Abstract
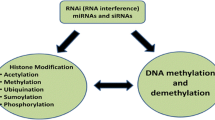
Plants are constantly exposed to hostile environments, which drive them to adopt several survival mechanisms including epigenetic modifications. Epigenetic modifications such as DNA methylation, histone modifications, and small RNA mediated silencing play significant roles in stress tolerance. Epigenetic modifications offer plants an advantage of long-term stress adaptation through either prolonged gene regulation or transgenerational inheritance. Among the epigenetic modifications, DNA methylation is a major epigenetic mark that stably inherit to multiple generations. Transgenerational epigenetic aspect of biotic stresses in plants is less explored compared to abiotic stresses. This review emphases on biotic-stress associated epigenetic changes and their inheritance in plants.

Similar content being viewed by others
References
Agorio A, Vera P (2007) ARGONAUTE4 is required for resistance to Pseudomonas syringae in Arabidopsis. Plant Cell 19:3778–3790. https://doi.org/10.1105/tpc.107.054494
Akimoto K, Katakami H, Kim HJ, Ogawa E, Sano CM, Wada Y, Sano H (2007) Epigenetic inheritance in rice plants. Ann Bot 100:205–217. https://doi.org/10.1093/aob/mcm110
Alonso C, Ramos-Cruz D, Becker C (2019) The role of plant epigenetics in biotic interactions. New Phytol 221:731–737. https://doi.org/10.1111/nph.15408
Amritha MS, Sridharan K, Puthur JT, Dhankher OP (2021) Priming with nanoscale materials for boosting abiotic stress tolerance in crop plants. J Agric Food Chem 69:10017–10035. https://doi.org/10.1021/acs.jafc.1c03673
Ando S, Jaskiewicz M, Mochizuki S, Koseki S, Miyashita S, Takahashi H, Conrath U (2021) Priming for enhanced ARGONAUTE2 activation accompanies induced resistance to cucumber mosaic virus in Arabidopsis thaliana. Mol Plant Pathol 22:19–30. https://doi.org/10.1111/mpp.13005
Arteaga-Vazquez MA, Chandler VL (2010) Paramutation in maize: RNA mediated trans-generational gene silencing. Curr Opin Genet Dev 20:156–163. https://doi.org/10.1016/j.gde.2010.01.008
Atighi MR, Verstraeten B, De Meyer T, Kyndt T (2021) Genome-wide shifts in histone modifications at early stage of rice infection with Meloidogyne graminicola. Mol Plant Pathol 22:440–455. https://doi.org/10.1101/2020.07.06.190538
Atkins CA, Smith PM, Rodriguez-Medina C (2011) Macromolecules in phloem exudates—a review. Protoplasma 248:165–172. https://doi.org/10.1007/s00709-010-0236-3
Baek D, Jiang J, Chung JS, Wang B, Chen J, Xin Z, Shi H (2010) Regulated AtHKT1 gene expression by a distal enhancer element and DNA methylation in the promoter plays an important role in salt tolerance. Plant Cell Physiol 52:149–161. https://doi.org/10.1093/pcp/pcq182
Baldrich P, Kakar K, Siré C, Moreno AB, Berger A, García-Chapa M, López-Moya JJ, Riechmann JL, San Segundo B (2014) Small RNA profiling reveals regulation of Arabidopsis miR168 and heterochromatic siRNA415 in response to fungal elicitors. BMC Genom 15:1–16. https://doi.org/10.1186/1471-2164-15-1083
Bartels A, Han Q, Nair P, Stacey L, Gaynier H, Mosley M, Huang Q, Pearson J, Hsieh TF, An YQ, Xiao W (2018) Dynamic DNA methylation in plant growth and development. Int J Mol Sci 19:2144–2161. https://doi.org/10.3389/fpls.2021.596236
Baulcombe D (2004) RNA silencing in plants. Nature 431:356–363. https://doi.org/10.1038/nature02874
Berger SL (2007) The complex language of chromatin regulation during transcription. Nature 447:407–412. https://doi.org/10.1038/nature05915
