Abstract
Fueled by the successes of genome-wide association studies, numerous studies have investigated the predictive ability of genetic risk models in type 2 diabetes. In this paper, we review these studies from a methodological perspective, focusing on the variables included in the risk models as well as the study designs and populations investigated. We argue and show that differences in study design and characteristics of the study population have an impact on the observed predictive ability of risk models. This observation emphasizes that genetic risk prediction studies should be conducted in those populations in which the prediction models will ultimately be applied, if proven useful. Of all genetic risk prediction studies to date, only a few were conducted in populations that might be relevant for targeting preventive interventions.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Type 2 diabetes (T2D) is a multifactorial disease, caused by a complex interplay between genetic and nongenetic risk factors. Compelling evidence has identified increasing age, higher body mass index (BMI), impaired fasting glucose, impaired glucose tolerance, higher glycated hemoglobin (HbA1c) level, and metabolic syndrome as important T2D risk factors (Table 1) [1–10]. These nongenetic factors have a substantial impact on disease risk and are frequent. For example, metabolic syndrome poses an eight times higher T2D risk and is present in more than 40% of the individuals over 50 years of age. The high impact and frequency make these risk factors suitable candidates for targeting preventive interventions, such as medication, weight loss, and increased physical activity that can slow down or even reverse the disease process [11, 12].
In the past 5 years, genome-wide association studies have identified and replicated over 40 single nucleotide polymorphisms (SNPs) that predispose to T2D [13, 14]. However, the effect sizes of the associated variants are very modest, with per allele odds ratios ranging from 1.05 to 1.35 [13]. Even the strongest susceptibility variant, rs7903146 in the TCF7L2 gene, is a weaker predictor of T2D risk than most nongenetic risk factors. Evidently, the low effect sizes make single genetic risk factors unsuitable for targeting preventive interventions, but there is increasing interest in investigating the extent to which genetic risk factors combined can improve the prediction of the disease.
An improvement in the early identification of high-risk groups is warranted because T2D imposes a great burden on human health and health care systems [15, 16]. An estimated 285 million people worldwide have diabetes [15] and this number is expected to increase by more than 50% in the next 20 years if no preventive strategies are implemented [15]. To identify high-risk individuals, many risk prediction models have been proposed.
Guidelines for T2D prevention advocate the use of clinical risk scores as primary screening tools, followed by blood glucose measurements to detect individuals with impaired fasting glucose, impaired glucose tolerance, or metabolic syndrome [17]. Examples of commonly used risk scores include the FINDRISC (Finnish Diabetes Risk Score) and the Diabetes Risk Calculator [18, 19]. The FINDRISC score is based on age, BMI, waist circumference, use of antihypertensive medication, history of elevated blood glucose, daily physical activity and daily intake of fruits or vegetables, and the Diabetes Risk Calculator on age, waist circumference, gestational diabetes, height, race/ethnicity, hypertension, family history of diabetes, and exercise.
The predictive ability of these clinical risk scores is modest, but satisfactory. The area under the receiver operating characteristic curve (AUC) is a commonly used measure to indicate the predictive ability; the AUC indicates the discriminative accuracy of a prediction model. To generate the curve, on the x-axis 1-specificity is plotted, and on the y-axis sensitivity is plotted. The AUC value represents the probability that the predicted risk of a random “patient” is higher than that of a random “nonpatient.” When predicted risks of individuals who will develop the disease are always higher than the risks of those who will not develop the disease, the AUC is 1.0. When their risks are higher for 50% of the random pairs, the AUC is 0.50, equaling the predictive performance of tossing a coin [20].
The AUC was 0.65 in men and 0.66 in women for the FINDRISC score predicting impaired fasting glucose, impaired glucose tolerance, or undiagnosed diabetes, and 0.72 and 0.75 for detecting metabolic syndrome [18]. The AUC of the Diabetes Risk Calculator was 0.70 for detecting impaired fasting glucose, impaired glucose tolerance, or undiagnosed diabetes [19]. These modest AUC values indicate that many people who will develop T2D are not identified as being at increased risk by these risk scores, and that many that will not develop the disease are labeled as increased risk. Although offering lifestyle modification programs to individuals who will not develop T2D may do no harm and may even provide other benefits by reducing the risk of other diseases, not recognizing the many who will develop diabetes would clearly be missed opportunities to reduce the serious burden of disease [12]. Some clinical risk models that include invasive measurements showed higher AUC values for detecting individuals who will develop T2D. An example is the Framingham Risk Score including age, sex, obesity, hypertension, parental history of diabetes, low levels of high-density lipoprotein cholesterol, elevated triglyceride levels, and impaired fasting glucose [21]. The AUC of this risk model was 0.85 for predicting T2D in middle-aged adults [21]. However, inclusion of invasive measurements that can change over time in clinical risk models might be inconvenient at the population level and these models still leave room for improvement.
