Abstract
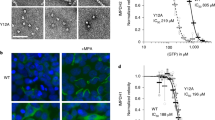
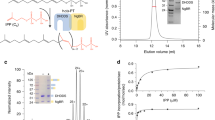
Defects in inosine monophosphate dehydrogenase-1 (IMPDH1) lead to insufficient biosyntheses of purine nucleotides. In eyes, these defects are believed to cause retinitis pigmentosa (RP). Major retinal isoforms of IMPDH1 are structurally distinct from those in other tissues, by bearing terminal extensions. Using recombinant mouse IMPDH1 (mH1), we evaluated the kinetics and oligomerization states of the retinal isoforms. Moreover, we adopted molecular simulation tools to study the possible effect of terminal tails on the function of major enzyme isoforms with the aim to find structural evidence in favor of contradictory observations on retinal IMPDH1 function. Our findings indicated higher catalytic activity for the major mouse retinal isoform (mH1603) along with lower fibrillation capacity under the influence of ATP. However, higher mass oligomerization products were formed by the mH1 (603) isoform in the presence of the enzyme inhibitors such as GTP and/or MPA. Collectively, our findings demonstrate that the structural differences between the retinal isoforms have led to functional variations possibly to justify the retinal cells’ requirements.






Similar content being viewed by others
References
Bowne, S., Sullivan, S., Blanton, S., Cepko, C. L., Blackshaw, S., Birch, D. G., Hughbanks-Wheaton, D., Heckenlively, J. R. & Daiger, S. P. (2002). Mutations in the inosine monophosphate dehydrogenase 1 gene (IMPDH1) cause the RP10 form of autosomal dominant retinitis pigmentosa. Human Molecular Genetics, 11(5), 559–568.
Sohocki, M. M., Daiger, S. P., Bowne, S., Rodriquez, J. A., Northrup, H., Heckenlively, J. R., Birch, D. G., Mintz-Hittner, H., Ruiz, R. S., Lewis, R. A., Saperstein, D. A. & Sullivan, L. S. (2001). Prevalence of mutations causing retinitis pigmentosa and other inherited retinopathies. Human Mutation, 17(1), 42–51.
Spellicy, C. J., Daiger, S. P., Sullivan, L. S., Zhu, J., Liu, Q., Pierce, E. A. & Bowne, S. J. (2007). Characterization of retinal inosine monophosphate dehydrogenase 1 in several mammalian species. Molecular Vision, 13, 1866–1872.
Gunter, J. H., Thomas, E. C., Lengefeld, N., Kruger, S. J., Worton, L., Gardiner, E. M., Jones, A., Barnett, N. L. & Whitehead, J. P. (2008). Characterisation of inosine monophosphate dehydrogenase expression during retinal development: differences between variants and isoforms. The International Journal of Biochemistry & Cell Biology, 40(9), 1716–1728.
Bateman, A. (1997). The structure of a domain common to archaebacteria and the homocystinuria disease protein. Trends in Biochemical Sciences, 22(1), 12–13.
Colby, T. D., Vanderveen, K., Strickler, M. D., Markham, G. D. & Goldstein, B. M. (1999). Crystal structure of human type II inosine monophosphate dehydrogenase: implications for ligand binding and drug design. Proceedings of the National Academy of Sciences of the United States of America, 96(7), 3531–3536.
Labesse, G., Alexandre, T., Vaupre, L., Salard-Arnaud, I., Him, J. L., Raynal, B., Bron, P., & Munier- Lehmann, H. (2013). MgATP regulates allostery and fiber formation in IMPDHs. Structure, 21(6), 975–985.
Buey, R. M., Ledesma-Amaro, R., Velazquez-Campoy, A., Balsera, M., Chagoyen, M., de Pereda, J. M. & & Revuelta, J. L. (2015). Guanine nucleotide binding to the Bateman domain mediates the allosteric inhibition of eukaryotic IMP dehydrogenases. Nature Communications, 6, 8923.
Andashti, B., Yazdanparast, R., Barzegari, E., & Galehdari, H. (2020). The functional impact of the C/N‑terminal extensions of the mouse retinal IMPDH1 isoforms: a kinetic evaluation. Molecular and Cellular Biochemistry, 465, 155–164.
Dwivedi, P., Rodriguez, J., Ibe, N. U., & Weers, P. M. (2016). Deletion of the N- or C‑terminal helix of apolipophorin III to create a four-helix bundle protein. Biochemistry, 55(26), 3607–15. https://doi.org/10.1021/acs.biochem.6b00381.