Berr A, McCallum EJ, Alioua A, Heintz D, Heitz T, Shen WH (2010) Arabidopsis histone methyltransferase SET DOMAIN GROUP8 mediates induction of the jasmonate/ethylene pathway genes in plant defense response to necrotrophic fungi. Plant Physiol 154:1403–1414. https://doi.org/10.1104/pp.110.161497
Bond DM, Baulcombe DC (2014) Small RNAs and heritable epigenetic variation in plants. Trends Cell Biol 24:100–107. https://doi.org/10.1016/j.tcb.2013.08.001
Bossdorf O, Richards CL, Pigliucci M (2008) Epigenetics for ecologists. Ecol Lett 11:106–115. https://doi.org/10.1111/j.1461-0248.2007.01130.x
Boyko A, Kathiria P, Zemp FJ, Yao Y, Pogribny I, Kovalchuk I (2007) Transgenerational changes in the genome stability and methylation in pathogen-infected plants: (Virus-induced plant genome instability). Nucleic Acids Res 35:1714–1725. https://doi.org/10.1093/nar/gkm029
Bruce TJA, Matthes MC, Napier JA, Pickett JA (2007) Stressful “memories” of plants: evidence and possible mechanisms. Plant Sci 173:603–608. https://doi.org/10.1016/j.plantsci.2007.09.002
Buchmann RC, Asad S, Wolf JN, Mohannath G, Bisaro DM (2009) Geminivirus AL2 and L2 proteins suppress transcriptional gene silencing and cause genome-wide reductions in cytosine methylation. J Virol 83:5005–5013. https://doi.org/10.1128/jvi.01771-08
Calarco JP, Borges F, Donoghue MT, Van Ex F, Jullien PE, Lopes T, Gardner R, Berger F, Feijó JA, Becker JD, Martienssen RA (2012) Reprogramming of DNA methylation in pollen guides epigenetic inheritance via small RNA. Cell 151:194–205. https://doi.org/10.1016/j.cell.2012.09.001
Cao Y, Dai Y, Cui S, Ma L (2008) Histone H2B monoubiquitination in the chromatin of FLOWERING LOCUS C regulates flowering time in Arabidopsis. Plant Cell 20:2586–2602. https://doi.org/10.1105/tpc.108.062760
Cedar H, Bergman Y (2009) Linking DNA methylation and histone modification: patterns and paradigms. Nat Rev Genet 10:295–304. https://doi.org/10.1038/nrg2540
Çelik Ö, Meriç S, Ayan A, Atak Ç (2020) Biotic stress-tolerant plants through small RNA technology. In: Guleria P, Kumar V (eds) Plant small RNA, 1st edn. Academic Press, pp 435–468. https://doi.org/10.1016/b978-0-12-817112-7.00020-1
Chen X (2009) Small RNAs and their roles in plant development. Annu Rev Cell Dev 25:21–44. https://doi.org/10.1146/annurev.cellbio.042308.113417
Cokus SJ, Feng S, Zhang X, Chen Z, Merriman B, Haudenschild CD, Pradhan S, Nelson SF, Pellegrini M, Jacobsen SE (2008) Shotgun bisulphite sequencing of the Arabidopsis genome reveals DNA methylation patterning. Nature 452:215–219. https://doi.org/10.1038/nature06745
Conrath U, Beckers GJ, Flors V, García-Agustín P, Jakab G, Mauch F, Newman MA, Pieterse CM, Poinssot B, Pozo MJ, Pugin A (2006) Priming: getting ready for battle. Mol Plant Microbe Interact 19:1062–1071. https://doi.org/10.1094/mpmi-19-1062
Conrath U, Pieterse CM, Mauch-Mani B (2002) Priming in plant–pathogen interactions. Trends Plant Sci 7:210–216. https://doi.org/10.1016/S1360-1385(02)02244-6
Crespo-Salvador Ó, Escamilla-Aguilar M, López-Cruz J, López-Rodas G, González-Bosch C (2018) Determination of histone epigenetic marks in Arabidopsis and tomato genes in the early response to Botrytis cinerea. Plant Cell Rep 37:153–166. https://doi.org/10.1007/s00299-017-2218-9
Dalakouras A, Dadami E, Zwiebel M, Krczal G, Wassenegger M (2012) Transgenerational maintenance of transgene body CG but not CHG and CHH methylation. Epigenetics 7:1071–1078. https://doi.org/10.4161/epi.21644
De-La-Peña C, Rangel-Cano A, Alvarez-Venegas R (2012) Regulation of disease-responsive genes mediated by epigenetic factors: interaction of Arabidopsis–Pseudomonas. Mol Plant Pathol 13:388–398. https://doi.org/10.1111/j.1364-3703.2011.00757.x