Recent studies have investigated the predictive ability of risk models that include genetic variants only or genetic variants added to clinical risk factors. A study that investigated a genetic risk score based on 34 diabetes-associated variants showed a significant association of the risk score with risk of developing diabetes [22•]. This risk was attenuated by lifestyle interventions, also in individuals in the highest genetic risk quartile, suggesting that detecting individuals at high risk of developing T2D based on genetic variants and offering them lifestyle modification programs is useful. In this paper, we review genetic risk prediction studies from a methodologic perspective by focusing on factors in the choice of study design and population that may have impacted the observed predictive ability.
Genetic Risk Prediction Studies
The number of studies that investigate the predictive ability of genetic variants in T2D has increased rapidly (Table 2; [23, 24•, 25–38, 39•, 40–42, 43•]). These studies assessed risk models that were based on genetic variants only or on a combination of both genetic and nongenetic variants. The table shows that the number of SNPs included in the genetic models has increased from 3 in 2005 to 40 in 2011. The models show considerable overlap in the genetic variants that were considered, but there also are many differences. Since its discovery, all but one of the studies had included TCF7L2 and the majority additionally investigated PPARG, CDKN2A/B, KCNJ11, IGF2BP2, SLC30A8, and HHEX-IDE-KIF11. Yet, most other SNPs were included in one or two models only [43•]. The same was observed for the clinical models. Most clinical models included at least age, sex, and BMI, but they differed in the other factors that were added, such as blood pressure, family history of T2D, and fasting plasma glucose level.
Table 2 shows that, almost without exception, the genetic risk models had lower AUC values than the clinical models. The AUC values for the genetic models ranged from 0.55 to 0.68 and for the clinical models from 0.61 to 0.92. Table 2 also shows that the addition of genetic factors either did not or only marginally improved the AUC beyond that of the clinical risk models.
Predictive Ability of Clinical Risk Models
The differences in the predictive ability of clinical risk models are explained by how many and which risk factors are included in the model and by differences in study design and study population. This is nicely illustrated by three studies that had investigated largely the same 18 genetic variants. The AUCs of the genetic risks models in these studies were similar (0.58–0.60), but the AUCs of the clinical models were 0.66, 0.78, and 0.90 [33–35]. The clinical models with AUC values of 0.66 and 0.78 included age, sex, and BMI, but the model with an AUC value of 0.90 also included T2D family history, fasting plasma glucose, systolic blood pressure, high-density lipoprotein cholesterol, and triglycerides. The excellent predictive ability was likely due to the inclusion of fasting plasma glucose, as individuals with impaired fasting glucose have a very high risk of developing T2D (Table 1). Table 2 shows that AUC values tend to be higher when more risk factors are included in the model, particularly when fasting plasma glucose was included.
Yet, also the two studies that both investigated age, sex, and BMI in the clinical model had markedly different AUC values (0.66 and 0.78). The difference in these AUC values was likely explained by differences in the study design and population. The AUC of 0.66 was obtained in a prospective cohort study, the Rotterdam Study, and the AUC of 0.78 in a case–control study, consisting of case and control subjects from the GoDARTS (Genetics of Diabetes Audit and Research Tayside Study). Participants in the Rotterdam Study were older and less often men (Table 2), but the two populations predominantly differed in BMI. The mean BMI of the cases in the GoDARTS study was higher than the mean BMI of cases in the Rotterdam Study (31.5 vs 28.0 kg/m2). Also, the difference in mean BMI between cases and controls was much larger in the GoDARTS study compared with the Rotterdam Study (4.6 vs 2.0 kg/m2). In general and by definition, the predictive ability of risk models is higher when there are larger differences between cases and controls on the risk factors included in the risk model. Along the same lines, study design and population characteristics may have influenced the observed AUC values of the other clinical models, and also of AUC values of the genetic risk models.
