Abstract
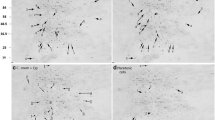
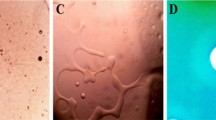
Lipopolysaccharide (LPS) is the main surface constituent of Gram-negative bacteria. Lipid A, the hydrophobic moiety, outer monolayer of the outer cell membrane forms the major component of LPS. Immunogenic Lipid A is recognized by the innate immune system through the TLR 4/MD-2 complex. Pseudomonas aeruginosa PAO1, a Gram-negative bacterium is known to cause nosocomial infection and known for its adaptation to adverse environmental conditions. Pseudomonas aeruginosa can infect a broad host spectrum including Caenorhabditis elegans, a simple free living soil nematode. Here, we reveal that PAO1 modifies its Lipid A during the host interaction with C. elegans. The penta-acylated form of Lipid A was identified by using matrix assisted laser desorption ionization–time of flight analysis and the β-(1,6)-linked disaccharide of glucosamine with phosphate groups, 2 and 2′ amide linked fatty acid chain and 3 and 3′ ester linked fatty acids were investigated for the modification using the non destructive 1H NMR, spin–lattice (T 1) relaxation measurement, differential scanning calorimetry. T 1 relaxation measurements showed that the 2 and 2′ amide linked fatty acid chain, –CH in the glucosamine disaccharide of PAO1 lipid A, in an exposed host had a different spin lattice relaxation time compared to an unexposed host and the findings were reconfirmed using in vitro human corneal epithelial cells cell lines. Furthermore, scanning electron microscope and confocal laser scanning microscopy analysis revealed that the P. aeruginosa PAO1 biofilm formation was disturbed in the exposed host condition. The daf-12, daf-16, tol-1, pmk-1, ins-7 and ilys3 immune genes of C. elegans were examined with live bacterial and isolated lipid moiety infection and the expression was found to be highly specific. Overall, the present study revealed that PAO1 modified its 2 and 2′ amide linked fatty acid chain in the lipid A of PAO1 LPS during the exposed host condition.














Similar content being viewed by others
Abbreviations
- °C:
-
Degree Celsius
- cDNA:
-
Complementary deoxyribonucleic acid
- CLSM:
-
Confocal laser scanning microscope
- CO2 :
-
Carbon-di-oxide
- CPA:
-
Common polysaccharide antigen
- daf :
-
Abnormal dauer formation (gene)
- DHB:
-
2,5-Dihydroxybenzoic acid
- DSC:
-
Differential scanning calorimetry
- FTIR:
-
Fourier transform infrared
- h:
-
Hour
- HCEC:
-
Human corneal epithelial cells
- HEGF:
-
Human epithelial growth factor
- ilys :
-
Invertebrate lysozyme (gene)
- ins :
-
Insulin (gene)
- KBr:
-
Potassium bromide
- L4:
-
Larva stage 4
- LPS:
-
Lipopolysaccharide
- Lys:
-
Lysozyme
- M:
-
Molar
- MALDI–TOF:
-
Matrix assisted laser desorption ionization–time of flight
- MAMP:
-
Microbe associated molecular pattern
- m:
-
minute
- mRNA:
-
Messenger ribonucleic acid
- NGM:
-
Nematode growth medium
- OD:
-
Optical density
- OSA:
-
O-specific antigen(s)
- PAMP:
-
Pathogen associated molecular pattern(s)
- PBS:
-
Phosphate buffer saline
- PCR:
-
Polymerase chain reaction
- pmk :
-
P38 map kinase (gene)
- qPCR:
-
Quantitative polymerase chain reaction
- RNA:
-
Ribonucleic acid
- rpm:
-
Rotation per minute
- s:
-
Second
- SDS-PAGE:
-
Sodium dodecyl sulfate-polyacrylamide gel electrophoresis
- SEM:
-
Scanning electron microscope
- tol :
-
Toll like receptor (gene)
- UV–VIS:
-
Ultra violet–visible
References
Raetz CR, Whitfield C (2002) Lipopolysaccharide endotoxins. Annu Rev Biochem 71:635–700
Gallo RL, Hooper LV (2012) Epithelial antimicrobial defence of the skin and intestine. Nat Rev Immunol 12(7):503–516
Lewis LA, Shafer WM, Ray TD, Ram S, Rice PA (2012) Phosphoethanolamine residues on the lipid A moiety of Neisseria gonorrhoeae lipooligosaccharide modulate binding of complement inhibitors and resistance to complement killing. Infect Immun 81(1):33–42
