Abstract
Purpose of Review
Insulin is at the heart of diabetes mellitus (DM). DM alters cardiac metabolism causing cardiomyopathy, ultimately leading to heart failure. Polyamines, organic compounds synthesized by cardiomyocytes, have an insulin-like activity and effect on glucose metabolism, making them metabolites of interest in the DM heart. This review sheds light on the disrupted microRNA network in the DM heart in relation to developing novel therapeutics targeting polyamine biosynthesis to prevent/mitigate diabetic cardiomyopathy.
Recent Findings
Polyamines prevent DM-induced upregulation of glucose and ketone body levels similar to insulin. Polyamines also enhance mitochondrial respiration and thereby regulate all major metabolic pathways. Non-coding microRNAs regulate a majority of the biological pathways in our body by modulating gene expression via mRNA degradation or translational repression. However, the role of miRNA in polyamine biosynthesis in the DM heart remains unclear.
Summary
This review discusses the regulation of polyamine synthesis and metabolism, and its impact on cardiac metabolism and circulating levels of glucose, insulin, and ketone bodies. We provide insights on potential roles of polyamines in diabetic cardiomyopathy and putative miRNAs that could regulate polyamine biosynthesis in the DM heart. Future studies will unravel the regulatory roles these miRNAs play in polyamine biosynthesis and will open new doors in the prevention/treatment of adverse cardiac remodeling in diabetic cardiomyopathy.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Impaired glucose metabolism due to insulin deficiency (T1DM) or insulin resistance (T2DM) causes cardiac metabolic remodeling in diabetes mellitus (DM) [1]. The DM heart develops a cardiac muscle disorder called diabetic cardiomyopathy (DMCM), which leads to heart failure [2•]. DMCM causes steady progressive remodeling in the heart, exhibiting different features at different stages: early stage — increased oxidative stress and inflammation; middle stage — increased cell death, fibrosis, and diastolic dysfunction; and late stage — diastolic and systolic dysfunction [3]. At the crossroad of these pathologies is metabolic dysregulation, a significant part of which is consists of impaired polyamine biosynthesis.
Polyamines are charged organic compounds with insulin-like actions that affect the metabolism of both lipids and glucose [4•, 5, 6]. The rate-limiting enzyme of polyamine biosynthesis is ornithine decarboxylase (ODC). In streptozotocin-induced T1DM rats, levels of ODC are markedly lower in the heart, liver, and skeletal muscles but retain normal levels in insulin-treated T1DM rats [7]. Thus, polyamine biosynthesis may be regulated by insulin. High glucose decreases levels of the polyamine spermine in primary neonatal rat cardiomyocytes, while simultaneously impairing mitochondrial function by reducing mitochondrial membrane potential and increasing ATP leakage. Supplementation of spermine, another polyamine, in these cardiomyocytes prevented high glucose-induced loss of mitochondrial membrane potential and improved ATP levels [8]. Therefore, polyamines, such as spermine, could be important in preserving mitochondrial energetics in a high glucose environment. Moreover, supplementation of spermine improves glucose uptake, reduces myocardial cell death and oxidative stress, and attenuates cardiac dysfunction in the streptozotocin-induced T1DM heart [9]. In alloxan-induced DM rats, separate treatments with insulin and polyamines were compared in their ability to maintain serum levels of glucose and beta-hydroxybutyrate — a ketone body associated with mitochondrial damage and cardiotoxicity. Polyamines reduced the levels of glucose and beta-hydroxybutyrate comparable to that of insulin, suggesting that polyamines and insulin signaling work in parallel for controlling the circulating levels of glucose and ketone bodies in DM [10••]. These findings suggest that polyamines have critical protective roles in myocardial metabolism and adverse cardiac remodeling in DM. However, it remains unclear as to how the biosynthesis of polyamines is regulated in the DM heart.
The majority of biological functions are regulated by microRNAs-tiny (18–25 nucleotides long), endogenous, non-coding RNAs that fine tune gene expression by mRNA degradation or translational repression [11•, 12]. A single microRNA (miRNA) can regulate several genes in a biological pathway and thus could have therapeutic potential for cardiovascular diseases [13•]. Levels of many miRNAs that regulate metabolism are altered in the DM heart [14,15,16]. How miRNA regulates cardiac polyamine biosynthesis in the DM heart is unknown.
Here, we searched for potential miRNAs that are present in the heart and have predicted targets in the polyamine biosynthesis pathway. We also elaborated the potential roles of polyamines in cardiac metabolism and maintaining ketone body levels — a crucial step in metabolism to control ketoacidosis in DM. This review sets a platform to explore the regulatory roles of polyamines in mitochondrial dysfunction and cardiac metabolic derangement during DMCM and investigate the potential of miRNAs to regulate levels of polyamines in the DM heart.
Regulation of Polyamine Levels
Polyamines are organic compounds with two or more amino groups that render them a high positive charge, thus also gaining the term polycations [17•]. They were first discovered in seminal fluids [18•]. One important function of polyamines is the translational regulation of genes [19, 20]. The uptake, biosynthesis, and catabolism of polyamines are briefly described below:
Polyamine Uptake
Polyamines from the diet are transported through specific uptake mechanisms; however, there is a lack of unifying theory behind this [21]. Some transporters that have been suggested so far include the solute carrier SLC3A2, with selectivity towards putrescine polyamine [22, 23], and the vesicular solute carrier SLC18B, with selectivity towards spermidine and spermine polyamines [24]. An alternate transport pathway utilizes endocytic uptake of polyamines through glypicans, which are heparan sulfate groups within plasma membrane proteins [25]. The lysosomal/endosomal transporter ATP13A3 has also been suggested for the movement of putrescine, spermine, and spermidine into the cytosol [26•].
Polyamine Biosynthesis
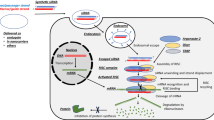
While the primary exogenous source of mammalian polyamines is through nutrition, most of the intracellular polyamine pool is regulated through cellular synthesis or synthesis by the gut microbiome [27, 28]. Gut microbiome-triggered polyamine production is predominantly responsible for colorectal cancer [29]. Polyamines are synthesized from amino acids — arginine, ornithine, and methionine — and rely on unique enzymes to harness each amino acid pool separately [30, 31]. Arginase converts arginine to ornithine and exists in two isoforms-arginase 1 (cytoplasmic) and arginase 2 (mitochondrial) [32]. Although predominantly found in the liver, arginase 1 is also present in endothelium, smooth muscle, and neuronal cells [33,34,35,36]. Ornithine is metabolized by ODC to produce the first polyamine, putrescine. ODC is the rate-limiting enzyme of the polyamine biosynthesis pathway, and as such is tightly regulated at the transcriptional and post-transcriptional levels [37, 38]. Spermidine synthase biosynthesizes the second polyamine, spermidine, from putrescine. Spermidine is converted into the final polyamine, spermine, via the enzyme spermine synthase (Fig. 1).

source of ornithine is from the urea cycle in the liver. S-adenosylmethionine metabolism is a parallel pathway that contributes to synthesis of polyamines. This is a part of the methionine cycle which occurs in the cytoplasm of all cells, including cardiomyocytes. Figure created using BioRender
The intracellular polyamine metabolic pathway. The primary polyamine metabolic pathway occurs in the cytoplasm of all cells and involves the formation of polyamines (putrescine, spermidine, and spermine) from ornithine through ornithine decarboxylase enzyme. The intracellular
Another pathway for polyamine biosynthesis is the methionine cycle, which is present in most cells [39]. Methionine is shuttled as S-adenosyl methionine via S-adenosylmethionine synthase (SAMS). This is further metabolized through S-adenosylmethionine decarboxylase to decarboxylated S-adenosylmethionine, which can replenish polyamines pool through spermidine synthase (for spermidine) and spermine synthase (for spermine) (Fig. 1). In several pathological conditions, such as hepatic carcinoma, lung cancer, and prostate cancer, the levels of S-adenosylmethionine are elevated [40, 41, 42•].
Polyamine Catabolism
While non-reversible, it is possible to recycle spermine and spermidine through the action of N-acetyltransferase intermediates [43, 44]. Spermine can be converted to spermidine through the action of spermine oxidase (SMOX). Alternatively, spermine or spermidine can be converted to lower polyamines through a two-step reaction involving spermidine/spermine N1-acetyltransferase (SAT1) and acetylpolyamine oxidase. This is an important step in polyamine catabolism as SAT1 is the primary driving force for maintaining intracellular polyamine pools. Upregulation of SAT1 drives spermine catabolism, which activates the biosynthesis of spermidine. Spermine is also catabolized by spermine oxidase to form spermidine, thus maintaining polyamine concentrations [42•] (Fig. 1). Oxidative deamination of polyamines by Cu2+-dependent diamine oxidase plays an important role in the terminal catabolism of polyamines, which has been discussed at length in a previous review [43].