Predictive Ability of Genetic Risk Models
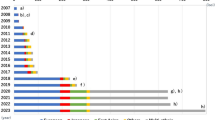
The AUC values of the genetic risk models ranged from 0.55 to 0.68, a range that was much smaller than that of the clinical models. Similar as for the clinical risk models and given that all SNPs approximately have the same low effect size, one would expect better predictive ability for models that included a higher number of SNPs, but Figure 1 shows that this was not observed for the studies listed in Table 2. The differences in the AUC values of the genetic risk scores cannot be explained by the number of polymorphisms included in the risk models. In fact, the highest genetic AUC (0.68) was found for a model that included 11 SNPs, and the lowest for a model that included these exact 11 SNPs plus an additional 8 others. The explanation for the absence of this relationship is likely in the low effect sizes of the genetic variants. A higher number of SNPs only yields a slightly higher AUC, a combined effect that could easily be outweighed by the influence of other factors, such as study design and study population.
Genetic risk prediction models have been investigated in prospective cohort studies, in case–control studies and in cross-sectional studies, and in study populations that differed in age, sex, and BMI (Table 2). These methodologic aspects may have impacted the observed AUC values in a similar way as they impact the AUC values of the clinical models. First, clinical and demographic characteristics of the study population may have influenced the observed predictive ability of the genetic risk models. There are two ways in which these characteristics may impact the predictive ability: the clinical and demographic characteristics of the study population itself and the differences in these characteristics between patients and nonpatients.
Table 2 describes mean age and BMI and the percentage of men in published genetic risk prediction studies for T2D. Mean age varied from 42.1 to 68.9 years, mean BMI from 23.4 to 29.1 kg/m2, and the percentage of men from 0% to 100%. It is often hypothesized that genetic risk factors may be more predictive in populations in which nongenetic T2D risk factors are not yet present (eg, in younger or normal weight cohorts), but AUC values of the genetic models were not markedly higher when populations were younger, had lower BMI, or had a lower percentage of men. However, because of the heterogeneity between the studies and their relatively small number, conclusions must be drawn with caution. Moreover, one study that had investigated the predictive performance in two age categories (<50 years vs ≥50 years) did find higher AUC values for the genetic risk score in younger people (AUC 0.66 vs 0.59) [40]. The observation that a stratified analysis within a single study did show differences in predictive ability suggests that the absence of a clear relation of age, BMI, and sex with AUC values across studies is likely explained by the presence of other differences between the studies.
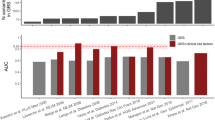
The other way in which clinical and demographic characteristics of the study population impact the predictive ability of risk models is through differences in these characteristics between patients and nonpatients. This specifically holds for characteristics that are included as risk factors in the prediction models, and for characteristics that are associated with these risk factors. Evidently and by definition, the presence of risk factors will differ between patients and nonpatients, but the difference can also be enlarged as a result of selection procedures. For example, patients who are recruited through hospitals may have more unfavorable risk profiles than patients randomly selected from the total patient population. Consequently, differences in risk factors between hospital-based cases and population-based controls will be larger and the impact of these risk factors on the predictive ability higher. For the studies listed in Table 2, differences in mean age ranged from −6.2 to 16.9 years, in mean BMI from 0.3 to 5.5 kg/m2, and differences in the percentage of men from −0.1% to 21.8%. Figure 2 shows that larger differences in mean age and BMI between patients and nonpatients were associated with higher AUC values for the clinical risk models, and, although less apparent, lower AUC values for the genetic models. No relation was observed between clinical AUC values and the percentage of men included in the studies, but this may be because male sex only marginally increases T2D risk compared with age and BMI (Table 1).
A second methodological aspect that may impact the predictive ability of risk models is study design. Genetic risk prediction studies are preferably conducted in prospective follow-up studies, but cross-sectional and case–control studies have been used as well (Table 2). The impact of study design on AUC values of T2D risk prediction models is in part related to the impact of population characteristics. Selection procedures for cases and controls may affect differences in clinical and demographic characteristics between patients and nonpatients. Case–control studies may demonstrate AUC values that deviate from those observed in prospective cohort and cross-sectional studies when cases and controls are recruited from different sources.
