Abstract
The aim of this study was to investigate molecular basis of germination inhibition of Triticosecale under the influence of exogenous ABA. Seedlings were isolated from seeds after 48 h of germination in water (control sample) and 100 μM ABA solution. It was observed that the applied concentration of phytohormone caused a significant inhibition of germination and a suppressed contribution of polysomes in the total ribosomal fraction. To identify proteins whose expression was affected by ABA, a proteomic approach employing 2D electrophoresis was used. Proteome maps for both control and ABA-treated samples displayed over 2,200 Coomassie Brilliant Blue-stained spots each. Protein spots showing either increase or decrease in spot intensity under the ABA influence were identified by mass spectrometry. Many of the identified proteins whose expression was stimulated by ABA proved to be stress-related, involved in nucleotide metabolism or playing a regulatory and signal transduction role, whereas proteins whose expression decreased are known to be involved in numerous metabolic pathways (including energy, carbohydrate or nucleotide metabolism) as well as related to genetic information processing and protein synthesis. Our results show that the germination inhibition caused by ABA is a result of multigene, diversified actions leading together to triticale growth suppression.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Annual survival of plant species depends on its ability to withstand unfavorable environmental conditions and on its efficiency of reproduction. Seeds do not only play an extremely important role as dispersal units in the plant life cycle but are also one of the most vital components of the world’s diet. Triticale has a high crop potential and is readily used in animal breeding as a fodder of high alimentary values, good digestibility and yield efficiency (Boulter and Parthier 1982).
Physiological and biochemical processes in plant organisms are under the continuous control of phytohormone regulation. Abscisic acid (ABA) is a universal plant growth regulator implicated in the control of many developmental and physiological processes during the whole plant life. It is commonly known as a growth inhibitor and plays an important role in embryo development, stress response, senescence, etc. By the ability to induce and, most likely, maintain dormancy, ABA is known to circumvent seed germination and seedling growth in the early stages of its growth (Kucera et al. 2005; Bewley 1997; Koornneef et al. 2002; Pawłowski 2007), which naturally prevents precocious germination when still on the mother plant. Pre-harvest sprouting may cause severe economic problems for agriculture by yield drop as well as diminishing grain functionality and viability.
It was observed before that by incubating mature seeds in exogenous solution of abscisic acid, the prevention of radicle extension and therefore the completion of germination may be strongly delayed as long as the inhibitor is present (Schopfer and Plachy 1985). Although many studies have been performed to gain the understanding of ABA involvement in seed germination and dormancy (Garciarubrio et al. 1997; Ananieva and Ananiev 1997, Pawłowski 2009), the process is still not fully understood.
During the whole plant life, ABA causes desiccation tolerance acquisition, which is essential during seed maturation. It was observed that stress conditions such as dehydration or excessive salinity may also cause similar proteins accumulation (Close 1997; Borovskii et al. 2002). The level of protein synthesis at the time is determined by the mRNA synthesis and the efficiency of translation. The dynamics of this process may be correlated with the formation of polysomes. Investigation of polyribosome contribution gives some valuable information concerning the overall capacity of protein synthesis in a tissue.
2D PAGE analysis allows a global analysis of the changes in proteins expression. Some papers have already been published on the subject of plant proteomics concerning seed germination, dormancy breaking (Gallardo et al. 2001; Cangahuala Inocente et al. 2009; Yang et al. 2007; Pawłowski 2007, 2009), or the influence of various growth regulators on proteome alterations (Pawłowski 2007). However, the complicated mechanisms of germination and dormancy breaking are still not fully understood. Proteomic studies give a powerful tool for identifying alterations in protein expression in response to a variety of factors, including hormonal stimuli. In combination with mass spectrometry and accessible databases, it gives enormous possibilities in the attempt to understand many physiological processes occurring in cells (Pawłowski 2007).
The main aim of this experiment was to investigate the molecular basis of ABA involvement in germination suppression. In order to achieve this aim, the dynamics of protein translation and alterations in proteome profiles of triticale embryos were examined.
Materials and methods
Plant material and germination conditions
In this study, caryopses of cv Ugo triticale (Triticosecale Witt.) isolated from germinating seeds were used. Seeds were surface sterilized in 0.5 % sodium hypochlorite for 20 min, washed thoroughly with tap water and afterwards rinsed with distilled water. 50 grains were placed on a filter paper layer in 15 cm Petri dishes containing 20 ml of distilled water (control conditions) or 100 μM aqueous solution of abscisic acid (ABA). Seeds were germinating in the dark at 21 °C for 24, 48 and 72 h. After these time periods, embryos were isolated from seeds. All the experiments were performed in triplicates.
Polyribosomes isolation
After 24, 48 or 72 h of germination, caryopses of 2 g fresh weight were ground using mortar and pestle in 16 ml of buffer U (2 % PTE, 1 % DOC, 200 mM Tris–HCl pH 8.5, 50 mM potassium acetate, 25 mM magnesium acetate, 2 mM EGTA and 100 mg/ml Heparin), according to the method of Abe et al. (1992), and pelleted at 12,000×g for 20 min.