Protective Roles of Polyamines in Diabetes Mellitus
Polyamine metabolism affects DM by improving insulin action and preserving pancreas function. Polyamine biosynthesis is stimulated by glucose uptake in islet cells [44, 45]. Increases in the level of insulin due to elevated glucose levels is paralleled by an increase in polyamine synthesis that inhibits pro-insulin transcription [46]. It is known that polyamines, specifically spermidine and spermine, regulate insulin signaling via interaction with insulin-like-growth factor-1 and by increasing the transcription of the proinsulin gene [46, 47]. SAT1 knockout mice exhibit insulin resistance upon aging, while SAT1 overexpressing mice demonstrate improved glucose tolerance [48]. Increasing catabolism by depletion of internal polyamine pools also protects against the development of hyperglycemia by preserving pancreatic beta cell survival in chemically-induced DM in preclinical models [49]. In addition to this, deoxyhypusine synthase utilizes spermine to induce adaptive pancreatic beta cell proliferation under insulin resistant conditions [50]. Furthermore, polyamine oxidase is increased in the serum of diabetic children, providing a potential role of polyamine oxidase in increased levels of circulatory oxidative stress [51•]. The serum levels of putrescine are elevated in patients with T2DM, corroborating that polyamine metabolism is dysregulated in DM [52••].
Evidence of Polyamines Involvement in Cardiomyopathy
Polyamines are involved in the cardiac development of rats and mice [53•, 54••]. Polyamine dysregulation is linked to cardiovascular diseases caused by ischemia/reperfusion (IR) injuries, suggesting a potential of polyamines in adverse cardiac remodeling [54••]. However, a direct link between polyamines and cardiomyopathy has not been established. A distinct hallmark in ischemic cardiomyopathy is increased expression of N8-acetylspermidine (N8AS) [55•]. N8AS is formed from spermidine by the enzyme SAT1 and converted into spermidine by histone deacetylase [55•]. Overexpression of polyamine metabolic enzymes such as ODC and SAT1 has been linked to ischemic cardiomyopathy, and polyamines like N8AS serve as a potential biomarker for ischemic cardiomyopathy. Other polyamines such as spermidine and spermine are downregulated in cardiomyopathy, possibly due to consumption of spermine in preventing myocardial apoptosis [56•, 57]. In cirrhotic cardiomyopathy, an increase in spermidine offers a protective effect by reducing inflammation and increasing antioxidant enzymes [58]. Other studies on rats have shown that spermidine provides protection against heart injury by preventing oxidative stress [59•]. These findings support the potential cardioprotective roles of polyamines in cardiovascular diseases and cardiomyopathy.
Polyamines and Cardiac Metabolism
The pathway of polyamine biosynthesis is tied upstream to the TCA cycle, linking it to all major metabolic pathways. In patients with ischemic cardiomyopathy, plasma levels of N8AS correlate with an increase in metabolites in the carnitine shuttle and an oversaturation of the fatty acid β‐oxidation pathway [55•]. RNA sequencing performed in rat hearts revealed that spermine and spermidine treatment prevented cardiac age-related deterioration by restoring pathways associated with fatty acid metabolism [60]. In neonatal rat cardiomyocytes, spermidine treatment restored hypoxia-impaired state 3 and state 4 mitochondrial respiration [59•]. Spermidine feeding also enhanced mitochondrial respiration in aged mice and hypertensive rats, leading to improved cardiomyocyte composition and cardiac function [61••]. Heart failure, which typically decreases fatty acid metabolism, is associated with a significant increase in levels of putrescine, spermine, ornithine, and arginine along with a decrease in levels of several lipid intermediates in mouse hearts [62]. Mass spectrophotometric analysis conducted in 12/15LOX−/− mice undergoing ischemic heart failure indicated metabolic reprograming via an upregulation in levels of the amino acids arginine, ornithine, methionine, and the polyamines spermidine and spermine, suggesting a preference for amino acid entry directly into polyamine biosynthesis over the Krebs cycle in failing hearts with the impaired presence of fatty acid-metabolizing enzymes [63]. Ketogenesis, upregulated during fatty acid metabolism over-saturation, was found to be transiently activated, while arginine, methionine, and S-adenosylmethionine were found to be consistently decreased across postnatal mouse heart development [64]. Interestingly, treatment with L-arginine, spermidine, spermine, and putrescine in pregnant diabetic mice significantly lowered circulating level of beta-hydroxybutyrate (a ketone body), suggesting a ketogenic regulatory role of polyamines [10••].
Potential Roles of Polyamines in Diabetic Cardiomyopathy
DM induces DMCM, leading to an increased risk of heart failure. T2DM increases the risk of heart failure in men 2.4-fold and in women 5.1-fold [65], while T1DM increases the risk of heart failure fourfold independent of sex [66]. While there is limited clinical data examining polyamines in DMCM, there are several preclinical studies focusing on determining the role of polyamine biosynthesis in the DM heart. Exogenous spermine attenuates DMCM by suppressing ROS-p53-mediated downregulation of calcium-sensitive receptors and inhibiting Wnt/β-catenin signaling [9, 67•]. Downregulated calcium-sensitive receptor expression has been tied to DMCM, supporting the concept that downregulated spermine could contribute to DMCM [68]. Furthermore, ODC activity is 77% lower in the hearts of T1DM rats and is rescued by treatment via insulin [7, 69•]. Cardiac insulin binding is improved by treatment with spermine and spermidine via increased cardiac insulin receptor tyrosine kinase activity, corroborating a role of impaired polyamine biosynthesis in T2DM-induced insulin resistance [70]. Additional studies have shown that high glucose was able to induce fibrosis in rat cardiac fibroblasts through increased arginase activity. Upregulation of arginase activity is attributed to increased levels of intracellular ROS in hyperglycemic H9C2 cardiac myoblasts, while inhibition of arginase via remote ischemic preconditioning significantly preserved myocardial infarct size in T1DM, suggesting a potential maladaptive mechanism to the over activation of upstream enzymes in the polyamine biosynthesis pathway in DMCM [71, 72]. Furthermore, spermidine consumption has been shown to worsen myocardial lipid deposition, a precursor of DMCM, in high fat-diet-induced obese male mice [73].
A summary of expression levels of various polyamines and polyamine metabolic enzymes that have potential to contribute to DMCM is presented in Table 1. One of the challenges with linking polyamines to DMCM is the change in expression levels of polyamines over the course of disease progression. An increase or decrease in polyamine levels may either increase or decrease injury in a dose-dependent fashion in ischemia–reperfusion injury [74•]. There is a lack of human studies determining the role of polyamines in cardiac dysfunction and the regulation of polyamine biosynthesis and metabolism.
Potential miRNAs Involved in Regulation of Cardiac Polyamines in Diabetic Cardiomyopathy
DMCM is characterized by a dysregulation of the miRNA regulatory network. Out of the total cataloged 2000 mature miRNAs, approximately 400 are expressed in the human heart [75]. Of these 400 miRNAs, the expression of approximately 2 dozen is changed in the human heart (Table 2). Of these miRNAs, miR-1, miR-34a, miR-34b, miR-34c, miR-199a, miR-199b, miR-204, miR-208a, miR-210, miR-216a, miR-223, miR-320, miR-372, and miR-485 are upregulated while miR-9, miR-23b, miR-29b, miR-126, miR-132, miR-133a, miR-181a, miR-182, miR-214, and miR-221 are downregulated. Not all miRNA expression follows the same change in trend between T1DM and T2DM hearts. miR-21, miR-34a, miR-141, and miR-499 have opposite changes in rodent cardiac tissue expression between T1DM and T2DM hearts. Furthermore, in vitro studies have allowed a better understanding of the effects of dysregulated miRNAs on DM-induced cardiac remodeling pathways. Many cardiac miRNAs are involved in regulating cell death, with miR-1, miR-19, miR-34a, miR-195, miR-320, and miR-483 acting as pro-apoptotic whereas miR-30d as pro-pyroptotic. miR-26a, miR-26b, miR-30c, miR-144, and miR-181a are anti-apoptotic, while miR-9 is anti-pyroptotic. miR-195 induces oxidative stress, while miR-21, miR-144, and miR-373 each prevent the generation of oxidative stress. miR-133a, miR-150, miR-181a, and miR-373 are anti-hypertrophic, while miR-195 and miR-451 are pro-hypertrophic. Lastly, miR-29 and miR-141 promote DM-induced cardiac fibrosis whereas both miR-30c and miR-200b work to restore metabolism in the diabetic myocardium (Table 3).
Predicted miRNA and Their Targets in Polyamine Biosynthesis Pathway in Diabetic Cardiomyopathy
While there are no studies showing a direct relationship between miRNAs and polyamine biosynthesis in DMCM, there is some data supporting roles of those miRNAs that are deregulated during DMCM in the regulation of polyamine biosynthesis in other diseases. miR-34a and miR-34b, which are upregulated in DMCM, target methionine adenosyl transferase 2A (MAT2A). S-adenosylmethionine (SAM) and methylthioadenosine (MTA) upregulate miR-34a and miR-34b in colorectal cancer cells [76]. Cellular levels of polyamines in intestinal epithelial cells decrease miR-29b via decreasing JunD-expression levels, mimicking miR-29b’s expression in the DM heart [77]. S-adenosyl-L-methionine (AdoMet) upregulates miR-34c in triple negative breast cancer cell lines while inducing a pro-apoptotic cell death effect. miR-34c is also increased in the DM heart, while miR-34a has been established to be both upregulated in the DM heart and be pro-apoptotic [78, 79]. Decreased levels of miR-199a-5p are also correlated with increased aggressive pituitary tumor growth via targeted inhibition of SAT1. miR-199a is also increased in the DM heart [80]. miR-210, which is both pro-cancerous and overexpressed in DMCM, significantly increased levels of all metabolites in HEK293 cells, including putrescine and spermidine [80, 81]. miR-372, another oncogenic miRNA upregulated during DMCM, was the first established miRNA to be inhibited via polyamine derivatives [80, 82]. Furthermore, knockdown of miR-485-3p in Hep3B and HepG2 liver cell lines induced methionine adenosyl-transferase 1A (MAT1A) expression [83]. miR-200a targets Keap1 to allow NRF2 binding to SAT1 in non-small-cell lung carcinoma cell lines. While miR-200a expression is not changed in the DM heart, miR-200b and miR-200c expressions are different [84, 85]. This remains relevant, as the miR-200a and miR-200b/c seed sequence varies by only a single nucleotide [86]. In addition to these potential miRNA regulators of polyamine metabolism, there are extensive mRNA gene-interaction analysis conducted utilizing microarray and immunoprecipitation showing interactions between miRNAs disrupted during DMCM and polyamine biosynthesis enzymes (Table 4).
