Abstract
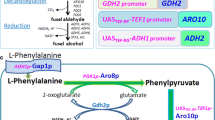
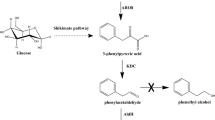
2-Phenylethanol (2-PE) is an important aromatic alcohol with a rose-like odor and has wide applications. The present work aims to construct a synthetic pathway for 2-PE synthesis from glucose in Escherichia coli. First, the genes adh1 (encoding alcohol dehydrogenase) and kdc (encoding phenylpyruvate decarboxylase) from Saccharomyces cerevisiae S288c and Pichia pastoris GS115 were investigated in E. coli, respectively, and single overexpression of adh1 or kdc significantly increased 2-PE accumulation. When co-overexpressing adh1 and kdc, 2-PE was increased up to 130 from 57 mg/L. Furthermore, by optimizing coordinated expression of the four committed genes aroF, pheA, adh1 and kdc, 2-PE was improved to 285 mg/L which was the highest production of 2-PE by the recombinant E. coli system. In addition, our results also demonstrated that the tyrB gene, which encodes aromatic-amino-acid transaminase, plays an important role on 2-PE synthesis.





Similar content being viewed by others
References
Etschmann, M. M., Bluemke, W., Sell, D., & Schrader, J. (2002). Biotechnological production of 2-phenylethanol. Applied Microbiology and Biotechnology, 59, 1–8.
Atsumi, S., Hanai, T., & Liao, J. C. (2008). Non-fermentative pathways for synthesis of branched-chain higher alcohols as biofuels. Nature, 451, 86–89.
Xu, P., Hua, D., & Ma, C. (2007). Microbial transformation of propenylbenzenes for natural flavour production. Trends in Biotechnology, 25, 571–576.
Hua, D., & Xu, P. (2011). Recent advances in biotechnological production of 2-phenylethanol. Biotechnology Advances, 29, 654–660.
Jin, J. Z., Li, H., & Zhang, J. (2010). Improved synthesis of (S)-1-phenyl-2-propanol in high concentration with coupled whole cells of Rhodococcus erythropolis and Bacillus subtilis on preparative scale. Applied Biochemistry and Biotechnology, 162, 2075–2086.
Jollivet, N., Bézenger, M. C., Vayssier, Y., & Belin, J. M. (1992). Production of volatile compounds in liquid cultures by six strains of coryneform bacteria. Applied Microbiology and Biotechnology, 36, 790–794.
Lomascolo, A., Lesage-Meessen, L., Haon, M., Navarro, D., Antona, C., Faulds, C., & Marcel, A. (2001). Evaluation of the potential of Aspergillus niger species for the bioconversion of L-phenylalanine into 2-phenylethanol. World Journal of Microbiology and Biotechnology, 17, 99–102.
Stark, D., Zala, D., Münch, T., Sonnleitner, B., Marison, I., & Von Stockar, U. (2003). Inhibition aspects of the bioconversion of l-phenylalanine to 2-phenylethanol by Saccharomyces cerevisiae. Enzyme and Microbial Technology, 32, 212–223.
Eshkol, N., Sendovski, M., Bahalul, M., Katz-Ezov, T., Kashi, Y., & Fishman, A. (2009). Production of 2-phenylethanol from L-phenylalanine by a stress tolerant Saccharomyces cerevisiae strain. Journal of Applied Microbiology, 106, 534–542.
Seo, W. T., & Ahn, K. C. (2003). Production of phenylethanol from L-phenylalanine by Candida sp S-8. Food Science and Biotechnology, 12, 644–648.
Serp, D., von Stockar, U., & Marison, I. W. (2003). Enhancement of 2-phenylethanol productivity by Saccharomyces cerevisiae in two-phase fed-batch fermentations using solvent immobilization. Biotechnology and Bioengineering, 82, 103–110.