Dhawan R, Luo H, Foerster AM, AbuQamar S, Du HN, Briggs SD, Scheid OM, Mengiste T (2009) HISTONE MONOUBIQUITINATION1 interacts with a subunit of the mediator complex and regulates defense against necrotrophic fungal pathogens in Arabidopsis. Plant Cell 21:1000–1019. https://doi.org/10.1105/tpc.108.062364
Dowen RH, Pelizzola M, Schmitz RJ, Lister R, Dowen JM, Nery JR, Dixon JE, Ecker JR (2012) Widespread dynamic DNA methylation in response to biotic stress. Proc Natl Acad Sci 109:E2183–E2191. https://doi.org/10.1073/pnas.1209329109
Dunoyer P, Himber C, Voinnet O (2006) Induction, suppression and requirement of RNA silencing pathways in virulent Agrobacterium tumefaciens infections. Nat Genet 38:258–263. https://doi.org/10.1038/ng1722
Eichten SR, Schmitz RJ, Springer NM (2014) Epigenetics: beyond chromatin modifications and complex genetic regulation. Plant Physiol 65:933–947. https://doi.org/10.1104/pp.113.234211
Erdmann RM, Picard CL (2020) RNA-directed DNA methylation. PLoS Genet 16:e1009034. https://doi.org/10.1371/journal.pgen.1009034
Galis I, Gaquerel E, Pandey SP, Baldwin IT (2009) Molecular mechanisms underlying plant memory in JA-mediated defence responses. Plant Cell Environ 32:617–627. https://doi.org/10.1111/j.1365-3040.2008.01862.x
Gallego-Bartolomé J (2020) DNA methylation in plants: mechanisms and tools for targeted manipulation. New Phytol 227:38–44. https://doi.org/10.1111/nph.16529
Genre A, Bonfante P (2002) Epidermal cells of a symbiosis-defective mutant of Lotus japonicus show altered cytoskeleton organisation in the presence of a mycorrhizal fungus. Protoplasma 219:43–50. https://doi.org/10.1007/s007090200004
Gohlke J, Deeken R (2014) Plant responses to Agrobacterium tumefaciens and crown gall development. Front Plant Sci 5:155–166. https://doi.org/10.3389/fpls.2014.00155
Guseinov VA, Vanyushin BF (1975) Content and localisation of 5-methylcytosine in DNA of healthy and wilt-infected cotton plants. Biochim Biophys Acta 395:229–238. https://doi.org/10.1016/0005-2787(75)90193-8
Hanania U, Furman-Matarasso N, Ron M, Avni A (1999) Isolation of a novel SUMO protein from tomato that suppresses EIX‐induced cell death. Plant J 19:533–541. https://doi.org/10.1046/j.1365-313x.1999.00547.x
Herceg Z, Murr R (2011) Mechanisms of histone modifications. In: Tollefsbol T (ed) Handbook of epigenetics: the new molecular and medical genetics, 1st edn. Academic Press, pp 25–45. https://doi.org/10.1016/b978-0-12-375709-8.00003-4
Hewezi T, Lane T, Piya S, Rambani A, Rice JH, Staton M (2017) Cyst nematode parasitism induces dynamic changes in the root epigenome. Plant Physiol 174:405–420. https://doi.org/10.1104/pp.16.01948
Holeski LM, Jander G, Agrawal AA (2012) Transgenerational defense induction and epigenetic inheritance in plants. Trends Ecol Evol 27:618–626. https://doi.org/10.1016/j.tree.2012.07.011
Hu M, Pei BL, Zhang LF, Li YZ (2014) Histone H2B monoubiquitination is involved in regulating the dynamics of microtubules during the defense response to Verticillium dahliae toxins in Arabidopsis. Plant Physiol 164:1857–1865. https://doi.org/10.1104/pp.113.234567
Jagadeeswaran G, Saini A, Sunkar R (2009) Biotic and abiotic stress down-regulate miR398 expression in Arabidopsis. Planta 229:1009–1014. https://doi.org/10.1007/s00425-009-0889-3
Jaskiewicz M, Conrath U, Peterhänsel C (2011) Chromatin modification acts as a memory for systemic acquired resistance in the plant stress response. EMBO Rep 12:50–55. https://doi.org/10.3410/f.720681943.793516343