The extract was purified and enriched by pelleting through 50 % sucrose pads at 330,000×g for 90 min in a Beckman SW 55 Ti rotor. The pellet containing the total extractable fraction of polyribosomes was resuspended in buffer U. 200 μl of aliquot (about 10 OD units) was fractionated on a gradient of 15–60 % (w/v) solution of sucrose in buffer B (50 mM Tris–HCl pH 8.5, 20 mM potassium acetate, 10 mM magnesium acetate) at 330,000×g for 45 min. The ribosome profile was monitored by measuring the absorbance at 254 nm using a UA-5 flow recorder (ISCO, Lincoln, NE, USA) according to Davies and Abe (1995). All steps were carried at 0–4 °C. The level of polysomes in the overall ribosomal fraction was determined by excising the area under the polyribosome profile from the total fraction of polysomes, together with monosomes and ribosomal subunits. All chemicals, if not indicated otherwise, were acquired from Sigma-Aldrich.
Proteome extraction
Proteome was extracted from embryonic tissue after 48 h of germination, as recommended by Gallardo et al. (2002a, b). 300 mg of tissue was powdered in liquid nitrogen and suspended in chilled lysate buffer containing 7.5 M urea, 2.2 M thiourea, 63 mM CHAPS, 0.25 % Triton X-100 (Bio-Rad), 5 mM Trizma base, 21 mM Trizma hydrochloride, 1.1 % ampholytes (pH 3–10), 60 U/ml DNase I, 72 μg/ml RNase A (5.8 Kunitz units/ml), 14 % protease inhibitors (Cocktail complete mini, Roche) and 8 mM DTT (Bio-Rad). Each sample was mixed, shaken on ice for 45 min and centrifuged several times at 12,000×g for 10 min in order to obtain clear supernatant. The protein concentration was measured at 280 nm using a NanoDrop 1000 spectrophotometer (Thermo Scientific).
Protein 2-D PAGE electrophoresis
Two dimensional electrophoresis in polyacrylamide gel (2D PAGE) can separate a complex protein mixture according to their isoelectric point (pI) in the first dimension and molecular mass in the second. First dimension of 2D PAGE was performed in a 24 cm Immobiline DryStrip Gel strip with a linear pH gradient from 3 to 10 (GE Healthcare) in an Ettan IPGphor 3 system (GE Healthcare), following the manufacturer’s instructions. The protein extract was purified using 2D Clean-Up Kit (GE Healthcare). Prior to isoelectrofocusing, 2 mg of proteins were solubilized in rehydration buffer (7 M urea, 2 M thiourea, 2 % CHAPS, 0.5 % ampholytes (pH 3–10) (Amersham), 10 mM DTT and 0.002 % bromophenol blue) to a total volume 450 μl. Directly before the second dimension, the strip (containing separated proteins) was equilibrated in a glass tube in equilibration buffer containing 6 M urea, 75 mM Tris–HCl (pH 6.8), 30 % glycerol and 26 mM DTT (in the first step) or 220 mM iodoacetamide instead of DTT (in the second step) for 15 min each. In order to separate the proteins of the same pI according to their molecular mass, the strip was placed on top of a Precast DaltGel 12.5 (GE Healthcare) and sealed with 10 % agarose solubilized in electrode buffer containing 25 mM Trizma base, 195 mM glycine and 0.1 % SDS (Laemmli 1970). Electrophoresis was carried in an EttanDalt Six electrophoresis unit (GE Healthcare) using Laemmli buffer system. To estimate molecular masses of proteins, Low Molecular Mass Standards (Bio-Rad) were used.
Analysis of 2DE gels
After 2D PAGE, the gel was stained with Coomassie Brilliant Blue G-250 according to Neuhoff et al. (1988) and scanned on an Image Scanner using Labscan 3.0 software (GE Healthcare). The protein pattern was analyzed using ImageMaster 2D Platinum 6.0 programme. After spot detection the quantitative determination of spot intensity, including a normalization following the method of principal component analysis (Wibig 2007), was performed, taking into consideration the average values of background optical density (OD) and spots intensity on each gel. The obtained data were subjected to a t test statistical analysis. 12 of up- and 12 of down-regulated proteins, who showed at least twofold difference in spot intensity in response to ABA, were cut from the gel and subjected to a mass spectrometry identification.