Utilizing RNA binding prediction software, it is possible to examine the feasibility of cardiac miRNA targets within the polyamine biosynthesis pathway in silico, such as the predicted binding between miR-1 and ODC (Fig. 2). miR-1 is overexpressed and ODC activity is drastically decreased in the DM heart [69•, 87]. These findings along with in silico prediction results suggest a potential role of the dysregulated miRNA-polyamine axis in DMCM (Fig. 3).
RNA-RNA binding prediction and interaction of hsa-miR-1-3p and ODC. RNA-RNA binding prediction was performed using the IntaRNA online tool (http://rna.informatik.uni-freiburg.de) to measure the hybridization energy between ODC1 (NM_001287189.2) and the hsa-miR-1-3P (MI0000651). The binding prediction of the microRNA and the target mRNA sequence revealed a hybridization energy of − 15.86 kcal/mol in exon 2 position 77 to position 94 with two mismatching nucleotides at position 87 and 88 (A, B). A secondary structure analysis was performed using Forna (Kerpedjiev P, Hammer S, Hofacker IL, 2015). Forna (force-directed RNA): Simple and effective online RNA secondary structure diagrams. Bioinformatics 31(20):3377–9.) server in Vienna’s RNA lab tool to reveal the binding position of the microRNA in the target sequence. The top structure of C represents color-coded mRNA secondary structure of ODC1 Exon 2. The nucleotide colors represent the type of structure they form, green: stems (canonical helices), red: multiloops (junctions), yellow: interior loops, blue: hairpin loops, orange: 5′ and 3′ unpaired region. The bottom structure represents the binding site of the microRNA, the green color represents mismatching Nucleotide, while the red color represents matching nucleotides at the binding site
Gene interactions of hsa-miR-1-3p within the polyamine biosynthesis pathway. This figure shows hsa-miR-1-3p’s known interactions in the polyamine biosynthesis pathway from microarray and co-immunoprecipitation experiments collected from the Tarbase v8 database. This data analysis resulted in hsa-miR-1-3p 3′ UTR binding association with Ornithine Decarboxylase (ODC), Spermine synthase (SPMS), and Spermidine/spermine N1-acetyltransferase (SAT1). Other major polyamine biosynthesis enzymes and metabolites depicted are spermidine synthase (SPDS), acetylated polyamine oxidase (AcPAO), S-adenosylmethionine synthase (SAMS), S-adenosylmethionine decarboxylase (AdoMetDC), and Spermine Oxidase (SMOX)
Summary and Future Direction
In this review, we summarized potential roles of polyamines in the progression of DMCM and the regulatory roles of miRNA in maintaining the synthesis and metabolism of polyamines. We have also provided a list of putative candidate miRNAs that are involved in the regulation of cardiac polyamines in DMCM. As DM is a metabolic disease, we elaborated the specific roles of polyamines in cardiac metabolism. In the DM heart, fat metabolism is increased, and mitochondrial function is impaired. Thus, we have elaborated how polyamines are involved in the regulation of fatty acid beta-oxidation and mitochondrial respiration. We have also elaborated how polyamines are involved in ketogenesis.
Despite intensive glycemic control, the risk of heart failure is high in DM patients [88•]. Thus, there is no cure for DM-induced heart failure, and a novel therapeutic approach is warranted. Regulation of cardiac polyamines could prove to be a potential novel avenue to ameliorate DMCM and prevent DM-induced heart failure. MiRNAs and their predicted targets described in this article warrant empirical studies to determine their specific roles in polyamine biosynthesis and cardiac metabolism in DMCM.
References
Papers of particular interest, published recently, have been highlighted as: • Of importance •• Of major importance
Bugger H, Abel ED. Molecular mechanisms of diabetic cardiomyopathy. Diabetologia. 2014;57(4):660–71. https://doi.org/10.1007/s00125-014-3171-6.
• Rubler S, Dlugash J, Yuceoglu YZ, Kumral T, Branwood AW, Grishman A (1972) New type of cardiomyopathy associated with diabetic glomerulosclerosis. Am J Cardiol 30 (6):595–602. This is the first paper that discovered the effect of diabetes mellitus on heart dysfunction and lead to development of new area — diabetic cardiomyopathy.
Chavali V, Tyagi SC, Mishra PK. Predictors and prevention of diabetic cardiomyopathy. Diabetes Metab Syndr Obes. 2013;6:151–60. https://doi.org/10.2147/DMSO.S30968.
• Lockwood DH, East LE (1974) Studies of the insulin-like actions of polyamines on lipid and glucose metabolism in adipose tissue cells. J Biol Chem 249 (24):7717–7722. This paper demonstrates dose-response relationship for effect of polyamines on glucose oxidation and insulin-like actions of polyamines.
Sadasivan SK, Vasamsetti B, Singh J, Marikunte VV, Oommen AM, Jagannath MR, Pralhada Rao R. Exogenous administration of spermine improves glucose utilization and decreases bodyweight in mice. Eur J Pharmacol. 2014;729:94–9. https://doi.org/10.1016/j.ejphar.2014.01.073.
Pirinen E, Kuulasmaa T, Pietila M, Heikkinen S, Tusa M, Itkonen P, Boman S, Skommer J, Virkamaki A, Hohtola E, Kettunen M, Fatrai S, Kansanen E, Koota S, Niiranen K, Parkkinen J, Levonen AL, Yla-Herttuala S, Hiltunen JK, Alhonen L, Smith U, Janne J, Laakso M. Enhanced polyamine catabolism alters homeostatic control of white adipose tissue mass, energy expenditure, and glucose metabolism. Mol Cell Biol. 2007;27(13):4953–67. https://doi.org/10.1128/MCB.02034-06.
Conover CA, Rozovski SJ, Belur ER, Aoki TT, Ruderman NB. Ornithine decarboxylase activity in insulin-deficient states. Biochem J. 1980;192(2):725–32. https://doi.org/10.1042/bj1920725.
Wang Y, Wang Y, Li F, Zhang X, Li H, Yang G, Xu C, Wei C. Spermine protects cardiomyocytes from high glucose-induced energy disturbance by targeting the CaSR-gp78-ubiquitin proteasome system. Cardiovasc Drugs Ther. 2021;35(1):73–85. https://doi.org/10.1007/s10557-020-07064-z.
Wang Y, Chen J, Li S, Zhang X, Guo Z, Hu J, Shao X, Song N, Zhao Y, Li H, Yang G, Xu C, Wei C. Exogenous spermine attenuates rat diabetic cardiomyopathy via suppressing ROS-p53 mediated downregulation of calcium-sensitive receptor. Redox Biol. 2020;32:101514. https://doi.org/10.1016/j.redox.2020.101514.
•• Mendez JD, Palomar-Morales M (1999) Prevention by L-arginine and polyamines of delayed development and embryotoxicity caused by chemically-induced diabetes in rats. Reprod Toxicol 13 (6):501-509. https://doi.org/10.1016/s0890-6238(99)00039-8. This paper provides empirical evidence for insulin-like effects of polyamines on restoring the levels of glucose and ketone body in diabetic rats.
• Bartel DP (2004) MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116 (2):281–297. This is a classical paper on miRNA biology.
Bartel DP. MicroRNAs: target recognition and regulatory functions. Cell. 2009;136(2):215–33. https://doi.org/10.1016/j.cell.2009.01.002.
• Mishra PK, Tyagi N, Kumar M, Tyagi SC (2009) MicroRNAs as a therapeutic target for cardiovascular diseases. J Cell Mol Med 13(4):778-789. https://doi.org/10.1111/j.1582-4934.2009.00744.xThis review focuses on multifaceted roles of miRNA in cardiovascular diseases.
Chavali V, Tyagi SC, Mishra PK. Differential expression of dicer, miRNAs, and inflammatory markers in diabetic Ins2+/- Akita hearts. Cell Biochem Biophys. 2014;68(1):25–35. https://doi.org/10.1007/s12013-013-9679-4.
Kesherwani V, Shahshahan HR, Mishra PK. Cardiac transcriptome profiling of diabetic Akita mice using microarray and next generation sequencing. PLoS ONE. 2017;12(8):e0182828. https://doi.org/10.1371/journal.pone.0182828.
Belenchia AM, Gavini MP, Toedebusch RG, DeMarco VG, Pulakat L. Comparison of cardiac miRNA transcriptomes induced by diabetes and rapamycin treatment and identification of a rapamycin-associated cardiac MicroRNA signature. Oxid Med Cell Longev. 2018;2018:8364608. https://doi.org/10.1155/2018/8364608.
