 2
(PNG 35 kb)
ESM 3
(PNG 34 kb)
ESM 4
(PNG 36 kb)
ESM 5
(PNG 36 kb)
ESM 6
(PNG 36 kb)
ESM 7
(PNG 32 kb)
ESM 8
(PNG 39 kb)
ESM 9
(PNG 39 kb)
ESM 10
(PNG 29 kb)
ESM 11
(PNG 29 kb)
ESM 12
(PNG 32 kb)
ESM 13
(PNG 589 kb)
ESM 14
(PNG 456 kb)
ESM 15
(PNG 596 kb)
ESM 16
(PNG 520 kb)
ESM 17
(PNG 510 kb)
ESM 18
(PNG 501 kb)
ESM 19
(PNG 512 kb)
ESM 20
(PNG 405 kb)
ESM 21
(PNG 474 kb)
ESM 22
(PNG 504 kb)
ESM 23
(PNG 527 kb)
ESM 24
(PNG 503 kb)
ESM 25
(PNG 522 kb)
ESM 26
(PNG 525 kb)
ESM 27
(PNG 420 kb)
ESM 28
(PNG 544 kb)
ESM 29
(PNG 510 kb)
ESM 30
(PNG 436 kb)
ESM 31
(PNG 506 kb)
ESM 32
(PNG 502 kb)
ESM 33
(PNG 438 kb)
ESM 34
(PNG 479 kb)
ESM 35
(PNG 493 kb)
ESM 36
(PNG 411 kb)
ESM 37
(PNG 472 kb)
ESM 38
(PNG 533 kb)
ESM 39
(PNG 509 kb)
ESM 40
(PNG 511 kb)
ESM 41
(PNG 516 kb)
ESM 42
(PNG 59 kb)
ESM 43
(PNG 474 kb)
ESM 44
(DOCX 14 kb)
ESM 45
(DOCX 15 kb)
ESM 46
(DOCX 14 kb)
Rights and permissions
About this article
Cite this article
Song, Y., Pan, Y., Wu, M. et al. METTL3-Mediated lncRNA m6A Modification in the Osteogenic Differentiation of Human Adipose-Derived Stem Cells Induced by NEL-Like 1 Protein. Stem Cell Rev and Rep 17, 2276–2290 (2021). https://doi.org/10.1007/s12015-021-10245-4
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12015-021-10245-4