Giangrande C, Colarusso L, Lanzetta R, Molinaro A, Pucci P, Amoresano A (2012) Innate immunity probed by lipopolysaccharides affinity strategy and proteomics. Anal Bioanal Chem 405(2–3):775–784
Tan LA, Yang AC, Kishore U, Sim RB (2011) Interactions of complement proteins C1q and factor H with lipid A and Escherichia coli: further evidence that factor H regulates the classical complement pathway. Protein Cell 2(4):320–332
Needham BD, Trent MS (2013) Fortifying the barrier: the impact of lipid A remodelling on bacterial pathogenesis. Nat Rev Microbiol 11(7):467–481
Frimmersdorf E, Horatzek S, Pelnikevich A, Wiehlmann L, Schomburg D (2010) How Pseudomonas aeruginosa adapts to various environments: a metabolomic approach. Environ Microbiol 12(6):1734–1747
Cigana C, Lore NI, Bernardini ML, Bragonzi A (2011) Dampening host sensing and avoiding recognition in Pseudomonas aeruginosa pneumonia. J Biomed Biotechnol 2011:852513
Skiada A, Markogiannakis A, Plachouras D, Daikos GL (2011) Adaptive resistance to cationic compounds in Pseudomonas aeruginosa. Int J Antimicrob Agents 37(3):187–193
Carmeli Y, Troillet N, Eliopoulos GM, Samore MH (1999) Emergence of antibiotic-resistant Pseudomonas aeruginosa: comparison of risks associated with different antipseudomonal agents. Antimicrob Agents Chemother 43(6):1379–1382
Cullen TW, Giles DK, Wolf LN, Ecobichon C, Boneca IG, Trent MS (2011) Helicobacter pylori versus the host: remodeling of the bacterial outer membrane is required for survival in the gastric mucosa. PLoS Pathog 7(12):e1002454
Ciornei CD, Novikov A, Beloin C, Fitting C, Caroff M, Ghigo JM, Cavaillon JM, Adib-Conquy M (2010) Biofilm-forming Pseudomonas aeruginosa bacteria undergo lipopolysaccharide structural modifications and induce enhanced inflammatory cytokine response in human monocytes. Innate Immun 16(5):288–301
Roland KL, Martin LE, Esther CR, Spitznagel JK (1993) Spontaneous pmrA mutants of Salmonella typhimurium LT2 define a new two-component regulatory system with a possible role in virulence. J Bacteriol 175(13):4154–4164
Miller SI, Kukral AM, Mekalanos JJ (1989) A two-component regulatory system (phoP phoQ) controls Salmonella typhimurium virulence. Proc Natl Acad Sci USA 86(13):5054–5058
Prost LR, Miller SI (2008) The Salmonellae PhoQ sensor: mechanisms of detection of phagosome signals. Cell Microbiol 10(3):576–582
Gunn JS (2008) The Salmonella PmrAB regulon: lipopolysaccharide modifications, antimicrobial peptide resistance and more. Trends Microbiol 16(6):284–290
Fernandez L, Gooderham WJ, Bains M, McPhee JB, Wiegand I, Hancock RE (2010) Adaptive resistance to the “last hope” antibiotics polymyxin B and colistin in Pseudomonas aeruginosa is mediated by the novel two-component regulatory system ParR-ParS. Antimicrob Agents Chemother 54(8):3372–3382
Fernandez L, Jenssen H, Bains M, Wiegand I, Gooderham WJ, Hancock RE (2012) The two-component system CprRS senses cationic peptides and triggers adaptive resistance in Pseudomonas aeruginosa independently of ParRS. Antimicrob Agents Chemother 56(12):6212–6222
Ernst RK, Adams KN, Moskowitz SM, Kraig GM, Kawasaki K, Stead CM, Trent MS, Miller SI (2006) The Pseudomonas aeruginosa lipid A deacylase: selection for expression and loss within the cystic fibrosis airway. J Bacteriol 188(1):191–201
Burrows LL, Chow D, Lam JS (1997) Pseudomonas aeruginosa B-band O-antigen chain length is modulated by Wzz (Ro1). J Bacteriol 179(5):1482–1489
Wang J, Lory S, Ramphal R, Jin S (1996) Isolation and characterization of Pseudomonas aeruginosa genes inducible by respiratory mucus derived from cystic fibrosis patients. Mol Microbiol 22(5):1005–1012
Yi EC, Hackett M (2000) Rapid isolation method for lipopolysaccharide and lipid A from gram-negative bacteria. Analyst 125(4):651–656
El Hamidi A, Tirsoaga A, Novikov A, Hussein A, Caroff M (2005) Microextraction of bacterial lipid A: easy and rapid method for mass spectrometric characterization. J Lipid Res 46(8):1773–1778