Joseph TJ, Chandhini S, Das S, Mysore SK, Shah MJ (2021) Methylation status of Arabidopsis DNA repair gene promoters during Agrobacterium infection reveals epigenetic changes in three generations. Plant Mol Biol Rep 39:773–791. https://doi.org/10.1007/s11105-021-01287-6
Kasschau KD, Xie Z, Allen E, Llave C, Chapman EJ, Krizan KA, Carrington JC (2003) P1/HC-Pro, a viral suppressor of RNA silencing, interferes with Arabidopsis development and miRNA function. Dev Cell 4:205–217. https://doi.org/10.1016/s1534-5807(03)00025-x
Katiyar-Agarwal S, Gao S, Vivian-Smith A, Jin H (2007) A novel class of bacteria-induced small RNAs in Arabidopsis. Genes Dev 21:3123–3134. https://doi.org/10.1101/gad.1595107
Katiyar-Agarwal S, Morgan R, Dahlbeck D, Borsani O, Villegas A, Zhu JK, Staskawicz BJ, Jin H (2006) A pathogen-inducible endogenous siRNA in plant immunity. Proc Natl Acad Sci 103:18002–18007. https://doi.org/10.1073/pnas.0608258103
Kellenberger RT, Schlüter PM, Schiestl FP (2016) Herbivore-induced DNA demethylation changes floral signalling and attractiveness to pollinators in Brassica rapa. PLoS ONE 11:e0166646. https://doi.org/10.1371/journal.pone.0166646b
Khraiwesh B, Zhu JK, Zhu J (2012) Role of miRNAs and siRNAs in biotic and abiotic stress responses of plants. Biochim Biophys Acta 1819:137–148. https://doi.org/10.1016/j.bbagrm.2011.05.001
Kim S, Piquerez SJ, Ramirez-Prado JS, Mastorakis E, Veluchamy A, Latrasse D, Manza-Mianza D, Brik-Chaouche R, Huang Y, Rodriguez-Granados NY, Concia L (2020) GCN5 modulates salicylic acid homeostasis by regulating H3K14ac levels at the 5′ and 3′ ends of its target genes. Nucleic Acids Res 48:5953–5966. https://doi.org/10.1093/nar/gkaa369
Kinoshita T, Seki M (2014) Epigenetic memory for stress response and adaptation in plants. Plant Cell Physiol 55:1859–1863. https://doi.org/10.1093/pcp/pcu125
Kuć J (1982) Induced immunity to plant disease. Bioscience 32:854–860. https://doi.org/10.2307/1309008
Kumar V, Thakur JK, Prasad M (2021) Histone acetylation dynamics regulating plant development and stress responses. Cell Mol Life Sci 78:4467–4486. https://doi.org/10.1007/s00018-021-03794-x
Lal SK, Kumar S, Sheri V, Mehta S, Varakumar P, Ram B, Borphukan B, James D, Fartyal D, Reddy MK (2018) Seed priming: an emerging technology to impart abiotic stress tolerance in crop plants. In: Rakshit A, Singh HB (eds) Advances in seed priming, 1st edn. Springer, Singapore, pp 41–50. https://doi.org/10.1007/978-981-13-0032-5
Lämke J, Bäurle I (2017) Epigenetic and chromatin-based mechanisms in environmental stress adaptation and stress memory in plants. Genome Biol 18:124–135. https://doi.org/10.1186/s13059-017-1263-6
Le TN, Schumann U, Smith NA, Tiwari S, Au PCK, Zhu QH, Taylor JM, Kazan K, Llewellyn DJ, Zhang R, Dennis ES (2014) DNA demethylases target promoter transposable elements to positively regulate stress responsive genes in Arabidopsis. Genome Biol 15:458–476. https://doi.org/10.1186/s13059-014-0458-3
Li S, Lin M, Wang J, Zhang L, Lin M, Hu Z, Qi Z, Jiang H, Fu Y, Xin D, Liu C (2019) Regulation of soybean SUMOylation system in response to Phytophthora sojae infection and heat shock. Plant Growth Regul 87:69–82. https://doi.org/10.1007/s10725-018-0452-y
Li Y, Xia Q, Kou H, Wang D, Lin X, Wu Y, Xu C, Xing S, Liu B (2011) Induced Pib expression and resistance to Magnaporthe grisea are compromised by cytosine demethylation at critical promoter regions in rice. J Integr Plant Biol 53:814–823. https://doi.org/10.1111/j.1744-7909.2011.01070.x
López A, Ramírez V, García-Andrade J, Flors V, Vera P (2011) The RNA silencing enzyme RNA polymerase V is required for plant immunity. PLoS Genet 7:e1002434. https://doi.org/10.1371/journal.pgen.1002434