Wang, H., Dong, Q. F., Meng, C., Shi, X. A., & Guo, Y. H. (2011). A continuous and adsorptive bioprocess for efficient production of the natural aroma chemical 2-phenylethanol with yeast. Enzyme and Microbial Technology, 48, 404–407.
Stark, D., Munch, T., Sonnleitner, B., Marison, I. W., & von Stockar, U. (2002). Extractive bioconversion of 2-phenylethanol from L-phenylalanine by Saccharomyces cerevisiae. Biotechnology Progress, 18, 514–523.
Celinska, E., Kubiak, P., Bialas, W., Dziadas, M., & Grajek, W. (2013). Yarrowia lipolytica: the novel and promising 2-phenylethanol producer. Journal of Industrial Microbiology & Biotechnology, 40, 389–392.
Chen, X., Zhou, L., Kangming, T., Kumar, A., Singh, S., Prior, B. A., Wang, Z. (2013). Metabolic engineering of Escherichia coli: a sustainable industrial platform for bio-based chemical production. Biotechnology Advances.
Keasling, J. D. (2008). Synthetic biology for synthetic chemistry. ACS Chemical Biology, 3, 64–76.
Goss, R. J., Shankar, S., & Fayad, A. A. (2012). The generation of “unnatural” products: synthetic biology meets synthetic chemistry. Natural Product Reports, 29, 870–889.
Atsumi, S., Wu, T. Y., Machado, I. M., Huang, W. C., Chen, P. Y., Pellegrini, M., & Liao, J. C. (2010). Evolution, genomic analysis, and reconstruction of isobutanol tolerance in Escherichia coli. Molecular Systems Biology, 6, 449.
Smith, K. M., & Liao, J. C. (2011). An evolutionary strategy for isobutanol production strain development in Escherichia coli. Metabolic Engineering, 13, 674–681.
Woodruff, L. B., Boyle, N. R., & Gill, R. T. (2013). Engineering improved ethanol production in Escherichia coli with a genome-wide approach. Metabolic Engineering, 17, 1–11.
Koma, D., Yamanaka, H., Moriyoshi, K., Ohmoto, T., & Sakai, K. (2012). Production of aromatic compounds by metabolically engineered Escherichia coli with an expanded shikimate pathway. Applied and Environmental Microbiology, 78, 6203–6216.
Gosset, G. (2009). Production of aromatic compounds in bacteria. Current Opinion in Biotechnology, 20, 651–658.
Brynildsen, M. P., & Liao, J. C. (2009). An integrated network approach identifies the isobutanol response network of Escherichia coli. Molecular Systems Biology, 5, 277.
Zheng, Y. N., Li, L. Z., Xian, M., Ma, Y. J., Yang, J. M., Xu, X., & He, D. Z. (2009). Problems with the microbial production of butanol. Journal of Industrial Microbiology & Biotechnology, 36, 1127–1138.
Woodruff, L. B., Pandhal, J., Ow, S. Y., Karimpour-Fard, A., Weiss, S. J., Wright, P. C., & Gill, R. T. (2013). Genome-scale identification and characterization of ethanol tolerance genes in Escherichia coli. Metabolic Engineering, 15, 124–133.
Zhang, H. F., Chong, H. Q., Ching, C. B., Song, H., & Jiang. R. R. (2012). Engineering global transcription factor cyclic AMP receptor protein of Escherichia coli for improved 1-butanol tolerance. Applied Microbiology and Biotechnology, 94, 1107–1117.
Liu, H., Liu, K., Yan, M., Xu, L., & Ouyang, P. (2011). gTME for improved adaptation of Saccharomyces cerevisiae to corn cob acid hydrolysate. Applied Biochemistry and Biotechnology, 164, 1150–1159.
Dunlop, M. J., Dossani, Z. Y., Szmidt, H. L., Chu, H. C., Lee, T. S., Keasling, J. D., Hadi, M. Z., & Mukhopadhyay, A. (2011). Engineering microbial biofuel tolerance and export using efflux pumps. Molecular Systems Biology, 7, 487.