Mass spectrometry
Spots of interest were analyzed by liquid chromatography coupled with a Hybrid-2D-Linear Quadrupole Ion Trap-Fourier Transform Ion Cyclotron Resonance (LTQ-FT-ICR) mass spectrometer (Thermo Electron Corp., San Jose, CA, USA) in the Institute of Biochemistry and Biophysics (Polish Academy of Sciences, Warsaw, Poland). Prior to the analysis, gel slices were subjected to a standard in-gel digestion procedure (Shevchenko et al. 1996) during which gel pieces were washed with 50 % (v/v) acetonitrile, 10 mM ammonium bicarbonate solution, dehydrated with 100 % acetonitrile and completely dried in a Speedvac concentrator. Proteins were digested overnight with Sequencing Grade Modified Trypsin (Promega V5111). Peptides were eluted from gel with 0.1 % TFA, 2 % acetonitrile and applied to RP-18 precolumn (LC Packings) using 0.1 % TFA as a mobile phase. Afterwards, peptide mixtures were transferred to nano-HPLC RP-18 column (Waters), using 0–60 % acetonitrile gradient in 30 min in the presence of 0.05 % formic acid (150 nl/min flowrate). Column outlet was directly coupled with ion source of LTQ-FT-MS, working in the regime of data-dependent MS to MS/MS switch. The acquired raw data were processed by Mascot Distiller followed by Mascot Search (Matrix Science, London, UK, locally installed http://proteom.pl/mascot) against NCBI non-redundant database. Search parameters for precursor and product ions mass tolerance were, respectively, ±40 ppm and ±0.8 Da (Pappin et al. 1993).
Results
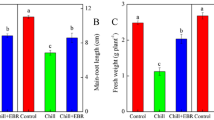
In this study, triticale seeds were subjected to germination in 100 μM aqueous solution of phytohormone abscisic acid (ABA). Measurements of the fresh weight content of embryonic tissues after 48 h of germination showed significant differences between control seedlings and those subjected to ABA. Their masses amounted to 23.79 and 10.45 mg per seedling, respectively. After 72 h of germination, the masses grew to 44.77 mg in control and 12.28 mg in ABA-treated seedlings (Fig. 1). Comparing the values of fresh weight content after both time periods, it is clear that ABA caused a significant suppression in seedlings fresh weight accumulation during early stages of triticale seed germination.
The tendency was similar in the polysome content in the total ribosomal fraction (monosomes, polysomes and ribosomal subunits) of embryonic tissues. Figure 2 presents sedimentation profiles of the total ribosomal fractions of triticale seedlings separated in a sucrose gradient 15–60 %. The analysis of the polysome population contribution in the total ribosomal fraction showed an increase from 18.6 % after 24 h of germination under the control conditions up to 65.6 % after 48 h and a slight decrease to 64.3 % in 72 h seedlings. ABA showed an inhibitory effect on the contribution of the polysome population content. The amount of polysomes in the total ribosomal fraction of caryopses treated with exogenous ABA was greatly reduced. After 24 h of germination, it constituted only 4.6 % of the total ribosomal fraction, 24 h later 22.3 % (nearly 3 times less than in the control at the same time) and after 72 h of germination it decreased to 18.1 %.
It has been observed that after 48 h of germination, caryopses displayed the first significant difference in the fresh weight content caused by the ABA application and the maximum of polysome formation. Due to this fact, the proteomes of caryopses were analyzed at that time. In order to test the protein expression alterations caused by ABA, the 2D PAGE method coupled with mass spectrometry was used. It was observed that ABA not only caused a decrease of some protein expression but also up-regulated the expression of others. The 2D proteome profiles in the pH range from 3 to 10 were established for control and ABA-treated caryopses. Figure 3 shows the achieved electrophoretic patterns with the identified proteins marked with arrows. Proteins whose expression was stimulated by ABA (spots 1–12) are presented in Table 1 and those whose expression decreased (spots 13–24) are shown in Table 2.
Two-dimensional electrophoretic patterns of embryonic proteins obtained from Triticosecale seedlings germinating for 48 h in water—control (a) and in 100 μM solution of abscisic acid (b). Isoelectrofocusing was performed in the 24 cm polyacrylamide strips with the immobilized pH gradient ranging from 3 to 10. Proteins were visualized using Coomassie Brilliant Blue G-250. Identified spots are marked with arrows. On the left side of each gel was added a marker of protein molecular weight (kDa)
Discussion
Seed germination is one of the most critical processes in plant life, when quiescent embryos transform into active, viable new organisms. It has already been noted that ABA blocks reserve mobilization by repressing α-amylase gene transcription and therefore restricts the availability of energy and metabolites for growing embryos, alters seed carbon and nitrogen metabolism (Garciarubrio et al. 1997), inhibits proteins and RNA synthesis (Ananieva and Ananiev 1997), water uptake, endosperm rupture, restrain further embryo extension and seedling growth after radicle emergence (Schopfer and Plachy 1985; Kucera et al. 2005). The suppression of germination is then a result of changes in many various processes occurring in tissues, often initiated by expression of the ABA-responsive genes. The present results show that an exogenously applied growth regulator, namely ABA in 100 μM concentration, significantly influences on the physiological and biochemical behavior of germinating triticale seeds.