Another way in which study design may impact the predictive ability of risk models is length of follow-up in prospective cohort studies. Longer follow-up increases the likelihood that clinical T2D risk factors change over time, and that as a result their baseline values will be less predictive for the development of disease, resulting in prediction models with lower AUC. The length of follow-up of the studies listed in Table 2 varied from 6 to 25 years. Again, the number of prospective cohort studies was too small to investigate the impact of follow-up duration, but one study investigated the predictive ability in quintiles of follow-up time. This study demonstrated that the AUC of the clinical risk model decreased with increasing duration of follow-up, whereas the AUC of the genetic risk model increased [29]. From the first to the fifth quintile, the clinical AUC value decreased from 0.75 to 0.67 and the genetic AUC value increased from 0.57 to 0.62 [29].
Conclusions
In this review, we showed that study design and population characteristics may have affected the observed predictive performance of risk models. AUC values of the clinical risk models were higher and, although weaker, AUC values of the genetic risk models were lower when there were larger differences in age and BMI between cases and controls. This observation has important implications for the design and health care relevance of genetic risk prediction studies.
The predictive ability of risk models is preferably investigated in prospective cohort studies, but in practice often only case–control or cross-sectional designs are available. Because clinical risk factors, particularly the difference in risk factors between cases and controls, impact AUC values, it is expected that AUC values for genetic risk models obtained in case–control or cross-sectional studies may be valid when the distribution of these risk factors does not differ from prospective studies. For case–control studies, this means that the selection of cases and controls is not affected by these risk factors. In case of selection, transparency about the methods is important to enable a correct interpretation of the scientific and health care relevance of the results. For this reason, the GRIPS (Genetic Risk Prediction Studies) statement, a recently published guideline for the reporting of genetic risk prediction studies, recommends to describe eligibility criteria for participants, and sources and methods of selection of participants [44•].
The observed impact of population characteristics implies that it is important to assess the predictive ability of risk scores in representative samples of the population in which the model is ultimately applied to get valid estimates of their performance in that population. The question then is: which populations do we want to target for the prevention of T2D? Evidently, these may include individuals with metabolic syndrome or overweight, but for genetic prediction this may particularly concern young individuals who have not developed clinical risk factors. To date none of the T2D risk prediction studies have been conducted in younger populations; all studies were conducted in populations who on average were older than 40 years of age, two even in populations over 60 years of age [33, 41]. The study that best approximates the desired study population has been conducted in a population with a mean age of 42 years, a mean BMI of 25.6 kg/m2, and an almost equal number of men and women [35]. Given the observed differences in AUC values, we must conclude that we do not know whether genetic variants are useful in predicting T2D risk in younger populations. None of the studies so far has started from a health care perspective when investigating the predictive ability of T2D risk models.
There is increasing interest in investigating the value of genetic risk factors in the prediction of T2D risk. In this review, we demonstrated that the choice of study design and predominantly the choice of study population impact the observed predictive ability of risk models. For this reason it is important that the planning of future genetic risk prediction studies in T2D starts from a health care perspective by asking in which population we want to predict T2D risk. It is the answer to this question that determines the population in which the predictive ability should be assessed and that determines whether the results of the study ultimately can be informative and change health care practice.
References
Papers of particular interest, published recently, have been highlighted as: • Of importance
U.S. Census Bureau: General demographic characteristics, 2009 population estimates, United States. Available at http://factfinder.census.gov/servlet/QTTable?-ds_name=PEP_2009_EST&-qr_name=PEP_2009_EST_DP1&-geo_id=01000US. Accessed June 2011.
Centers for Disease Control and Prevention: Percentage of civilian, noninstitutionalized population with diagnosed diabetes, by age, United States, 1980–2009. Available at http://www.cdc.gov/diabetes/statistics/prev/national/figbyage.htm. Accessed June 2011.
Centers for Disease Control and Prevention: Age-adjusted percentage of civilian noninstitutionalized population with diagnosed diabetes, by sex, United States, 1980–2009. Available at http://www.cdc.gov/diabetes/statstics/prev/national/figbysex.htm. Accessed June 2011.
Centers for Disease Control and Prevention: Prevalence of overweight, obesity, and extreme obesity among adults: United States, trends 1976–1980 through 2007–2008. Available at http://www.cdc.gov/nchs/data/hestat/obesity_adult_07_08/obesity_adult_07_08.pdf. Accessed June 2011.
Flegal KM, Carroll MD, Ogden CL, et al. Prevalence and trends in obesity among US adults, 1999–2008. JAMA. 2010;303:235–41.
Nguyen NT, Nguyen XT, Lane J, et al. Relationship between obesity and diabetes in a US adult population: findings from the National Health and Nutrition Examinaton Survey, 1999–2006. Obes Surg. 2011;21:351–5.