Reyes, L. H., Almario, M. P., & Kao, K. C. (2011). Genomic library screens for genes involved in n-butanol tolerance in Escherichia coli. Plos One, 6.
Minty, J. J., Lesnefsky, A. A., Lin, F., Chen, Y., Zaroff, T. A., Veloso, A. B., Xie, B., McConnell, C. A., Ward, R. J., Schwartz, D. R., Rouillard, J. M., Gao, Y., Gulari, E., & Lin, X. N. (2011). Evolution combined with genomic study elucidates genetic bases of isobutanol tolerance in Escherichia coli. Microbial Cell Factories, 10, 18.
Ting, C. N., Wu, J., Takahashi, K., Endo, A., & Zhao, H. (2012). Screened butanol-tolerant Enterococcus faecium capable of butanol production. Applied Biochemistry and Biotechnology, 168, 1672–1680.
Hirata, H., Ohnishi, T., Ishida, H., Tomida, K., Sakai, M., Hara, M., & Watanabe, N. (2012). Functional characterization of aromatic amino acid aminotransferase involved in 2-phenylethanol biosynthesis in isolated rose petal protoplasts. Journal of Plant Physiology, 169, 444–451.
Jarboe, L. R. (2011). YqhD: a broad-substrate range aldehyde reductase with various applications in production of biorenewable fuels and chemicals. Applied Microbiology and Biotechnology, 89, 249–257.
Russell, D. W., Smith, M., Williamson, V. M., & Young, E. T. (1983). Nucleotide sequence of the yeast alcohol dehydrogenase II gene. The Journal of Biological Chemistry, 258, 2674–2682.
Juminaga, D., Baidoo, E. E., Redding-Johanson, A. M., Batth, T. S., Burd, H., Mukhopadhyay, A., Petzold, C. J., & Keasling, J. D. (2012). Modular engineering of L-tyrosine production in Escherichia coli. Applied and Environmental Microbiology, 78, 89–98.
Kang, Z., Wang, Y., Gu, P., Wang, Q., & Qi, Q. (2011). Engineering Escherichia coli for efficient production of 5-aminolevulinic acid from glucose. Metabolic Engineering, 13, 492–498.
Wu, J., Du, G., Zhou, J., & Chen, J. (2013). Metabolic engineering of Escherichia coli for (2S)-pinocembrin production from glucose by a modular metabolic strategy. Metabolic Engineering, 16, 48–55.
Lerner, C. G., & Inouye, M. (1990). Low copy number plasmids for regulated low-level expression of cloned genes in Escherichia coli with blue/white insert screening capability. Nucleic Acids Research, 18, 4631.
Lim, H. N., Lee, Y., & Hussein, R. (2011). Fundamental relationship between operon organization and gene expression. Proceedings of the National Academy of Sciences of the United States of America, 108, 10626–10631.
Kovacs, K., Hurst, L. D., & Papp, B. (2009). Stochasticity in Protein Levels Drives Colinearity of Gene Order in Metabolic Operons of Escherichia coli. Plos Biology, 7.
Zhou, H., Liao, X., Wang, T., Du, G., & Chen, J. (2010). Enhanced l-phenylalanine biosynthesis by co-expression of pheA(fbr) and aroF(wt). Bioresource Technology, 101, 4151–4156.
Acknowledgments
This work was financially supported by the Major State Basic Research Development Program of China (973 Program, 2014CB745103, 2013CB733602), the National Natural Science Foundation of China (31200020), the National High Technology Research and Development Program of China (2012AA021201), the National Science Foundation for Post-doctoral Scientists of China (2013 M540414), the Jiangsu Planned Projects for Postdoctoral Research Funds (1301010B), the 111 Project and the Priority Academic Program Development of Jiangsu Higher Education Institutions.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kang, Z., Zhang, C., Du, G. et al. Metabolic Engineering of Escherichia coli for Production of 2-phenylethanol from Renewable Glucose. Appl Biochem Biotechnol 172, 2012–2021 (2014). https://doi.org/10.1007/s12010-013-0659-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-013-0659-3