In this work we show that ABA treatment has a strong inhibitory impact on polysome participation in protein synthesis during seed germination. Very few studies on this subject exist, mainly completed by Weidner et al. (1991, 2003, 2006). Formation of a polysome population in the presence of ABA proved to be suppressed more than threefold during the whole time of the experiment. Abscisic acid inhibits transcription but increases RNA stability (Weidner et al. 2006), therefore protein biosynthesis takes part mainly on the mRNAs preserved in desiccated embryos. Kawaguchi et al. (2004) showed that dehydration stress causes the reduction of mRNAs recruited to polysomes, what indicates that the effect of the restriction of translation mediated by ABA and dehydration may be related. The suppression of polysome formation may be a result of ABA-mediated reduction of translation initiation, diminished rate of elongation, termination or mRNA availability.
The results concerning changes in the polysome content show a slight decrease in the polysome content after 72 h of germination in comparison to 48 h seedlings both in control and ABA-treated samples. Diminishing amount of the transcripts incorporated into polysomes in the seedlings may be an indicator of triggering post-germination processes such as reduction of cell division in favor of higher extension rate and higher water uptake. Long exposition to the phytohormone may be one of the causes of low water input and polysome formation reduction in ABA-treated seedlings.
2D PAGE analysis of the proteomes of ABA-treated and non-treated seedling showed differences in proteins expression. Identified proteins were classified according to their functions into categories such as stress response, metabolism, signal transduction, protein synthesis and genetic information processing, protein degradation, cell structure.
Stress response
Many of the identified proteins can be included in the group of stress-related proteins. It is known that ABA enhances stress tolerance in plants, especially desiccation tolerance in the late stages of seed development, by induction of stress-related proteins production (Yang et al. 2007; Li et al. 2010). Because the whole process of germination might be described as the reverse of seed maturation, it is not surprising that some of the proteins involved in embryo development are still present under the influence of exogenously applied ABA, although normally during the imbibition stage of germination their abundance gradually diminishes (Han and Kermode 1996; Ostergaard et al. 2004). Abscisic acid in this work caused an increase in the expression of some LEA (late embryogenesis proteins—spots 1, 6, 7, 8), dehydrin DHN6 (spot 2), NAD-dependent formate dehydrogenase (spot 10), USP family protein (spot 4), and glutathione transferase (spot 12).
LEAs are known to be ABA responsive (Finklelstein and Rock 2002; Grelet et al. 2005) and protect developing embryos from stresses by replacing water and maintaining protein and membrane structure (Close 1997; Han and Kermode 1996). DHN6 was another stress protein up-regulated by ABA. The increase of this protein expression was especially high (more than threefold). Dehydrins are known to play protective functions, preventing protein coagulation and plasma membrane destabilization, what is important for embryo prevention during seed maturation. The up-regulation of DHN6 occurs not only after ABA treatment but also under low water availability or cold stress condition (Rinne et al. 1999; Borovskii et al. 2002) but we still lack the complete knowledge about their exact role in the cell. The up-regulation of NAD-dependent formate dehydrogenase and USP family protein gives another evidence of ABA cross-talk with stress response pathways. The former, due to the ability to oxidize accumulating under stress formate into CO2 in the presence of NAD+, may complement the respiration when the efficiency of Krebs cycle diminishes. Both the transcript and the enzyme were detected under many stresses such as hypoxia, drought, cold, wounding, and osmotic stress (Ambard-Bretteville et al. 2003; Kim et al. 2007; Li et al. 2010). The latter, USP family proteins, are also often expressed in many species under the unfavorable conditions, providing the organism with the ability to protect from various stresses and therefore this proteins constitute a useful target in stress tolerance engineering. Suggested function of the proteins containing USP domain is nucleotide binding and signal transduction (Nachin et al. 2005; Isokpehi et al. 2011). Its function under ABA conditions has not been studied yet.
Also glutathione transferases are members of a diversified group of enzymes protecting cells from a wide range of reactive substrates. Those genes were already found to be responsive to abscisic acid as well as many abiotic stresses (Xu et al. 2002). They are involved in detoxification and became a marker of plant response to stress (Dixon et al. 2002). On the other hand, we found that one of the proteins from this group, glutathione transferase F4 (spot 16), was down-regulated in response to applied abscisic acid. Peng et al. (2009) examined two strains of wheat differing in stress tolerance and obtained the substantial up-regulation of glutathione transferase F4 under stress conditions only in the tolerant strain. This data suggest that the ability to generate detoxification reactions with this protein involvement is dependent on the stress susceptibility of the species. It could be also concluded that not all of the stress and ABA-responsive pathways are common.
ABA caused also a negative effect on HSC70 (heat shock protein cognate) expression (spots 19, 20, 23). HSC70 belong to the group of chaperones, which bind to newly synthesized proteins, preventing from aggregation and improper folding during a very active time of plant development (Sung et al. 2001). The low level of heat shock proteins seems to coincide with the low-efficient protein yield and the germination inhibition in the presence of ABA.
On the basis of the previous results as well as the results of this study, we may observe an evident cross-talk between ABA and stress response. It is not surprising, since stress-inducible genes expression in many cases is conditioned by ABA-dependent phosphorylation of transcriptional factors binding to ABRE (ABA-responsive elements) regulatory sequences (Yamaguchi-Shinozaki and Shinozaki 2006). Abscisic acid is a main player in setting a seed dormancy and triggers the processes allowing the embryo to survive in a quiescent state. Considering the severe conditions naturally imposed on seeds during desiccation, the relationship between stress and the phytohormone is indispensable.