López-Galiano MJ, González-Hernández AI, Crespo-Salvador O, Rausell C, Real MD, Escamilla M, Camañes G, García-Agustín P, González-Bosch C, García-Robles I (2018) Epigenetic regulation of the expression of WRKY75 transcription factor in response to biotic and abiotic stresses in Solanaceae plants. Plant Cell Rep 37:167–176. https://doi.org/10.1007/s00299-017-2219-8
Luan Y, Cui J, Li J, Jiang N, Liu P, Meng J (2018) Effective enhancement of resistance to Phytophthora infestans by overexpression of miR172a and b in Solanum lycopersicum. Planta 247:127–138. https://doi.org/10.1007/s00425-017-2773-x
Luna E, Bruce TJ, Roberts MR, Flors V, Ton J (2012) Next-generation systemic acquired resistance. Plant Physiol 158:844–853. https://doi.org/10.1104/pp.111.187468
Luo JY, Pan XL, Peng TC, Chen YY, Zhao H, Mu L, Peng Y, He R, Tang H (2016) DNA methylation patterns of banana leaves in response to Fusarium oxysporum f. sp. cubense tropical race 4. J Integr Agric 15:2736–2744. https://doi.org/10.1016/s2095-3119(16)61495-8
Martin A, Troadec C, Boualem A, Rajab M, Fernandez R, Morin H, Pitrat M, Dogimont C, Bendahmane A (2009) A transposon-induced epigenetic change leads to sex determination in melon. Nature 461:1135–1138. https://doi.org/10.1038/nature08498
Mason G, Noris E, Lanteri S, Acquadro A, Accotto GP, Portis E (2008) Potentiality of methylation-sensitive amplification polymorphism (MSAP) in identifying genes involved in tomato response to tomato yellow leaf curl sardinia virus. Plant Mol Biol Rep 26:156–173. https://doi.org/10.1007/s11105-008-0031-x
Mathieu O, Reinders J, Čaikovski M, Smathajitt C, Paszkowski J (2007) Transgenerational stability of the Arabidopsis epigenome is coordinated by CG methylation. Cell 130:851–862. https://doi.org/10.1016/j.cell.2007.07.007
Mauch-Mani B, Baccelli I, Luna E, Flors V (2017) Defense priming: an adaptive part of induced resistance. Annu Rev Plant Biol 68:485–512. https://doi.org/10.1146/annurev-arplant-042916-041132
Miryeganeh M (2021) Plants’ epigenetic mechanisms and abiotic stress. Genes 12:1106–1123. https://doi.org/10.3390/genes12081106
Mohammadi P, Bahramnejad B, Badakhshan H, Kanouni H (2015) DNA methylation changes in fusarium wilt resistant and sensitive chickpea genotypes (Cicer arietinum L.). Physiol Mol Plant Pathol 91:72–80. https://doi.org/10.1016/j.pmpp.2015.06.001
Morán-Diez ME, Martinez de Alba AE, Rubio MB, Hermosa R, Monte E (2021) Trichoderma and the plant heritable priming responses. J Fungi 7:318–341. https://doi.org/10.3390/jof7040318
Mosher RA, Durrant WE, Wang D, Song J, Dong X (2006) A comprehensive structure–function analysis of Arabidopsis SNI1 defines essential regions and transcriptional repressor activity. Plant Cell 18:1750–1765. https://doi.org/10.1105/tpc.105.039677
Mosher RA, Melnyk CW (2010) siRNAs and DNA methylation: seedy epigenetics. Trends Plant Sci 15:204–210. https://doi.org/10.1016/j.tplants.2010.01.002
Navarro L, Dunoyer P, Jay F, Arnold B, Dharmasiri N, Estelle M, Voinnet O, Jones JD (2006) A plant miRNA contributes to antibacterial resistance by repressing auxin signaling. Science 312:436–439. https://doi.org/10.1126/science.1126088
Navarro L, Jay F, Nomura K, He SY, Voinnet O (2008) Suppression of the microRNA pathway by bacterial effector proteins. Science 321:964–967. https://doi.org/10.1126/science.1159505
Nazim Uddin M, Kim JY (2013) Intercellular and systemic spread of RNA and RNAi in plants. Wiley Interdiscip Rev RNA 4:279–293. https://doi.org/10.1002/wrna.1160