Karve A, Hayward RA. Prevalence, diagnosis, and treatment of impaired fasting glucose and impaired glucose tolerance in nondiabetic U.S. adults. Diabetes Care. 2010;33:2355–9.
Gerstein HC, Santaguida P, Raina P, et al. Annual incidence and relative risk of diabetes in people with various categories of dysglycemia: a systematic overview and meta-analysis of prospective studies. Diabetes Res Clin Pract. 2007;78:305–12.
Selvin E, Steffes MW, Zhu H, et al. Glycated hemoglobin, diabetes, and cardiovascular risk in nondiabetic adults. N Engl J Med. 2010;362:800–11.
Alexander CM, Landsman PB, Teutsch SM, et al. NCEP-defined metabolic syndrome, diabetes, and prevalence of coronary heart disease among NHANES III participants age 50 years and older. Diabetes. 2003;52:1210–4.
Gillies CL, Abrams KR, Lambert PC, et al. Pharmacological and lifestyle interventions to prevent or delay type 2 diabetes in people with impaired glucose tolerance: systematic review and meta-analysis. BMJ. 2007;334:299.
Colagiuri S. Epidemiology of prediabetes. Med Clin North Am. 2011;95:299–307.
McCarthy MI. From hype to hope? A journey through the genetics of type 2 diabetes. Diabet Med. 2011;28:132–40.
McCarthy MI. Genomics, type 2 diabetes, and obesity. N Engl J Med. 2010;363:2339–50.
International Diabetes Federation: Diabetes and impaired glucose tolerance. Global burden: prevalence and projections, 2010 and 2030. Available at: http://www.idf.org/diabetesatlas/diabetes-and-impaired-glucose-tolerance. Accessed June 2011.
Stumvoll M, Goldstein BJ, van Haeften TW. Type 2 diabetes: principles of pathogenesis and therapy. Lancet. 2005;365:1333–46.
Paulweber B, Valensi P, Lindström J, et al. A European evidence-based guideline for the prevention of type 2 diabetes. Horm Metab Res. 2010;42 Suppl 1:S3–36.
Saaristo T, Peltonen M, Lindström J, et al. Cross-sectional evaluation of the Finnish Diabetes Risk Score: a tool to identify undetected type 2 diabetes, abnormal glucose tolerance and metabolic syndrome. Diab Vasc Dis Res. 2005;2:67–72.
Heikes KE, Eddy DM, Arondekar B, et al. Diabetes Risk Calculator: a simple tool for detecting undiagnosed diabetes and prediabetes. Diabetes Care. 2008;31:1040–5.
Janssens ACJW, Khoury MJ. Assessment of improved prediction beyond traditional risk factors – when does a difference make a difference? Circ Cardiovasc Genet. 2010;3:3–5.
Wilson PWF, Meigs JB, Sullivan L, et al. Prediction of incident diabetes mellitus in middle-aged adults. Arch Intern Med. 2007;167:1068–74.
• Hivert MF, Jablonski KA, Perreault L, et al. Updated genetic score based on 34 confirmed type 2 diabetes loci is associated with diabetes incidence and regression to normoglycemia in the Diabetes Prevention Program. Diabetes 2011;60:1340–8. This study investigated effectiveness of preventive interventions in individuals in all quartiles of genetic risk estimated by a genetic risk score. This is very interesting from a public health perspective.
Balkau B, Lange C, Fezeu L, et al. Predicting diabetes: clinical, biological, and genetic approaches: data from the Epidemiological Study on the Insulin Resistance Syndrome (DESIR). Diabetes Care. 2008;31:2056–61.
• Lyssenko V, Almgren P, Anevski D, et al. Genetic prediction of future type 2 diabetes. PLoS Med. 2005;2:e345. This was the first paper in T2D that considered the combined effect of multiple susceptibility variants in the prediction of disease risk.
Janssens ACJW, Gwinn M, Subramonia-Iyer S, et al. Does genetic testing really improve the prediction of future type 2 diabetes? PLoS Med. 2006;3:e114.
Weedon MN, McCarthy MI, Hitman G, et al. Combining information from common type 2 diabetes risk polymorphisms improves disease prediction. PLoS Med. 2006;3:e374.
Vaxillaire M, Veslot J, Dina C, et al. Impact of common type 2 diabetes risk polymorphisms in the DESIR prospective study. Diabetes. 2008;57:244–54.