Metabolism
The results of our work indicate that abscisic acid regulates many aspects of cell energy, carbohydrate, and nucleotide metabolism and causes complex and diversified effects on cell functions. Our findings indicate that ABA-mediated germination inhibition rely to a large extent on metabolism suppression. Germination and subsequent steps of seedling growth require large amounts of energy released from the seed storage reserves and converted into energy mainly by glycolysis. Yang et al. (2007) identified many glycolytic enzymes up-regulated during barley germination. In our study, the applied concentration of ABA has shown an inhibitory effect on the synthesis of many proteins involved in metabolism such as glyceraldehyde-3-phosphate dehydrogenase (spot 15), glucose and ribitol dehydrogenase homolog (spot 17), dehydroascorbate reductase (spot 18), and ATP synthase beta subunit (spot 21). Glyceraldehyde-3-phosphate dehydrogenase (GAPDH) is the enzyme involved not only in glycolysis but also in other functions related to plant stress. It was demonstrated that the amount of GAPDH transcripts in rice grows significantly in response to exogenous ABA treatment and drops considerably after few hours (Zhang et al. 2011). Although most of this protein functions in relation to stress remain largely unknown, it was suggested that it may take part in protection against high ROS levels and generate NADPH when other proteins undergo inactivation or degradation (Chang et al. 2000; Bustos et al. 2008).
Down-regulation of ATP synthase beta subunit suggests the contribution of ABA to regulating the energy metabolism in germinating seedlings. Glucose and ribitol dehydrogenase is involved in an alternative carbohydrate metabolism and its gene is known to be ABA responsive (Ostergaard et al. 2004). Dehydroascorbate reductase (identified as spot 18) reduces dehydroascorbate to ascorbic acid, which protects cells against oxidative stress. Deficiency of this protein regulates cell elongation and cell cycle, what was already shown to lead to suppression of leaf expansion, deceleration of root growth and delay of flowering (Chen and Gallie 2006).
The up-regulated protein in this category is nucleoside diphosphate kinase (NDPK) (spot 5). As a housekeeping enzyme, it provides nucleoside phosphates for nucleic acid synthesis and plays a role in signal transduction. Its function is not entirely known yet. The results obtained in this work are not consistent with the results of Yano et al. (1995) or Kim et al. (2008), who detected an increase of NDPK during germination and seedling growth. However, Harris et al. (1994) detected the up-regulation of NDPK as a result of stress response, which suggests that the role of NDPK in plants is diversified and needs further investigation.
Signal transduction
GTP-binding proteins are important in a cell signal transduction chain initiated by phytohormones (Terryn et al. 1993), therefore the especially high (more than threefold) up-regulation of triticale GTP-binding protein (spot 9) expression by ABA is not surprising.
Protein synthesis/genetic information processing
Uwer et al. (1998) suggest that an arrest of embryo growth between globular and heart stages may be caused by the loss of Glycyl-tRNA synthetase (spot 13) during embryogenesis. Abscisic acid in our study caused a decrease in the protein showing similarity to this enzyme. This may suggest that ABA is directly responsible for maintenance of dormancy and inhibition of germination by the inhibition of specific protein expression.
A responsiveness of the ribosomal protein S7 (corresponding to spot 22) to abscisic is a novel finding. The protein reveals down-regulation in relation to the phytohormone. It is known as an initiator of 30S ribosomal subunit assembly and plays important role in protein synthesis by implication in tRNA and mRNA interactions (Donotsova et al. 1991). The down-regulation of this protein may be one of the explanations of the decrease of polysome formation and growth inhibition obtained in this work in the presence of exogenously applied ABA.
Protein degradation
The main player in a protein degradation pathway of plant cells is 26S proteasome. It may regulate numerous processes, such as division or development by degradation of unwanted or regulatory proteins; it can ensure proper protein folding and assembling of complexes by controlling ubiquitin-mediated degradation (Wolf and Hilt 2004; Borghetti et al. 2002). Expression of 26S proteasome regulatory particle triple-A ATPase subunit 4b (spot 3) was positively (more than threefold) modulated by ABA, indicating that protein cleavage is an especially important control point in processes connected to germination inhibition and dormancy breaking.