Nonomura K-I (2018) Small RNA pathways responsible for non-cell-autonomous regulation of plant reproduction. Plant Reprod 31:21–29. https://doi.org/10.1007/s00497-018-0321-x
Orth K, Xu Z, Mudgett MB, Bao ZQ, Palmer LE, Bliska JB, Mangel WF, Staskawicz B, Dixon JE (2000) Disruption of signaling by Yersinia effector YopJ, a ubiquitin-like protein protease. Science 290:1594–1597. https://doi.org/10.1126/science.290.5496.1594
Palma K, Thorgrimsen S, Malinovsky FG, Fiil BK, Nielsen HB, Brodersen P, Hofius D, Petersen M, Mundy J (2010) Autoimmunity in Arabidopsis acd11 is mediated by epigenetic regulation of an immune receptor. PLoS Pathog 6:e1001137. https://doi.org/10.1371/journal.ppat.1001137
Park HJ, Kim WY, Park HC, Lee SY, Bohnert HJ, Yun DJ (2011) SUMO and SUMOylation in plants. Mol Cells 32:305–316. https://doi.org/10.1007/s10059-011-0122-7
Park JH, Shin C (2015) The role of plant small RNAs in NB-LRR regulation. Brief Funct Genomics 14:268–274. https://doi.org/10.1093/bfgp/elv006
Paszkowski J, Grossniklaus U (2011) Selected aspects of transgenerational epigenetic inheritance and resetting in plants. Curr Opin Plant Biol 14:195–203. https://doi.org/10.1016/j.pbi.2011.01.002
Pavet V, Quintero C, Cecchini NM, Rosa AL, Alvarez ME (2006) Arabidopsis displays centromeric DNA hypomethylation and cytological alterations of heterochromatin upon attack by Pseudomonas syringae. Mol Plant Microbe Interact 19:577–587. https://doi.org/10.1094/mpmi-19-0577
Peng H, Zhang J (2009) Plant genomic DNA methylation in response to stresses: potential applications and challenges in plant breeding. Prog Nat Sci 19:1037–1045. https://doi.org/10.1016/j.pnsc.2008.10.014
Pickart CM, Eddins MJ (2004) Ubiquitin: structures, functions, mechanisms. Biochim Biophys Acta 1695:55–72. https://doi.org/10.1016/j.bbamcr.2004.09.019
Pikaard CS, Scheid OM (2014) Epigenetic regulation in plants. Cold Spring Harb Perspect Biol 6:a019315. https://doi.org/10.1101/cshperspect.a019315
Rambani A, Rice JH, Liu J, Lane T, Ranjan P, Mazarei M, Pantalone V, Stewart CN, Staton M, Hewezi T (2015) The methylome of soybbean roots during the compatible interaction with the soybean cyst nematode. Plant Physiol 168:1364–1377. https://doi.org/10.1104/pp.15.00826
Ramirez-Prado JS, Abulfaraj AA, Rayapuram N, Benhamed M, Hirt H (2018) Plant immunity: from signaling to epigenetic control of defense. Trends Plant Sci 23:833–844. https://doi.org/10.1016/j.tplants.2018.06.004
Rasmann S, de Vos M, Casteel CL, Tian D, Halitschke R, Sun JY, Agrawal AA, Felton GW, Jander G (2012) Herbivory in the previous generation primes plants for enhanced insect resistance. Plant Physiol 158:854–863. https://doi.org/10.1104/pp.111.187831
Riddle NC (2014) Heritable generational epigenetic effects through RNA. In: Tollefsbol T (ed) Transgenerational epigenetics, 1st edn. Academic Press, pp 105–119. https://doi.org/10.1016/b978-0-12-405944-3.00010-6
Roberts MR, Sánchez AL (2019) Plant epigenetic mechanisms in response to biotic stress. In: Alvarez-Venegas R, De-la-Peña C, Casas-Mollano JA (eds) Epigenetics in plants of agronomic importance: fundamentals and applications, 2nd edn. Springer, Cham, pp 65–113. https://doi.org/10.1007/978-3-030-14760-0_2
Roden J, Eardley L, Hotson A, Cao Y, Mudgett MB (2004) Characterization of the Xanthomonas AvrXv4 effector, a SUMO protease translocated into plant cells. Mol Plant Microbe Interact 17:633–643. https://doi.org/10.1094/mpmi.2004.17.6.633
Sako K, Nguyen HM, Seki M (2020) Advances in chemical priming to enhance abiotic stress tolerance in plants. Plant Cell Physiol 61:1995–2003. https://doi.org/10.1093/pcp/pcaa119