• Casero RA, Jr., Murray Stewart T, Pegg AE. Polyamine metabolism and cancer: treatments, challenges and opportunities. Nat Rev Cancer 2018;18(11):681–695. https://doi.org/10.1038/s41568-018-0050-3This is an important review on polyamine metabolism and cross-talk with other metabolic pathways.
•• Rosenheim O. The isolation of spermine phosphate from semen and testis. Biochem J. 1924;18(6):1253–1262 1251. https://doi.org/10.1042/bj0181253. This is the first paper that discovers polyamine.
Sakamoto A, Terui Y, Uemura T, Igarashi K, Kashiwagi K (2021) Translational regulation of clock genes BMAL1 and REV-ERBalpha by polyamines. Int J Mol Sci 22(3). https://doi.org/10.3390/ijms22031307
Ivanov IP, Shin BS, Loughran G, Tzani I, Young-Baird SK, Cao C, Atkins JF, Dever TE. Polyamine control of translation elongation regulates start site selection on antizyme inhibitor mRNA via ribosome queuing. Mol Cell. 2018;70(2):254-264 e256. https://doi.org/10.1016/j.molcel.2018.03.015.
Wang M, Phanstiel O, von Kalm L (2017) Evaluation of polyamine transport inhibitors in a drosophila epithelial model suggests the existence of multiple transport systems. Med Sci (Basel) 5 (4). https://doi.org/10.3390/medsci5040027
Uemura T, Stringer DE, Blohm-Mangone KA, Gerner EW. Polyamine transport is mediated by both endocytic and solute carrier transport mechanisms in the gastrointestinal tract. Am J Physiol Gastrointest Liver Physiol. 2010;299(2):G517-522. https://doi.org/10.1152/ajpgi.00169.2010.
Uemura T, Yerushalmi HF, Tsaprailis G, Stringer DE, Pastorian KE, Hawel L 3rd, Byus CV, Gerner EW. Identification and characterization of a diamine exporter in colon epithelial cells. J Biol Chem. 2008;283(39):26428–35. https://doi.org/10.1074/jbc.M804714200.
Hiasa M, Miyaji T, Haruna Y, Takeuchi T, Harada Y, Moriyama S, Yamamoto A, Omote H, Moriyama Y. Identification of a mammalian vesicular polyamine transporter. Sci Rep. 2014;4:6836. https://doi.org/10.1038/srep06836.
Ding K, Sandgren S, Mani K, Belting M, Fransson LA. Modulations of glypican-1 heparan sulfate structure by inhibition of endogenous polyamine synthesi. Mapping of spermine-binding sites and heparanase, heparin lyase, and nitric oxide/nitrite cleavage sites. J Biol Chem. 2001;276(50):46779–91. https://doi.org/10.1074/jbc.M105419200.
• Hamouda NN, Van den Haute C, Vanhoutte R, Sannerud R, Azfar M, Mayer R, Cortes Calabuig A, Swinnen JV, Agostinis P, Baekelandt V, Annaert W, Impens F, Verhelst SHL, Eggermont J, Martin S, Vangheluwe P (2021) ATP13A3 is a major component of the enigmatic mammalian polyamine transport system. J Biol Chem 296:100182. https://doi.org/10.1074/jbc.RA120.013908This paper establishes a new transporter of polyamines.
Seiler N, Sarhan S, Grauffel C, Jones R, Knodgen B, Moulinoux JP. Endogenous and exogenous polyamines in support of tumor growth. Cancer Res. 1990;50(16):5077–83.
Feng YL, Cao G, Chen DQ, Vaziri ND, Chen L, Zhang J, Wang M, Guo Y, Zhao YY. Microbiome-metabolomics reveals gut microbiota associated with glycine-conjugated metabolites and polyamine metabolism in chronic kidney disease. Cell Mol Life Sci. 2019;76(24):4961–78. https://doi.org/10.1007/s00018-019-03155-9.
Yang Y, Misra BB, Liang L, Bi D, Weng W, Wu W, Cai S, Qin H, Goel A, Li X, Ma Y. Integrated microbiome and metabolome analysis reveals a novel interplay between commensal bacteria and metabolites in colorectal cancer. Theranostics. 2019;9(14):4101–14. https://doi.org/10.7150/thno.35186.
de Oliveira LF, Navarro BV, Cerruti GV, Elbl P, Minocha R, Minocha SC, Dos Santos ALW, Floh EIS. Polyamine- and amino acid-related metabolism: the roles of arginine and ornithine are associated with the embryogenic potential. Plant Cell Physiol. 2018;59(5):1084–98. https://doi.org/10.1093/pcp/pcy049.
Morales TS, Avis EC, Paskowski EK, Shabar H, Nowotarski SL, DiAngelo JR (2021) The role of spermidine synthase (SpdS) and spermine synthase (Sms) in regulating triglyceride storage in Drosophila. Med Sci (Basel) 9 (2). doi:https://doi.org/10.3390/medsci9020027
Atawia RT, Bunch KL, Fouda AY, Lemtalsi T, Eldahshan W, Xu Z, Saul A, Elmasry K, Al-Shabrawey M, Caldwell RB, Caldwell RW (2020) Role of arginase 2 in murine retinopathy associated with Western diet-induced obesity. J Clin Med 9 (2). doi:https://doi.org/10.3390/jcm9020317
Chandra S, Fulton DJR, Caldwell RB, Caldwell RW, Toque HA. Hyperglycemia-impaired aortic vasorelaxation mediated through arginase elevation: role of stress kinase pathways. Eur J Pharmacol. 2019;844:26–37. https://doi.org/10.1016/j.ejphar.2018.11.027.
Rafnsson A, Matic LP, Lengquist M, Mahdi A, Shemyakin A, Paulsson-Berne G, Hansson GK, Gabrielsen A, Hedin U, Yang J, Pernow J. Endothelin-1 increases expression and activity of arginase 2 via ETB receptors and is co-expressed with arginase 2 in human atherosclerotic plaques. Atherosclerosis. 2020;292:215–23. https://doi.org/10.1016/j.atherosclerosis.2019.09.020.
Rodriguez-Guzman MJ, Peces-Barba Romero G, Perez Rial S, Serrano Del Castillo C, Palomero Rodriguez MA, Mahillo-Fernandez I, Villar-Alvarez F. Elevated levels of arginase activity are related to inflammation in patients with COPD exacerbation. BMC Pulm Med. 2021;21(1):271. https://doi.org/10.1186/s12890-021-01629-w.
Candadai AA, Liu F, Fouda AY, Alfarhan M, Palani CD, Xu Z, Caldwell RB, Narayanan SP. Deletion of arginase 2 attenuates neuroinflammation in an experimental model of optic neuritis. PLoS ONE. 2021;16(3):e0247901. https://doi.org/10.1371/journal.pone.0247901.
Nowotarski SL, Feehan RP, Presloid C, Shantz LM. Knockout of Raptor destabilizes ornithine decarboxylase mRNA and decreases binding of HuR to the ODC transcript in cells exposed to ultraviolet-B irradiation. Biochem Biophys Res Commun. 2018;505(4):1022–6. https://doi.org/10.1016/j.bbrc.2018.10.019.
Mai A, Nowotarski SL. Investigating ornithine decarboxylase posttranscriptional regulation via a pulldown assay using biotinylated transcripts. Methods Mol Biol. 2018;1694:299–308. https://doi.org/10.1007/978-1-4939-7398-9_25.
Soda K (2018) Polyamine metabolism and gene methylation in conjunction with one-carbon metabolism. Int J Mol Sci 19 (10). doi:https://doi.org/10.3390/ijms19103106
Kakehashi A, Suzuki S, Ishii N, Okuno T, Kuwae Y, Fujioka M, Gi M, Stefanov V, Wanibuchi H (2020) Accumulation of 8-hydroxydeoxyguanosine, L-arginine and glucose metabolites by liver tumor cells are the important characteristic features of metabolic syndrome and non-alcoholic steatohepatitis-associated hepatocarcinogenesis. Int J Mol Sci 21 (20). doi:https://doi.org/10.3390/ijms21207746
Lopez-Contreras F, Munoz-Uribe M, Perez-Laines J, Ascencio-Leal L, Rivera-Dictter A, Martin-Martin A, Burgos RA, Alarcon P, Lopez-Munoz R. Searching for drug synergy against cancer through polyamine metabolism impairment: insight into the metabolic effect of indomethacin on lung cancer cells. Front Pharmacol. 2019;10:1670. https://doi.org/10.3389/fphar.2019.01670.
• Affronti HC, Rowsam AM, Pellerite AJ, Rosario SR, Long MD, Jacobi JJ, Bianchi-Smiraglia A, Boerlin CS, Gillard BM, Karasik E, Foster BA, Moser M, Wilton JH, Attwood K, Nikiforov MA, Azabdaftari G, Pili R, Phillips JG, Casero RA, Jr., Smiraglia DJ (2020) Pharmacological polyamine catabolism upregulation with methionine salvage pathway inhibition as an effective prostate cancer therapy. Nat Commun 11 (1):52. https://doi.org/10.1038/s41467-019-13950-4. This paper inelaborates the therapeutic potential of maniulating endogenous polyamines.
Moinard C, Cynober L, de Bandt JP. Polyamines: metabolism and implications in human diseases. Clin Nutr. 2005;24(2):184–97. https://doi.org/10.1016/j.clnu.2004.11.001.
Minois N, Carmona-Gutierrez D, Madeo F. Polyamines in aging and disease Aging (Albany NY). 2011;3(8):716–32. https://doi.org/10.18632/aging.100361.