Irazoqui JE, Troemel ER, Feinbaum RL, Luhachack LG, Cezairliyan BO, Ausubel FM (2010) Distinct pathogenesis and host responses during infection of C. elegans by P. aeruginosa and S. aureus. PLoS Pathog 6:e1000982
Vigneshkumar B, Pandian SK, Balamurugan K (2012) Regulation of Caenorhabditis elegans and Pseudomonas aeruginosa machinery during interactions. Arch Microbiol 194(4):229–242
Wilkinson SG, Galbraith L, Anderton WJ (1983) Lipopolysaccharides from Pseudomonas maltophilia: composition of the lipopolysaccharide and structure of the side-chain polysaccharide from strain N.C.I.B. 9204. Carbohydr Res 112(2):241–252
Adone R, Muscillo M, La Rosa G, Francia M, Tarantino M (2011) Antigenic, immunologic and genetic characterization of rough strains B. abortus RB51, B. melitensis B115 and B. melitensis B18. PLoS One 6(10):e24073
Trent MS, Stead CM, Tran AX, Hankins JV (2006) Diversity of endotoxin and its impact on pathogenesis. J Endotoxin Res 12(4):205–223
Park BS, Song DH, Kim HM, Choi BS, Lee H, Lee JO (2009) The structural basis of lipopolysaccharide recognition by the TLR4-MD-2 complex. Nature 458(7242):1191–1195
Lam JS, MacDonald LA, Lam MY, Duchesne LG, Southam GG (1987) Production and characterization of monoclonal antibodies against serotype strains of Pseudomonas aeruginosa. Infect Immun 55(5):1051–1057
Lam JS, Taylor VL, Islam ST, Hao Y, Kocincova D (2011) Genetic and functional diversity of Pseudomonas aeruginosa lipopolysaccharide. Front Microbiol 2:118
Antebi A, Yeh WH, Tait D, Hedgecock EM, Riddle DL (2000) daf-12 Encodes a nuclear receptor that regulates the dauer diapause and developmental age in C. elegans. Genes Dev 14(12):1512–1527
Snow MI, Larsen PL (2000) Structure and expression of daf-12: a nuclear hormone receptor with three isoforms that are involved in development and aging in Caenorhabditis elegans. Biochim Biophys Acta 1494(1–2):104–116
Riddle DL, Swanson MM, Albert PS (1981) Interacting genes in nematode dauer larva formation. Nature 290(5808):668–671
Lin K, Hsin H, Libina N, Kenyon C (2001) Regulation of the Caenorhabditis elegans longevity protein DAF-16 by insulin/IGF-1 and germline signaling. Nat Genet 28(2):139–145
Kwon ES, Narasimhan SD, Yen K, Tissenbaum HA (2010) A new DAF-16 isoform regulates longevity. Nature 466(7305):498–502
Pujol N, Link EM, Liu LX, Kurz CL, Alloing G, Tan MW, Ray KP, Solari R, Johnson CD, Ewbank JJ (2001) A reverse genetic analysis of components of the Toll signaling pathway in Caenorhabditis elegans. Curr Biol 11(11):809–821
Acknowledgments
We sincerely acknowledge Dr. Michael Givskov, at the Faculty of Health Sciences, University of Copenhagen for providing the Pseudomonas aeruginosa PAO1 strain. We thank The Director, CECRI for permitting us to use the Central Instrumentation Facility for NMR analysis and also we thank Mr. V. M. Shanmugam, CECRI for his technical support. We are indebted to Prof. K. Dharmalingam, Director-Research and Dr. Chitra Thangavel, Aravind Medical Research Foundation, Madurai, India for providing human corneal epithelial cells for the study. Also we gratefully acknowledge the use of the Bioinformatics Infrastructure Facility, Alagappa University funded by the Department of Biotechnology, Ministry of Science and Technology, Government of India (No- BT/BI/25/001/2006). This work was supported in part by the Grants from DBT, DST, ICMR (stipend to BV), ITC-AU collaborative project and CSIR of India to KB.
Author information
Authors and Affiliations
Corresponding author
About this article
Cite this article
Vigneshkumar, B., Radhakrishnan, S. & Balamurugan, K. Analysis of Pseudomonas aeruginosa PAO1 Lipid A Changes During the Interaction with Model Organism, Caenorhabditis elegans . Lipids 49, 555–575 (2014). https://doi.org/10.1007/s11745-014-3898-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11745-014-3898-3