Sánchez LA, Stassen JH, Furci L, Smith LM, Ton J (2016) The role of DNA (de) methylation in immune responsiveness of Arabidopsis. Plant J 88:361–374. https://doi.org/10.1111/tpj.13252
Shah JM (2021) Epimutations and mutations, nurturing phenotypic diversity. Genetica. https://doi.org/10.1007/s10709-021-00124-8
Shahbazian MD, Grunstein M (2007) Functions of site-specific histone acetylation and deacetylation. Annu Rev Biochem 76:75–100. https://doi.org/10.1146/annurev.biochem.76.052705.162114
Shiba H, Kakizaki T, Iwano M, Tarutani Y, Watanabe M, Isogai A, Takayama S (2006) Dominance relationships between self-incompatibility alleles controlled by DNA methylation. Nat Genet 38:297–299. https://doi.org/10.1038/ng1734
Sicilia A, Catara V, Scialò E, Lo Piero AR (2021) Fungal infection induces anthocyanin biosynthesis and changes in DNA methylation configuration of Blood Orange [Citrus sinensis L.(Osbeck)]. Plants 10:244–254. https://doi.org/10.3390/plants10020244
Slaughter A, Daniel X, Flors V, Luna E, Hohn B, Mauch-Mani B (2012) Descendants of primed Arabidopsis plants exhibit resistance to biotic stress. Plant Physiol 158:835–843. https://doi.org/10.1104/pp.111.191593
Soto-Suárez M, Baldrich P, Weigel D, Rubio-Somoza I, San Segundo B (2017) The Arabidopsis miR396 mediates pathogen-associated molecular pattern-triggered immune responses against fungal pathogens. Sci Rep 7:1–14. https://doi.org/10.1038/srep44898
Stassen JH, López A, Jain R, Pascual-Pardo D, Luna E, Smith LM, Ton J (2018) The relationship between transgenerational acquired resistance and global DNA methylation in Arabidopsis. Sci Rep 8:1–13. https://doi.org/10.1038/s41598-018-32448-5
Takahashi S, Fukushima N, Osabe K, Itabashi E, Shimizu M, Miyaji N, Takasaki-Yasuda T, Suzuki Y, Seki M, Fujimoto R (2018) Identification of DNA methylated regions by using methylated DNA immunoprecipitation sequencing in Brassica rapa. Crop Pasture Sci 69:107–120. https://doi.org/10.1071/cp17394
Thiebaut F, Hemerly AS, Ferreira PCG (2019) A role for epigenetic regulation in the adaptation and stress responses of non-model plants. Front Plant Sci 10:246–253. https://doi.org/10.3389/fpls.2019.00246
Thomas TTD, Puthur JT (2017) UV radiation priming: a means of amplifying the inherent potential for abiotic stress tolerance in crop plants. Environ Ex Bot 138:57–66. https://doi.org/10.1016/j.envexpbot.2017.03.003
Turgut-Kara N, Arikan B, Celik H (2020) Epigenetic memory and priming in plants. Genetica 148:47–54. https://doi.org/10.1007/s10709-020-00093-4
van den Burg HA, Kini RK, Schuurink RC, Takken FL (2010) Arabidopsis small ubiquitin-like modifier paralogs have distinct functions in development and defense. Plant Cell 22:1998–2016. https://doi.org/10.1105/tpc.109.070961
Wada Y, Miyamoto K, Kusano T, Sano H (2004) Association between up-regulation of stress-responsive genes and hypomethylation of genomic DNA in tobacco plants. Mol Genet Genomics 271:658–666. https://doi.org/10.1007/s00438-004-1018-4
Walbot V (1996) Sources and consequences of phenotypic and genotypic plasticity in flowering plants. Trends Plant Sci 1:27–32. https://doi.org/10.1016/s1360-1385(96)80020-3
Wang L, Chen H, Li J, Shu H, Zhang X, Wang Y, Tyler BM, Dong S (2020) Effector gene silencing mediated by histone methylation underpins host adaptation in an oomycete plant pathogen. Nucleic Acids Res 48:1790–1799. https://doi.org/10.1093/nar/gkz1160
Weake VM, Workman JL (2008) Histone ubiquitination: triggering gene activity. Mol Cell 29:653–663. https://doi.org/10.1016/j.molcel.2008.02.014