Hougaard DM, Nielsen JH, Larsson LI. Localization and biosynthesis of polyamines in insulin-producing cells. Biochem J. 1986;238(1):43–7. https://doi.org/10.1042/bj2380043.
Welsh N. A role for polyamines in glucose-stimulated insulin-gene expression. Biochem J. 1990;271(2):393–7. https://doi.org/10.1042/bj2710393.
Huber M, Poulin R. Permissive role of polyamines in the cooperative action of estrogens and insulin or insulin-like growth factor I on human breast cancer cell growth. J Clin Endocrinol Metab. 1996;81(1):113–23. https://doi.org/10.1210/jcem.81.1.8550737.
Niiranen K, Keinanen TA, Pirinen E, Heikkinen S, Tusa M, Fatrai S, Suppola S, Pietila M, Uimari A, Laakso M, Alhonen L, Janne J. Mice with targeted disruption of spermidine/spermine N1-acetyltransferase gene maintain nearly normal tissue polyamine homeostasis but show signs of insulin resistance upon aging. J Cell Mol Med. 2006;10(4):933–45. https://doi.org/10.1111/j.1582-4934.2006.tb00536.x.
Tersey SA, Colvin SC, Maier B, Mirmira RG. Protective effects of polyamine depletion in mouse models of type 1 diabetes: implications for therapy. Amino Acids. 2014;46(3):633–42. https://doi.org/10.1007/s00726-013-1560-7.
Levasseur EM, Yamada K, Pineros AR, Wu W, Syed F, Orr KS, Anderson-Baucum E, Mastracci TL, Maier B, Mosley AL, Liu Y, Bernal-Mizrachi E, Alonso LC, Scott D, Garcia-Ocana A, Tersey SA, Mirmira RG (2019) Hypusine biosynthesis in beta cells links polyamine metabolism to facultative cellular proliferation to maintain glucose homeostasis. Sci Signal 12 (610). doi:https://doi.org/10.1126/scisignal.aax0715
• Bjelakovic G, Beninati S, Bjelakovic B, Sokolovic D, Jevtovic T, Stojanovic I, Rossi S, Tabolacci C, Kocic G, Pavlovic D, Saranac L, Zivic S (2010) Does polyamine oxidase activity influence the oxidative metabolism of children who suffer of diabetes mellitus? Mol Cell Biochem 341 (1-2):79-85. doi:https://doi.org/10.1007/s11010-010-0439-0 (This is a clinical study on T1DM children demonstrating the effect of polyamine metabolism on oxidative stress.)
•• Fernandez-Garcia JC, Delpino-Rius A, Samarra I, Castellano-Castillo D, Munoz-Garach A, Bernal-Lopez MR, Queipo-Ortuno MI, Cardona F, Ramos-Molina B, Tinahones FJ (2019) Type 2 diabetes is associated with a different pattern of serum polyamines: a case(-)control study from the PREDIMED-Plus trial. J Clin Med 8 (1). https://doi.org/10.3390/jcm8010071. This clinical trial-based paper demonstrates dysregulated polyamine metabolism in T2DM and reveals association of serum polyamines putrescine and spermine with glyccemic control and insulin levels, respectively.
Lin Y, Zhang X, Wang L, Zhao Y, Li H, Xiao W, Xu C, Liu J (2014) Polyamine depletion attenuates isoproterenol-induced hypertrophy and endoplasmic reticulum stress in cardiomyocytes. Cell Physiol Biochem 34 (5):1455-1465. https://doi.org/10.1159/000366350. This paper shows the effect of polyamine levels in cardiomyocyte’s hypertrophy.
•• Zhao YJ, Xu CQ, Zhang WH, Zhang L, Bian SL, Huang Q, Sun HL, Li QF, Zhang YQ, Tian Y, Wang R, Yang BF, Li WM (2007) Role of polyamines in myocardial ischemia/reperfusion injury and their interactions with nitric oxide. Eur J Pharmacol 562 (3):236-246. https://doi.org/10.1016/j.ejphar.2007.01.096. This paper provides evidence for cardioprotective effects of polyamine spermine in preventing cell death in ischemia/reperfusion injury model.
• Nayak A, Liu C, Mehta A, Ko YA, Tahhan AS, Dhindsa DS, Uppal K, Jones DP, Butler J, Morris AA, Quyyumi AA (2020) N8-acetylspermidine: a polyamine biomarker in ischemic cardiomyopathy with reduced ejection fraction. J Am Heart Assoc 9 (11):e016055. doi:https://doi.org/10.1161/JAHA.120.016055. This paper revealed polyamines could be a potential biomarker for heart failure.
• Wei C, Li H, Wang Y, Peng X, Shao H, Li H, Bai S, Xu C (2016) Exogenous spermine inhibits hypoxia/ischemia-induced myocardial apoptosis via regulation of mitochondrial permeability transition pore and associated pathways. Exp Biol Med (Maywood) 241 (14):1505–1515. doi:https://doi.org/10.1177/1535370216643417. This paper demonstrates the role of polyamines in preventing mitochondrial damage and apoptosis in the heart.)
Wei C, Wang Y, Li M, Li H, Lu X, Shao H, Xu C. Spermine inhibits endoplasmic reticulum stress-induced apoptosis: a new strategy to prevent cardiomyocyte apoptosis. Cell Physiol Biochem. 2016;38(2):531–44. https://doi.org/10.1159/000438648.
Sheibani M, Nezamoleslami S, Mousavi SE, Faghir-Ghanesefat H, Yousefi-Manesh H, Rezayat SM, Dehpour A. Protective effects of spermidine against cirrhotic cardiomyopathy in bile duct-ligated rats. J Cardiovasc Pharmacol. 2020;76(3):286–95. https://doi.org/10.1097/FJC.0000000000000872.
• Chai N, Zhang H, Li L, Yu X, Liu Y, Lin Y, Wang L, Yan J, Nikolaevna SE, Zhao Y (2019) Spermidine prevents heart injury in neonatal rats exposed to intrauterine hypoxia by inhibiting oxidative stress and mitochondrial fragmentation. Oxid Med Cell Longev 2019:5406468. https://doi.org/10.1155/2019/5406468. This paper shows impact of polyamines on mitochodrial damage in the neonatal heart.
Zhang H, Wang J, Li L, Chai N, Chen Y, Wu F, Zhang W, Wang L, Shi S, Zhang L, Bian S, Xu C, Tian Y, Zhao Y. Spermine and spermidine reversed age-related cardiac deterioration in rats. Oncotarget. 2017;8(39):64793–808. https://doi.org/10.18632/oncotarget.18334.
•• Eisenberg T, Abdellatif M, Schroeder S, Primessnig U, Stekovic S, Pendl T, Harger A, Schipke J, Zimmermann A, Schmidt A, Tong M, Ruckenstuhl C, Dammbrueck C, Gross AS, Herbst V, Magnes C, Trausinger G, Narath S, Meinitzer A, Hu Z, Kirsch A, Eller K, Carmona-Gutierrez D, Buttner S, Pietrocola F, Knittelfelder O, Schrepfer E, Rockenfeller P, Simonini C, Rahn A, Horsch M, Moreth K, Beckers J, Fuchs H, Gailus-Durner V, Neff F, Janik D, Rathkolb B, Rozman J, de Angelis MH, Moustafa T, Haemmerle G, Mayr M, Willeit P, von Frieling-Salewsky M, Pieske B, Scorrano L, Pieber T, Pechlaner R, Willeit J, Sigrist SJ, Linke WA, Muhlfeld C, Sadoshima J, Dengjel J, Kiechl S, Kroemer G, Sedej S, Madeo F (2016) Cardioprotection and lifespan extension by the natural polyamine spermidine. Nat Med 22 (12):1428–1438. https://doi.org/10.1038/nm.4222. This paper provides evidence that dietary spermidine polyamine protects the heart by enhancing autophagy, mitophagy, and mitochondrial respiration, and also reduces systemic blood pressure in high-salt diet-induced hypertensive congestive heart failure.
Sansbury BE, DeMartino AM, Xie Z, Brooks AC, Brainard RE, Watson LJ, DeFilippis AP, Cummins TD, Harbeson MA, Brittian KR, Prabhu SD, Bhatnagar A, Jones SP, Hill BG. Metabolomic analysis of pressure-overloaded and infarcted mouse hearts. Circ Heart Fail. 2014;7(4):634–42. https://doi.org/10.1161/CIRCHEARTFAILURE.114.001151.
Halade GV, Kain V, Tourki B, Jadapalli JK. Lipoxygenase drives lipidomic and metabolic reprogramming in ischemic heart failure. Metabolism. 2019;96:22–32. https://doi.org/10.1016/j.metabol.2019.04.011.
Talman V, Teppo J, Poho P, Movahedi P, Vaikkinen A, Karhu ST, Trost K, Suvitaival T, Heikkonen J, Pahikkala T, Kotiaho T, Kostiainen R, Varjosalo M, Ruskoaho H. Molecular atlas of postnatal mouse heart development. J Am Heart Assoc. 2018;7(20):e010378. https://doi.org/10.1161/JAHA.118.010378.
Kannel WB, Hjortland M, Castelli WP. Role of diabetes in congestive heart failure: the Framingham study. Am J Cardiol. 1974;34(1):29–34. https://doi.org/10.1016/0002-9149(74)90089-7.