Dixit, G., Sahu, I. D., Rynolds, W. D., Wadsworth, T. M., Harding, B. D., & Jaycox, C. K., et al. (2019). Probing the dynamics and structural topology of the reconstituted human KCNQ1 voltage sensor domain (Q1-VSD) in lipid bilayers using electron paramagnetic resonance spectroscopy. Biochemistry., 58(7), 965–973. https://doi.org/10.1021/acs.biochem.8b01042.
Herudek J. Structural and functional characterization of RNA-binding domain in human IMPDH proteins, Faculty of Science. Masaryk University, Central European Institute of Technology. 2013; 91.
Pettersen, E. F., Goddard, T. D., Huang, C. C., Couch, G. S., Greenblatt, D. M., Meng, E. C. & Ferrin, T. E. (2004). UCSF Chimera–a visualization system for exploratory research and analysis. Journal of Computational Chemistry, 25(13), 1605–1612.
Sintchak, M. D., Fleming, M. A., Futer, O., Raybuck, S. A., Chambers, S. P., Caron, P. R., Murcko, M. A., & Wilson, K. P. (1996). Structure and mechanism of inosine monophosphate dehydrogenase in complex with the immunosuppressant mycophenolic acid. Cell, 85(6), 921–930.
Trott, O. & Olson, A. J. (2010). AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization and multithreading. Journal of Computational Chemistry, 31, 455–461.
Van Zundert, G. C., Rodrigues, J. P., Trellet, M., Schmitz, C., Kastritis, P. L., Karaca, E., Melquiond, A. S., van Dijk, M., de Vries, S. J., & Bonvin, A. M. (2016). The HADDOCK2.2 web server: user1 friendly integrative modeling of biomolecular complexes. Journal of Molecular Biology, 428(4), 720–725.
Wassenaar, T. A., van Dijk, M., Loureiro-Ferreira, N., van der Schot, G., de Vries, S. J., Schmitz, C., van der Zwan, J., Boelens, R., Giachetti, A., & Ferella, L., et al. (2012). WeNMR: structural biology on the grid. J. Grid Computing, 10(4), 743–767.
De Vries, S. J., van Dijk, M. & Bonvin, A. M. (2010). The HADDOCK web server for data-driven biomolecular docking. Nature Protocols, 5(5), 883–897.
Moreira, I. S., Fernandes, P. A. & Ramos, M. J. (2010). Protein-protein docking dealing with the unknown. Journal of Computational Chemistry, 31(2), 317–342.
Risal D, Strickler MD, Goldstein BM. Crystal structure of the human type I inosine monophosphate dehydrogenase and implications for isoform specificity. 2003.
Porollo, A. A., Adamczak, R., & Meller, J. (2004). POLYVIEW: a flexible visualization tool for structural and functional annotations of proteins. Bioinformatics, 20(15), 2460–2462.
Eisenberg, D., Schwarz, E., Komarony, M., & Wall, R. (1984). Analysis of membrane and surface protein sequences with the hydrophobic moment plot. Journal of Molecular Biology, 179, 125–142.
Xu, D., Cobb, G., Spellicy, C. J., Bowne, S. J., Daiger, S. P. & Hedstrom, L. (2008). Retinal isoforms of inosine 5’-monophosphate dehydrogenase type 1 are poor nucleic acid binding proteins. Archives of Biochemistry and Biophysics, 472(2), 100–104.
Aherne, A., Kennan, A., Kenna, P. A., McNally, N., Lloyd, D. G., Alberts, I. L., Kiang, A. S., Humphries, M. M., Ayuso, C. & Engel, P. C. et al. (2004). On the molecular pathology of neurodegeneration in IMPDH1-based retinitis pigmentosa. Human Molecular Genetics, 13(6), 641–650.
Acknowledgements
This work was supported by the Research Council of University of Tehran, and Iranian National Science Foundation (INSF) under the project #94020269.
Author contributions
B.A. performed the experimental analyses and assisted in writing the paper, R.Y. supervised the research, designed the experiments and editted the manuscript, M.M. performed a part of experimental analyses, E B. done the bioinformatics model analysis and participated in the manuscript preparation and H.G. analyzed the molecular genetics data and sequences.
Funding
Research Council of University of Tehran.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare no competing interests.
Ethical Approval
All applicable international, national, and/or institutional guidelines for the care and use of animals were followed.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Andashti, B., Yazdanparast, R., Motahar, M. et al. Terminal Peptide Extensions Augment the Retinal IMPDH1 Catalytic Activity and Attenuate the ATP-induced Fibrillation Events. Cell Biochem Biophys 79, 221–229 (2021). https://doi.org/10.1007/s12013-021-00973-2
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12013-021-00973-2