Cell structure
Two of the identified proteins are involved in cell structure formation (Alpha-SNAP and α-tubulin). The up-regulated protein identified as Alpha-SNAP (spot 11) is important for proper cellular organization, division and expansion. It is also crucial for intracellular vesicle trafficking and fusion (Bachem et al. 2000; Leyman et al. 2000). The role of alpha-SNAP in ABA-mediated germination suppression has not been reported yet, but it was found to be important in a process of stomatal closure triggered by ABA on a non-transcriptional level (Geelen et al. 2002). It is involved in starch and soluble sugars accumulation in leaves (Bachem et al. 2000). The other protein from the group, down-regulated α-tubulin, marked here as spot 24, is one of the key components in controlling seed germination (Pawłowski 2009). Giani et al. (1998) indicated that a drop in α-tubulin transcript is correlated with the inhibition of coleoptile elongation in response to ABA in rice. Olinevich et al. (2002) observed a decrease in microtubules and α-tubulin content in wheat roots in the presence of the hormone, together with an enhancement in the stability of the microtubules, what affirms ABA-related suppression of cell extensibility, leading to growth retardation.
The down-regulated hypothetical protein Os09g0535300 has not been characterized yet, either for its responsiveness to the phytohormone or for its functions under stress conditions; however, the ethylene-responsive homolog protein XAP-5 (CIRCADIAN TIMEKEEPER XCT) from A. thaliana is suggested to play a role in ethylene-induced growth inhibition and hypocotyl shortening by modulating the activity of some transcription factors. It may also be connected to a circadian clock and signal transduction (Ellison et al. 2011).
Conclusions
As expected, exogenously applied abscisic acid caused inhibition of germination of triticale seeds. It has also been shown to play a suppressive role in polysome formation and to influence on expression of various proteins, altogether leading to the inhibition of growth and development. On the basis of our results we may conclude that the applied phytohormone mainly decelerates metabolism, protein translation, cell elongation and influences the expression of stress-related proteins in triticale.
Author contribution
Ewa Szypulska and Stanisław Weidner were responsible for experimental design, Sylwia Świgońska provided all the technical support, Anna Badowiec performed the research and prepared the manuscript under the guidance of Stanisław Weidner.
Abbreviations
- 2D PAGE:
-
Two dimensional electrophoresis in polyacrylamide gel
- ABA:
-
Abscisic acid
- CHAPS:
-
3-[(3-cholamidopropyl)dimethylammonio]-1-propanesulfonate
- DOC:
-
Sodium deoxycholate
- DTT:
-
Dithiothreitol
- EGTA:
-
Ethylenoglycol-O–O′-bis(2-aminoetyl)-N,N,N′,N′tetraacetic acid
- HEPES:
-
4-(2-hydroxyethyl)-1-piperazineethanesulfonic acid
- IEF:
-
Isoelectrofocusing
- M:
-
Monosomes
- pI:
-
Isoelectric point
- PTE:
-
Polyoxyethylene-10-tridecyl ether
- P:
-
Polysomes
- S:
-
Ribosome subunits
References
Abe S, Ito Y, Davies E (1992) Co-sedimentation of actin, tubulin and membranes in cytoskeleton fractions from peas and mouse 3T3 cells. J Exp Bot 43:941–949
Ambard-Bretteville F, Sorin C, Rébeillé F, Hourton-Cabassa C, Colas des Francs-Small C (2003) Repression of formate dehydrogenase in Solanum tuberosum increases steady-state levels of formate and accelerates the accumulation of proline in response to osmotic stress. Plant Mol Biol 52:1153–1168
Ananieva K, Ananiev ED (1997) Comparative study of the effects of methyl jasmonate and abscisic acid on RNA and protein synthesis in excised cotyledons of Cucurbita pepo L. (Zucchini). Bulg J Plant Physiol 23:80–90
Bachem CWB, Oomen RJFJ, Kuyt S, Horvath BM, Claassens MMJ, Vreugdenhil D, Visser RGF (2000) Antisense suppression of a potato alpha-SNAP homologue leads to alterations in cellular development and assimilate distribution. Plant Mol Biol 43:473–482
Bewley JD (1997) Seed germination and dormancy. Plant Cell 9:1055–1066
Borghetti F, Noda FN, Sa CM (2002) Possible involvement of proteasome activity in ethylene-induced germination of dormant sunflower embryos. Braz J Plant Physiol 14(2):125–131
Borovskii GB, Stupnikova IV, Antipina AI, Vladimirova SV, Voinikov VK (2002) Accumulation of dehydrin-like proteins in the mitochondria of cereals in response to cold, freezing, drought and ABA treatment. Plant Biol 2:5
Boulter D, Parthier B (eds) (1982) Nucleic acids and proteins in plants. Structure, biochemistry and physiology of proteins. Springer, Berlin