Rosengren A, Vestberg D, Svensson AM, Kosiborod M, Clements M, Rawshani A, Pivodic A, Gudbjornsdottir S, Lind M. Long-term excess risk of heart failure in people with type 1 diabetes: a prospective case-control study. Lancet Diabetes Endocrinol. 2015;3(11):876–85. https://doi.org/10.1016/S2213-8587(15)00292-2.
• Hu J, Lu X, Zhang X, Shao X, Wang Y, Chen J, Zhao B, Li S, Xu C, Wei C (2020) Exogenous spermine attenuates myocardial fibrosis in diabetic cardiomyopathy by inhibiting endoplasmic reticulum stress and the canonical Wnt signaling pathway. Cell Biol Int 44 (8):1660-1670. https://doi.org/10.1002/cbin.11360. This paper reveals the mechanism by which spermine polyamine mitigates adverse remodeling in the T1DM heart.
Bai SZ, Sun J, Wu H, Zhang N, Li HX, Li GW, Li HZ, He W, Zhang WH, Zhao YJ, Wang LN, Tian Y, Yang BF, Yang GD, Wu LY, Wang R, Xu CQ. Decrease in calcium-sensing receptor in the progress of diabetic cardiomyopathy. Diabetes Res Clin Pract. 2012;95(3):378–85. https://doi.org/10.1016/j.diabres.2011.11.007.
• Sochor M, Zaheer Baquer N, Hothersall JS, McLean P (1978) Effect of experimental diabetes on ornithine decarboxylase activity of rat tissues. Biochem Biophys Res Commun 80 (3):533-539. https://doi.org/10.1016/0006-291x(78)91601-7. This classical paper showed the impact of alloxan-induced T1DM on levels of ornithin decarboxylase in insulin sensitive liver and heart and insulin independent brain and kidney tissues.
Paxton R, Ye L. Regulation of heart insulin receptor tyrosine kinase activity by magnesium and spermine. Mol Cell Biochem. 2005;277(1–2):7–17. https://doi.org/10.1007/s11010-005-5755-4.
Zhou L, Sun CB, Liu C, Fan Y, Zhu HY, Wu XW, Hu L, Li QP. Upregulation of arginase activity contributes to intracellular ROS production induced by high glucose in H9c2 cells. Int J Clin Exp Pathol. 2015;8(3):2728–36.
Kiss A, Tratsiakovich Y, Gonon AT, Fedotovskaya O, Lanner JT, Andersson DC, Yang J, Pernow J. The role of arginase and rho kinase in cardioprotection from remote ischemic perconditioning in non-diabetic and diabetic rat in vivo. PLoS ONE. 2014;9(8):e104731. https://doi.org/10.1371/journal.pone.0104731.
Muhlfeld C, Pfeiffer C, Schneider V, Bornemann M, Schipke J. Voluntary activity reverses spermidine-induced myocardial fibrosis and lipid accumulation in the obese male mouse. Histochem Cell Biol. 2021;155(1):75–88. https://doi.org/10.1007/s00418-020-01926-1.
• Chen HY, Jia XL, Zhao SQ, Zheng WH, Mei ZG, Yang HW, Zhang SZ (2019) Dual role of polyamines in heart ischemia/reperfusion injury through regulation of mitochondrial permeability transition pore. Sheng Li Xue Bao 71 (5):681–688. This paper shows the effect of polyamine treatment on mitochondrial permeability transition pore opening in the ischemia/reperfusion injury heart.
Ludwig N, Leidinger P, Becker K, Backes C, Fehlmann T, Pallasch C, Rheinheimer S, Meder B, Stahler C, Meese E, Keller A. Distribution of miRNA expression across human tissues. Nucleic Acids Res. 2016;44(8):3865–77. https://doi.org/10.1093/nar/gkw116.
Tomasi ML, Cossu C, Spissu Y, Floris A, Ryoo M, Iglesias-Ara A, Wang Q, Pandol SJ, Bhowmick NA, Seki E, Posadas EM, Lu SC. S-adenosylmethionine and methylthioadenosine inhibit cancer metastasis by targeting microRNA 34a/b-methionine adenosyltransferase 2A/2B axis. Oncotarget. 2017;8(45):78851–69. https://doi.org/10.18632/oncotarget.20234.
Zou T, Rao JN, Liu L, Xiao L, Chung HK, Li Y, Chen G, Gorospe M, Wang JY. JunD enhances miR-29b levels transcriptionally and posttranscriptionally to inhibit proliferation of intestinal epithelial cells. Am J Physiol Cell Physiol. 2015;308(10):C813-824. https://doi.org/10.1152/ajpcell.00027.2015.
Coppola A, Ilisso CP, Stellavato A, Schiraldi C, Caraglia M, Mosca L, Cacciapuoti G, Porcelli M (2020) S-adenosylmethionine inhibits cell growth and migration of triple negative breast cancer cells through upregulating MiRNA-34c and MiRNA-449a. Int J Mol Sci 22(1). doi:https://doi.org/10.3390/ijms22010286
Fomison-Nurse I, Saw EEL, Gandhi S, Munasinghe PE, Van Hout I, Williams MJA, Galvin I, Bunton R, Davis P, Cameron V, Katare R. Diabetes induces the activation of pro-ageing miR-34a in the heart, but has differential effects on cardiomyocytes and cardiac progenitor cells. Cell Death Differ. 2018;25(7):1336–49. https://doi.org/10.1038/s41418-017-0047-6.
Greco S, Fasanaro P, Castelvecchio S, D’Alessandra Y, Arcelli D, Di Donato M, Malavazos A, Capogrossi MC, Menicanti L, Martelli F. MicroRNA dysregulation in diabetic ischemic heart failure patients. Diabetes. 2012;61(6):1633–41. https://doi.org/10.2337/db11-0952.
Nakada C, Hijiya N, Tsukamoto Y, Yano S, Kai T, Uchida T, Kimoto M, Takahashi M, Daa T, Matsuura K, Shin T, Mimata H, Moriyama M. A transgenic mouse expressing miR-210 in proximal tubule cells shows mitochondrial alteration: possible association of miR-210 with a shift in energy metabolism. J Pathol. 2020;251(1):12–25. https://doi.org/10.1002/path.5394.
Staedel C, Tran TPA, Giraud J, Darfeuille F, Di Giorgio A, Tourasse NJ, Salin F, Uriac P, Duca M. Modulation of oncogenic miRNA biogenesis using functionalized polyamines. Sci Rep. 2018;8(1):1667. https://doi.org/10.1038/s41598-018-20053-5.
Yang H, Cho ME, Li TW, Peng H, Ko KS, Mato JM, Lu SC. MicroRNAs regulate methionine adenosyltransferase 1A expression in hepatocellular carcinoma. J Clin Invest. 2013;123(1):285–98. https://doi.org/10.1172/JCI63861.
Xu L, Chen W, Ma M, Chen A, Tang C, Zhang C, Cai L. Microarray profiling analysis identifies the mechanism of miR-200b-3p/mRNA-CD36 affecting diabetic cardiomyopathy via peroxisome proliferator activated receptor-gamma signaling pathway. J Cell Biochem. 2019;120(4):5193–206. https://doi.org/10.1002/jcb.27795.
Singh GB, Raut SK, Khanna S, Kumar A, Sharma S, Prasad R, Khullar M. MicroRNA-200c modulates DUSP-1 expression in diabetes-induced cardiac hypertrophy. Mol Cell Biochem. 2017;424(1–2):1–11. https://doi.org/10.1007/s11010-016-2838-3.
Humphries B, Yang C. The microRNA-200 family: small molecules with novel roles in cancer development, progression and therapy. Oncotarget. 2015;6(9):6472–98. https://doi.org/10.18632/oncotarget.3052.
Al-Kafaji G, Al-Muhtaresh HA, Salem AH. Expression and clinical significance of miR-1 and miR-133 in pre-diabetes. Biomed Rep. 2021;14(3):33. https://doi.org/10.3892/br.2021.1409.
•• Castagno D, Baird-Gunning J, Jhund PS, Biondi-Zoccai G, MacDonald MR, Petrie MC, Gaita F, McMurray JJ (2011) Intensive glycemic control has no impact on the risk of heart failure in type 2 diabetic patients: evidence from a 37,229 patient meta-analysis. Am Heart J 162 (5):938–948 e932. https://doi.org/10.1016/j.ahj.2011.07.030.This paper provides evidence from clinical trials that tight controlling of blood glucose levels does not reduce the risk of heart failure in diabetic patients.
Jeyabal P, Thandavarayan RA, Joladarashi D, Suresh Babu S, Krishnamurthy S, Bhimaraj A, Youker KA, Kishore R, Krishnamurthy P. MicroRNA-9 inhibits hyperglycemia-induced pyroptosis in human ventricular cardiomyocytes by targeting ELAVL1. Biochem Biophys Res Commun. 2016;471(4):423–9. https://doi.org/10.1016/j.bbrc.2016.02.065.
Torella D, Ellison GM, Torella M, Vicinanza C, Aquila I, Iaconetti C, Scalise M, Marino F, Henning BJ, Lewis FC, Gareri C, Lascar N, Cuda G, Salvatore T, Nappi G, Indolfi C, Torella R, Cozzolino D, Sasso FC. Carbonic anhydrase activation is associated with worsened pathological remodeling in human ischemic diabetic cardiomyopathy. J Am Heart Assoc. 2014;3(2):e000434. https://doi.org/10.1161/JAHA.113.000434.