Xia S, Cheng YT, Huang S, Win J, Soards A, Jinn TL, Jones JD, Kamoun S, Chen S, Zhang Y, Li X (2013) Regulation of transcription of nucleotide-binding leucine-rich repeat-encoding genes SNC1 and RPP4 via H3K4 trimethylation. Plant Physiol 162:1694–1705. https://doi.org/10.1104/pp.113.214551
Yaish MW (2017) Epigenetic modifications associated with abiotic and biotic stresses in plants: an implication for understanding plant evolution. Front Plant Sci 8:1–3. https://doi.org/10.3389/fpls.2017.01983
Yakura H (2020) Cognitive and memory functions in plant immunity. Vaccines 8:541–556. https://doi.org/10.3390/vaccines8030541
Yang L, Mu X, Liu C, Cai J, Shi K, Zhu W, Yang Q (2015) Overexpression of potato miR482e enhanced plant sensitivity to Verticillium dahliae infection. J Integr Plant Biol 57:1078–1088. https://doi.org/10.1111/jipb.12348
Yang X, Zhang L, Yang Y, Schmid M, Wang Y (2021) miRNA mediated regulation and interaction between plants and pathogens. Int J Mol Sci 22:2913–2926. https://doi.org/10.3390/ijms22062913
Yi HS, Yang JW, Choi HK, Ghim SY, Ryu CM (2012) Benzothiadiazole-elicited defense priming and systemic acquired resistance against bacterial and viral pathogens of pepper under field conditions. Plant Biotechnol Rep 6:373–380. https://doi.org/10.1007/s11816-012-0234-3
Yu A, Lepère G, Jay F, Wang J, Bapaume L, Wang Y, Abraham AL, Penterman J, Fischer RL, Voinnet O, Navarro L (2013) Dynamics and biological relevance of DNA demethylation in Arabidopsis antibacterial defense. Proc Natl Acad Sci 110:2389–2394. https://doi.org/10.1073/pnas.1211757110
Zeng W, Huang H, Lin X, Zhu C, Kosami KI, Huang C, Zhang H, Duan CG, Zhu JK, Miki D (2021) Roles of DEMETER in regulating DNA methylation in vegetative tissues and pathogen resistance. J Integr Plant Biol 63:691–706. https://doi.org/10.1111/jipb.13037
Zhang X, Ménard R, Li Y, Coruzzi GM, Heitz T, Shen WH, Berr A (2020) Arabidopsis SDG8 potentiates the sustainable transcriptional induction of the pathogenesis-related genes PR1 and PR2 during plant defense response. Front Plant Sci 11:277–291. https://doi.org/10.3389/fpls.2020.00277
Zhao K, Kong D, Jin B, Smolke CD, Rhee SY (2021) A novel bivalent chromatin associates with rapid induction of camalexin biosynthesis genes in response to a pathogen signal in Arabidopsis. Elife 10:e69508. https://doi.org/10.7554/eLife.69508
Zhao T, Zhan Z, Jiang D (2019) Histone modifications and their regulatory roles in plant development and environmental memory. J Genet Genomics 46:467–476. https://doi.org/10.1016/j.jgg.2019.09.005
Zhou C, Zhang L, Duan J, Miki B, Wu K (2005) HISTONE DEACETYLASE19 is involved in jasmonic acid and ethylene signaling of pathogen response in Arabidopsis. Plant Cell 17:1196–1204. https://doi.org/10.1105/tpc.104.028514
Zhu QH, Shan WX, Ayliffe MA, Wang MB (2016) Epigenetic mechanisms: an emerging player in plant-microbe interactions. Mol Plant Microbe Interact 29:187–196. https://doi.org/10.1094/mpmi-08-15-0194-fi
Zou B, Yang DL, Shi Z, Dong H, Hua J (2014) Monoubiquitination of histone 2B at the disease resistance gene locus regulates its expression and impacts immune responses in Arabidopsis. Plant Physiol 165:309–318. https://doi.org/10.1104/pp.113.227801
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Authors declare no conflicts of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Joseph, J.T., Shah, J.M. Biotic stress-induced epigenetic changes and transgenerational memory in plants. Biologia 77, 2007–2021 (2022). https://doi.org/10.1007/s11756-022-01053-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11756-022-01053-3