Cornelis MC, Qi L, Zhang C, et al. Joint effects of common genetic variants on the risk for type 2 diabetes in U.S. men and women of European ancestry. Ann Intern Med. 2009;150:541–50.
Lyssenko V, Jonsson A, Almgren P, et al. Clinical risk factors, DNA variants, and the development of type 2 diabetes. N Engl J Med. 2008;359:2220–32.
Cauchi S, Meyre D, Durand E, et al. Post genome-wide association studies of novel genes associated with type 2 diabetes show gene-gene interaction and high predictive value. PLoS One. 2008;3:e2031.
Lin X, Song K, Lim N, et al. Risk prediction of prevalent diabetes in a Swiss population using a weighted genetic score–the CoLaus Study. Diabetologia. 2009;52:600–8.
Fontaine-Bisson B, Renström F, Rolandsson O, et al. Evaluating the discriminative power of multi-trait genetic risk scores for type 2 diabetes in a northern Swedish population. Diabetologia. 2010;53:2155–62.
van Hoek M, Dehghan A, Witteman JCM, et al. Predicting type 2 diabetes based on polymorphisms from genome-wide association studies: a population-based study. Diabetes. 2008;57:3122–8.
Lango H, Consortium UKTDG, Palmer CNA, et al. Assessing the combined impact of 18 common genetic variants of modest effect sizes on type 2 diabetes risk. Diabetes. 2008;57:3129–35.
Meigs JB, Shrader P, Sullivan LM, et al. Genotype score in addition to common risk factors for prediction of type 2 diabetes. N Engl J Med. 2008;359:2208–19.
Sparso T, Grarup N, Andreasen C, et al. Combined analysis of 19 common validated type 2 diabetes susceptibility gene variants shows moderate discriminative value and no evidence of gene-gene interaction. Diabetologia. 2009;52:1308–14.
Wang J, Stancakova A, Kuusisto J, et al. Identification of undiagnosed type 2 diabetic individuals by the finnish diabetes risk score and biochemical and genetic markers: a population-based study of 7232 Finnish men. J Clin Endocrinol Metab. 2010;95:3858–62.
Schulze MB, Weikert C, Pischon T, et al. Use of multiple metabolic and genetic markers to improve the prediction of type 2 diabetes: the EPIC-Potsdam Study. Diabetes Care. 2009;32:2116–9.
• Talmud PJ, Hingorani AD, Cooper JA, et al. Utility of genetic and non-genetic risk factors in prediction of type 2 diabetes: Whitehall II prospective cohort study. BMJ 2010;340:b4838. This study is a very good example of a genetic risk prediction study, with a very complete description of the study population and a thorough assessment of the predictive ability.
de Miguel-Yanes JM, Shrader P, Pencina MJ, et al. Genetic risk reclassification for type 2 diabetes by age below or above 50 years using 40 type 2 diabetes risk single nucleotide polymorphisms. Diabetes Care. 2011;34:121–5.
Miyake K, Yang W, Hara K, et al. Construction of a prediction model for type 2 diabetes mellitus in the Japanese population based on 11 genes with strong evidence of the association. J Hum Genet. 2009;54:236–41.
Hu C, Zhang R, Wang C, et al. PPARG, KCNJ11, CDKAL1, CDKN2A-CDKN2B, IDE-KIF11-HHEX, IGF2BP2 and SLC30A8 are associated with type 2 diabetes in a Chinese population. PLoS One. 2009;4:e7643.
• Mihaescu R, Meigs J, Sijbrands E, et al. Genetic risk profiling for type 2 diabetes. PloS Curr. 2011;3:RRN1208. This paper is regularly updated and gives an overview of all genetic risk prediction studies conducted in T2D.
• Janssens AC, Ioannidis JP, van Duijn CM, et al. Strengthening the reporting of genetic risk prediction studies: the GRIPS statement. Eur J Epidemiol. 2011;26:255–9. This paper describes reporting guidelines to enhance the transparency, quality, and completeness of the reporting of genetic risk prediction studies. A longer paper with examples and explanations was published simultaneously.
Disclosure
No potential conflicts of interest relevant to this article were reported.
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Willems, S.M., Mihaescu, R., Sijbrands, E.J.G. et al. A Methodological Perspective on Genetic Risk Prediction Studies in Type 2 Diabetes: Recommendations for Future Research. Curr Diab Rep 11, 511–518 (2011). https://doi.org/10.1007/s11892-011-0235-6
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11892-011-0235-6