Schellinger IN, Wagenhauser M, Chodisetti G, Mattern K, Dannert A, Petzold A, Jakubizka-Smorag J, Emrich F, Haunschild J, Schuster A, Schwob E, Schulz K, Maegdefessel L, Spin JM, Stumvoll M, Hasenfuss G, Tsao PS, Raaz U. MicroRNA miR-29b regulates diabetic aortic remodeling and stiffening. Mol Ther Nucleic Acids. 2021;24:188–99. https://doi.org/10.1016/j.omtn.2021.02.021.
Ni T, Lin N, Lu W, Sun Z, Lin H, Chi J, Guo H. Dihydromyricetin prevents diabetic cardiomyopathy via miR-34a suppression by activating autophagy. Cardiovasc Drugs Ther. 2020;34(3):291–301. https://doi.org/10.1007/s10557-020-06968-0.
Jansen F, Wang H, Przybilla D, Franklin BS, Dolf A, Pfeifer P, Schmitz T, Flender A, Endl E, Nickenig G, Werner N. Vascular endothelial microparticles-incorporated microRNAs are altered in patients with diabetes mellitus. Cardiovasc Diabetol. 2016;15:49. https://doi.org/10.1186/s12933-016-0367-8.
Rawal S, Munasinghe PE, Shindikar A, Paulin J, Cameron V, Manning P, Williams MJ, Jones GT, Bunton R, Galvin I, Katare R. Down-regulation of proangiogenic microRNA-126 and microRNA-132 are early modulators of diabetic cardiac microangiopathy. Cardiovasc Res. 2017;113(1):90–101. https://doi.org/10.1093/cvr/cvw235.
Nandi SS, Duryee MJ, Shahshahan HR, Thiele GM, Anderson DR, Mishra PK. Induction of autophagy markers is associated with attenuation of miR-133a in diabetic heart failure patients undergoing mechanical unloading. Am J Transl Res. 2015;7(4):683–96.
Raut SK, Singh GB, Rastogi B, Saikia UN, Mittal A, Dogra N, Singh S, Prasad R, Khullar M. miR-30c and miR-181a synergistically modulate p53–p21 pathway in diabetes induced cardiac hypertrophy. Mol Cell Biochem. 2016;417(1–2):191–203. https://doi.org/10.1007/s11010-016-2729-7.
Rawal S, Nagesh PT, Coffey S, Van Hout I, Galvin IF, Bunton RW, Davis P, Williams MJA, Katare R. Early dysregulation of cardiac-specific microRNA-208a is linked to maladaptive cardiac remodelling in diabetic myocardium. Cardiovasc Diabetol. 2019;18(1):13. https://doi.org/10.1186/s12933-019-0814-4.
Lu H, Buchan RJ, Cook SA. MicroRNA-223 regulates Glut4 expression and cardiomyocyte glucose metabolism. Cardiovasc Res. 2010;86(3):410–20. https://doi.org/10.1093/cvr/cvq010.
Li H, Fan J, Zhao Y, Zhang X, Dai B, Zhan J, Yin Z, Nie X, Fu XD, Chen C, Wang DW. Nuclear miR-320 mediates diabetes-induced cardiac dysfunction by activating transcription of fatty acid metabolic genes to cause lipotoxicity in the heart. Circ Res. 2019;125(12):1106–20. https://doi.org/10.1161/CIRCRESAHA.119.314898.
Shan ZX, Lin QX, Deng CY, Zhu JN, Mai LP, Liu JL, Fu YH, Liu XY, Li YX, Zhang YY, Lin SG, Yu XY. miR-1/miR-206 regulate Hsp60 expression contributing to glucose-mediated apoptosis in cardiomyocytes. FEBS Lett. 2010;584(16):3592–600. https://doi.org/10.1016/j.febslet.2010.07.027.
Katare R, Caporali A, Zentilin L, Avolio E, Sala-Newby G, Oikawa A, Cesselli D, Beltrami AP, Giacca M, Emanueli C, Madeddu P. Intravenous gene therapy with PIM-1 via a cardiotropic viral vector halts the progression of diabetic cardiomyopathy through promotion of prosurvival signaling. Circ Res. 2011;108(10):1238–51. https://doi.org/10.1161/CIRCRESAHA.110.239111.
Gao L, Wang X, Guo S, Xiao L, Liang C, Wang Z, Li Y, Liu Y, Yao R, Liu Y, Zhang Y. LncRNA HOTAIR functions as a competing endogenous RNA to upregulate SIRT1 by sponging miR-34a in diabetic cardiomyopathy. J Cell Physiol. 2019;234(4):4944–58. https://doi.org/10.1002/jcp.27296.
Gao L, Liu Y, Guo S, Xiao L, Wu L, Wang Z, Liang C, Yao R. Zhang Y (2018) LAZ3 protects cardiac remodeling in diabetic cardiomyopathy via regulating miR-21/PPARa signaling. Biochim Biophys Acta Mol Basis Dis. 1864;10:3322–38. https://doi.org/10.1016/j.bbadis.2018.07.019.
Zhu C, Zhang H, Wei D, Sun Z (2021) Silencing lncRNA GAS5 alleviates apoptosis and fibrosis in diabetic cardiomyopathy by targeting miR-26a/b-5p. Acta Diabetol. https://doi.org/10.1007/s00592-021-01745-3
Zhang Y, Wang JH, Zhang YY, Wang YZ, Wang J, Zhao Y, Jin XX, Xue GL, Li PH, Sun YL, Huang QH, Song XT, Zhang ZR, Gao X, Yang BF, Du ZM, Pan ZW. Deletion of interleukin-6 alleviated interstitial fibrosis in streptozotocin-induced diabetic cardiomyopathy of mice through affecting TGFbeta1 and miR-29 pathways. Sci Rep. 2016;6:23010. https://doi.org/10.1038/srep23010.
Yin Z, Zhao Y, He M, Li H, Fan J, Nie X, Yan M, Chen C, Wang DW. MiR-30c/PGC-1beta protects against diabetic cardiomyopathy via PPARalpha. Cardiovasc Diabetol. 2019;18(1):7. https://doi.org/10.1186/s12933-019-0811-7.
Li X, Du N, Zhang Q, Li J, Chen X, Liu X, Hu Y, Qin W, Shen N, Xu C, Fang Z, Wei Y, Wang R, Du Z, Zhang Y, Lu Y. MicroRNA-30d regulates cardiomyocyte pyroptosis by directly targeting foxo3a in diabetic cardiomyopathy. Cell Death Dis. 2014;5:e1479. https://doi.org/10.1038/cddis.2014.430.
Zhao F, Li B, Wei YZ, Zhou B, Wang H, Chen M, Gan XD, Wang ZH, Xiong SX. MicroRNA-34a regulates high glucose-induced apoptosis in H9c2 cardiomyocytes. J Huazhong Univ Sci Technolog Med Sci. 2013;33(6):834–9. https://doi.org/10.1007/s11596-013-1207-7.
Feng B, Chen S, George B, Feng Q, Chakrabarti S. miR133a regulates cardiomyocyte hypertrophy in diabetes. Diabetes Metab Res Rev. 2010;26(1):40–9. https://doi.org/10.1002/dmrr.1054.
Che H, Wang Y, Li H, Li Y, Sahil A, Lv J, Liu Y, Yang Z, Dong R, Xue H, Wang L. Melatonin alleviates cardiac fibrosis via inhibiting lncRNA MALAT1/miR-141-mediated NLRP3 inflammasome and TGF-beta1/Smads signaling in diabetic cardiomyopathy. FASEB J. 2020;34(4):5282–98. https://doi.org/10.1096/fj.201902692R.
Yu M, Liu Y, Zhang B, Shi Y, Cui L, Zhao X. Inhibiting microRNA-144 abates oxidative stress and reduces apoptosis in hearts of streptozotocin-induced diabetic mice. Cardiovasc Pathol. 2015;24(6):375–81. https://doi.org/10.1016/j.carpath.2015.06.003.
Duan Y, Zhou B, Su H, Liu Y, Du C. miR-150 regulates high glucose-induced cardiomyocyte hypertrophy by targeting the transcriptional co-activator p300. Exp Cell Res. 2013;319(3):173–84. https://doi.org/10.1016/j.yexcr.2012.11.015.
Zheng D, Ma J, Yu Y, Li M, Ni R, Wang G, Chen R, Li J, Fan GC, Lacefield JC, Peng T. Silencing of miR-195 reduces diabetic cardiomyopathy in C57BL/6 mice. Diabetologia. 2015;58(8):1949–58. https://doi.org/10.1007/s00125-015-3622-8.
Wang XH, Qian RZ, Zhang W, Chen SF, Jin HM, Hu RM. MicroRNA-320 expression in myocardial microvascular endothelial cells and its relationship with insulin-like growth factor-1 in type 2 diabetic rats. Clin Exp Pharmacol Physiol. 2009;36(2):181–8. https://doi.org/10.1111/j.1440-1681.2008.05057.x.
Shen E, Diao X, Wang X, Chen R, Hu B. MicroRNAs involved in the mitogen-activated protein kinase cascades pathway during glucose-induced cardiomyocyte hypertrophy. Am J Pathol. 2011;179(2):639–50. https://doi.org/10.1016/j.ajpath.2011.04.034.