Bustos DM, Bustamante CA, Iglesias AA (2008) Involvement of non-phosphorylating glyceraldehyde-3 phosphate dehydrogenase in response to oxidative stress. J Plant Physiol 165(4):456–461
Cangahuala Inocente GC, Villarinon A, Seixa D, Dumas-Gaudot E, Terenzi H, Guerra MP (2009) Differential proteomic analysis of developmental stages of Acca sellowiana somatic embryos. Acta Physiologiae Plantarum 33:4–53
Chang WWP, Huang L, Shen M, Webster C, Burlingame AL, Roberts JKM (2000) Patterns of protein synthesis and tolerance of anoxia in root tips of maize seedlings acclimated to a low-oxygen environment, and identification of proteins by mass spectrometry. Plant Physiol 122:295–318
Chen Z, Gallie DR (2006) Dehydroascorbate reductase affects leaf growth, development, and function. Plant Physiol 142:775–787
Close TJ (1997) Dehydrins: a commonality in the response of plants of dehydration and low temperature. Physiol Plant 100:291–296
Davies E, Abe S (1995) Methods for isolation and analysis of polyribosomes. In: erl-Hahn D (ed) Methods in plant cell biology, Part B in methods in cell biology, vol 50. Academic press Inc., USA, pp 209–222
Dixon DP, Lapthorn A, Edwards R (2002) Plant glutathione transferases. Genome Biol 3(3):3004.1–3004.10
Donotsova O, Kopylov A, Brimacombe R (1991) The location of mRNA in the ribosomal 30S initiation complex; site-directed cross-linking of mRNA analogues carrying several photo-reactive labels simultaneously on either side of the AUG start codon. EMBO J 10(9):2613–2620
Ellison CT, Vandenbussche F, Van Der Straeten D, Harmer S (2011) Xap5 circadian timekeeper regulates ethylene responses in aerial tissues of Arabidopsis. Plant Physiol 155:988–999
Finklelstein RR, Rock CD (2002) Abscisic acid biosynthesis and response. In: Somervile CR, Meyerovitz EM (eds) The Arabidopsis book. American Society of plant Biologists, Rockville, pp 1–18
Gallardo K, Claudette J, Groot SPC, Puype M, Demol H, Vandekerckhove J, Job D (2001) Proteomic analysis of Arabidopsis seed germination and priming. Plant Physiol 126:835–848
Gallardo K, Job C, Groot SPC, Puype M, Demol H, Vandekerckhove J, Job D (2002a) Importance of methionine biosynthesis for Arabidopsis seed germination and seedling growth. Physiol Plant 116:238–247
Gallardo K, Job C, Groot SPC, Puype M, Demol H, Vandekerckhove J, Job D (2002b) Proteomics of Arabidopsis seed germination. A comparative study of wild type and gibberellin deficient seeds. Plant Physiol 129:823–837
Garciarubrio A, Legaria JP, Covarrubias AA (1997) Abscisic acid inhibits germination of mature Arabidopsis seeds by limiting the availability of energy and nutrients. Planta 203:182–187
Geelen D, Leyman B, Batoko H, Di Sansebastiano GP, Moore I, Blatt MR (2002) The abscisic acid–related SNARE homolog NtSyr1 contributes to secretion and growth. Evidence from competition with its cytosolic domain. Plant Cell 14:387–406
Giani S, Qin X, Faoro F, Breviario D (1998) In rice, Oryzalin and abscisic acid differentially affect tubulin mRNA and protein levels. Planta 205:334–341
Grelet J, Benamar A, Teyssier E, Avelange-Marcherel MH, Grunwald D, Macherel D (2005) Identification in pea seed mitochondria of a late-embryogenesis abundant protein able to protect enzymes from drying. Plant Physiol 137:157–167
Han B, Kermode AR (1996) Dehydrin-like proteins in castor bean seeds and seedlings are differentially produced in response to ABA and water-deficit-related stress. J Exp Bot 42:933–939
Harris N, Taylor JE, Roberts JA (1994) Isolation of mRNA encoding a nucleoside diphosphate kinase from tomato that is up-regulated by wounding. Plant Mol Biol 25:739–742
Isokpehi RD, Simmons SS, Cohly HHP, Ekunwe SIN, Begonia GB, Ayensu WK (2011) Identification of drought-responsive universal stress proteins in Viridiplantae. Bioinform Biol Insights 5:41–58
Kawaguchi R, Girke T, Bray EA, Bailey-Serres J (2004) Differential mRNA translation contributes to gene regulation under non-stress and dehydration stress conditions in Arabidopsis thaliana. Plant J 38:823–839
Kim JK, Cho MR, Baek HJ, Ryu TH, Kim JB, Kim JH, Kim MJ, Yu CY, Fukusaki EI, Kobayashi A (2007) Quantitation of formate in plants and its enhancement in response to environmental stresses. J Appl Biol Chem 50(4):211–214
Kim ST, Kang SY, Wang Y, Kim SG, Hwang DH, Kang KY (2008) Analysis of embryonic proteome modulation by GA and ABA from germinating rice seeds. Proteomics 8:3577–3587
Koornneef M, Bentsink L, Hilhorst H (2002) Seed dormancy and germination. Curr Opin Plant Biol 5:33–36
Kucera B, Cohn MA, Leubner-Metzger G (2005) Plant hormone interactions during seed dormancy release and germination. Seed Sci Res 15:281–307