Kuwabara Y, Horie T, Baba O, Watanabe S, Nishiga M, Usami S, Izuhara M, Nakao T, Nishino T, Otsu K, Kita T, Kimura T, Ono K. MicroRNA-451 exacerbates lipotoxicity in cardiac myocytes and high-fat diet-induced cardiac hypertrophy in mice through suppression of the LKB1/AMPK pathway. Circ Res. 2015;116(2):279–88. https://doi.org/10.1161/CIRCRESAHA.116.304707.
Qiao Y, Zhao Y, Liu Y, Ma N, Wang C, Zou J, Liu Z, Zhou Z, Han D, He J, Sun Q, Liu Y, Xu C, Du Z, Huang H. miR-483-3p regulates hyperglycaemia-induced cardiomyocyte apoptosis in transgenic mice. Biochem Biophys Res Commun. 2016;477(4):541–7. https://doi.org/10.1016/j.bbrc.2016.06.051.
Grimson A, Farh KK, Johnston WK, Garrett-Engele P, Lim LP, Bartel DP. MicroRNA targeting specificity in mammals: determinants beyond seed pairing. Mol Cell. 2007;27(1):91–105. https://doi.org/10.1016/j.molcel.2007.06.017.
Kameswaran V, Bramswig NC, McKenna LB, Penn M, Schug J, Hand NJ, Chen Y, Choi I, Vourekas A, Won KJ, Liu C, Vivek K, Naji A, Friedman JR, Kaestner KH. Epigenetic regulation of the DLK1-MEG3 microRNA cluster in human type 2 diabetic islets. Cell Metab. 2014;19(1):135–45. https://doi.org/10.1016/j.cmet.2013.11.016.
Haecker I, Gay LA, Yang Y, Hu J, Morse AM, McIntyre LM, Renne R. Ago HITS-CLIP expands understanding of Kaposi’s sarcoma-associated herpesvirus miRNA function in primary effusion lymphomas. PLoS Pathog. 2012;8(8):e1002884. https://doi.org/10.1371/journal.ppat.1002884.
Eichhorn SW, Guo H, McGeary SE, Rodriguez-Mias RA, Shin C, Baek D, Hsu SH, Ghoshal K, Villen J, Bartel DP. mRNA destabilization is the dominant effect of mammalian microRNAs by the time substantial repression ensues. Mol Cell. 2014;56(1):104–15. https://doi.org/10.1016/j.molcel.2014.08.028.
Karginov FV, Hannon GJ. Remodeling of Ago2-mRNA interactions upon cellular stress reflects miRNA complementarity and correlates with altered translation rates. Genes Dev. 2013;27(14):1624–32. https://doi.org/10.1101/gad.215939.113.
Lechman ER, Gentner B, Ng SWK, Schoof EM, van Galen P, Kennedy JA, Nucera S, Ciceri F, Kaufmann KB, Takayama N, Dobson SM, Trotman-Grant A, Krivdova G, Elzinga J, Mitchell A, Nilsson B, Hermans KG, Eppert K, Marke R, Isserlin R, Voisin V, Bader GD, Zandstra PW, Golub TR, Ebert BL, Lu J, Minden M, Wang JCY, Naldini L, Dick JE. miR-126 regulates distinct self-renewal outcomes in normal and malignant hematopoietic stem cells. Cancer Cell. 2016;29(4):602–6. https://doi.org/10.1016/j.ccell.2016.03.015.
Navarro F, Gutman D, Meire E, Caceres M, Rigoutsos I, Bentwich Z, Lieberman J. miR-34a contributes to megakaryocytic differentiation of K562 cells independently of p53. Blood. 2009;114(10):2181–92. https://doi.org/10.1182/blood-2009-02-205062.
Kishore S, Jaskiewicz L, Burger L, Hausser J, Khorshid M, Zavolan M. A quantitative analysis of CLIP methods for identifying binding sites of RNA-binding proteins. Nat Methods. 2011;8(7):559–64. https://doi.org/10.1038/nmeth.1608.
Hafner M, Landthaler M, Burger L, Khorshid M, Hausser J, Berninger P, Rothballer A, Ascano M Jr, Jungkamp AC, Munschauer M, Ulrich A, Wardle GS, Dewell S, Zavolan M, Tuschl T. Transcriptome-wide identification of RNA-binding protein and microRNA target sites by PAR-CLIP. Cell. 2010;141(1):129–41. https://doi.org/10.1016/j.cell.2010.03.009.
Balakrishnan I, Yang X, Brown J, Ramakrishnan A, Torok-Storb B, Kabos P, Hesselberth JR, Pillai MM. Genome-wide analysis of miRNA-mRNA interactions in marrow stromal cells. Stem Cells. 2014;32(3):662–73. https://doi.org/10.1002/stem.1531.
Boudreau RL, Jiang P, Gilmore BL, Spengler RM, Tirabassi R, Nelson JA, Ross CA, Xing Y, Davidson BL. Transcriptome-wide discovery of microRNA binding sites in human brain. Neuron. 2014;81(2):294–305. https://doi.org/10.1016/j.neuron.2013.10.062.
Kiga K, Mimuro H, Suzuki M, Shinozaki-Ushiku A, Kobayashi T, Sanada T, Kim M, Ogawa M, Iwasaki YW, Kayo H, Fukuda-Yuzawa Y, Yashiro M, Fukayama M, Fukao T, Sasakawa C. Epigenetic silencing of miR-210 increases the proliferation of gastric epithelium during chronic Helicobacter pylori infection. Nat Commun. 2014;5:4497. https://doi.org/10.1038/ncomms5497.
Romay MC, Che N, Becker SN, Pouldar D, Hagopian R, Xiao X, Lusis AJ, Berliner JA, Civelek M. Regulation of NF-kappaB signaling by oxidized glycerophospholipid and IL-1beta induced miRs-21-3p and -27a-5p in human aortic endothelial cells. J Lipid Res. 2015;56(1):38–50. https://doi.org/10.1194/jlr.M052670.
Chang TC, Wentzel EA, Kent OA, Ramachandran K, Mullendore M, Lee KH, Feldmann G, Yamakuchi M, Ferlito M, Lowenstein CJ, Arking DE, Beer MA, Maitra A, Mendell JT. Transactivation of miR-34a by p53 broadly influences gene expression and promotes apoptosis. Mol Cell. 2007;26(5):745–52. https://doi.org/10.1016/j.molcel.2007.05.010.
Pillai MM, Gillen AE, Yamamoto TM, Kline E, Brown J, Flory K, Hesselberth JR, Kabos P. HITS-CLIP reveals key regulators of nuclear receptor signaling in breast cancer. Breast Cancer Res Treat. 2014;146(1):85–97. https://doi.org/10.1007/s10549-014-3004-9.
Xue Y, Ouyang K, Huang J, Zhou Y, Ouyang H, Li H, Wang G, Wu Q, Wei C, Bi Y, Jiang L, Cai Z, Sun H, Zhang K, Zhang Y, Chen J, Fu XD. Direct conversion of fibroblasts to neurons by reprogramming PTB-regulated microRNA circuits. Cell. 2013;152(1–2):82–96. https://doi.org/10.1016/j.cell.2012.11.045.
Li J, Wan Y, Guo Q, Zou L, Zhang J, Fang Y, Zhang J, Zhang J, Fu X, Liu H, Lu L, Wu Y. Altered microRNA expression profile with miR-146a upregulation in CD4+ T cells from patients with rheumatoid arthritis. Arthritis Res Ther. 2010;12(3):R81. https://doi.org/10.1186/ar3006.
• Lopez JP, Fiori LM, Gross JA, Labonte B, Yerko V, Mechawar N, Turecki G (2014) Regulatory role of miRNAs in polyamine gene expression in the prefrontal cortex of depressed suicide completers. Int J Neuropsychopharmacol 17 (1):23–32. doi:https://doi.org/10.1017/S1461145713000941. (This paper elaborates relationship between miRNA and polyamine gene expression in the suicide brain.)
Funding
This work was supported in part by the National Institute of General Medical Sciences 1U54GM115458, National Institutes of Health (NIH) grant P20GM104320, UNMC Center for Heart and Vascular Research, and Nebraska Center for the Prevention of Obesity Diseases to PKM, GM103427 (INBRE grant) to SC, Nebraska Research Initiative Collaborative Seed Grant to PKM and SC, and NIH F31 HL156402 to TNK. HT was supported as graduate research assistant provided by the Division of Research and the Office of Graduate Studies and Academic Outreach at University of Nebraska at Kearney. The content is solely the responsibility of the author and does not necessarily represent the official view of the funding agencies.
Author information
Authors and Affiliations
Contributions
SC and PKM conceptualized the manuscript, TNK and HMNT made initial draft where TNK contributed to DMCM and miRNA, and HMNT contributed to polyamines section, HMNT prepared Fig. 1 and Table 1, FIG prepared Fig. 2, TNK prepared Fig. 3, TNK and FIG designed Tables 2 and 3, and TNK created Table 4. Both SC and PKM prepared final manuscript. All authors read the manuscript and authorize submission.
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article is part of the Topical collection on Macrovascular Complications in Diabetes
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kambis, T.N., Tofilau, H.M.N., Gawargi, F.I. et al. Regulating Polyamine Metabolism by miRNAs in Diabetic Cardiomyopathy. Curr Diab Rep 21, 52 (2021). https://doi.org/10.1007/s11892-021-01429-w
Accepted:
Published:
DOI: https://doi.org/10.1007/s11892-021-01429-w