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227(5259):680–685
Leyman B, Geelen D, Blatt MR (2000) Localization and control of expression of Nt-Syr1, a tobacco SNARE protein. Plant J 24(3):369–381
Li XJ, Yang MF, Chen H, Qu LQ, Chen F, Shen SH (2010) Abscisic acid pretreatment enhances salt tolerance of rice seedlings: proteomic evidence. Biochim Biophys Acta 1804:929–940
Nachin L, Nannmark U, Nystrom T (2005) Differential roles of the universal stress proteins of Escherichia coli in oxidative stress resistance, adhesion, and motility. J Bacteriol 187:6265–6272
Neuhoff V, Arold N, Taube D, Ehrhard W (1988) Improved staining in proteins in polyacrylamide gels including isoelectric focusing gels with clear background at nanogram sensitivity using Coomassie Brilliant Blue G-250 and R-250. Electrophoresis 9(6):255–262
Olinevich OV, Khokhlova JP, Raudaskoski M (2002) The microtubule stability increases in abscisic acid-treated and cold-acclimated differentiating vascular root tissues of wheat. J Plant Physiol 159:465–472
Ostergaard O, Finnie Ch, Laugesen S, Roepstorf P, Svensson B (2004) Proteome analysis of barley seeds: identification of major proteins from two-dimensional gels (pI 4–7). Proteomics 4:2437–2447
Pappin DJC, Hojrup P, Bleasby AJ (1993) Rapid identification of proteins by peptide-mass fingerprinting. Curr Biol 3(6):327–332
Pawłowski TA (2007) Proteomics of Fagus sylvatica L. seed dormancy breaking: influence of abscisic and gibberellic acids. Proteomics 7:2226–2257
Pawłowski TA (2009) Proteome analysis of Norway maple (Acer platanoides L.) seeds dormancy breaking and germination: influence of abscisic and gibberellic acids. BMC Plant Biol 9:48
Peng Z, Wang M, Li F, Lv H, Li C, Xia G (2009) A proteomic study of the response to salinity and drought stress in an introgression strain of bread wheat. Mol Cell Proteomics 8(12):2676–2686
Rinne PLH, Kaikuranta PLM, Plas LHW, Schoot Ch (1999) Dehydrins in cold-acclimated apices of birch (Betula pubescens Ehrh.): production, localization and potential role in rescuing enzyme function during dehydration. Planta 209:377–388
Schopfer P, Plachy C (1985) Control of seed germination by abscisic acid. III. Effect on embryo growth potential (minimum turgor pressure) and growth coefficient (cell wall extensibility) in Brassica napus L. Plant Physiol 77:676–686
Shevchenko A, Wilm M, Vorm O, Mann M (1996) Mass spectrometric sequencing of proteins from silver stained polyacrylamide gels. Anal Chem 68:850–858
Sung DY, Vierling E, Guy CL (2001) Comprehensive expression profile analysis of the Arabidopsis Hsp70 gene family. Plant Physiol 126:789–800
Terryn N, Van Montagu M, Inze D (1993) GTP-binding proteins in plants. Plant Mol Biol 22:143–152
Uwer U, Willmitzer L, Altmann T (1998) Inactivation of a glycyl-tRNA synthetase leads to an arrest in plant embryo development. Plant Cell 10:1277–1294
Weidner S, Deckert J, Gwóźdź E (1991) Translational activity of polysomes from embryos during germination of wheat caryopses in the presence of abscisic acid. Acta Physiologiae Plantarum 13:175–182
Weidner W, Frączek E, Romanowska M, Amarowicz R, Abe S, Davies E (2003) The influence of abscisic acid on different polysomal populations in embryonal tissue during pea seeds germination. Acta Physiologiae Plantarum 25:5–12
Weidner S, Każarnowicz M, Frączek E, Amarowicz R, Karamać M (2006) Exogenous abscisic acid increases stability of polysomes in embryos of triticale caryopses during germination. Acta Physiologiae Plantarum 28:627–634
Wibig T (2007) Analiza danych PCA. Katedra Fizyki Teoretycznej, Wyd. Uniwersytetu Łódzkiego
Wolf DH, Hilt W (2004) The proteasome: a proteolytic nanomachine of cell regulation and waste disposal. Biochim Biophys Acta 1695:19–31
Xu F, Lagudah ES, Moose SP, Riechers DE (2002) Tandemly duplicated safener-induced glutathione S-transferase genes from Triticum tauschii contribute to genome- and organ-specific expression in hexaploid wheat. Plant Physiol 130:362–373
Yamaguchi-Shinozaki K, Shinozaki K (2006) Transcriptional regulatory networks in cellular responses and tolerance to dehydration and cold stresses. Annu Rev Plant Biol 57:781–803
Yang P, Li X, Wang X, Chen H, Chen F, Shen S (2007) Proteomic analysis of rice (Oryza sativa) seeds during germination. Proteomics 7(18):3358–3368
Yano A, Umeda M, Uchimiya H (1995) Expression of functional proteins of cDNA encoding rice nucleoside diphosphate kinase (NDPK) in Escherichia coli and organ-related alteration of NDK activities during rice seed germination (Oryza sativa L.). Plant Mol Biol 27:1053–1058
Zhang XH, Rao XL, Shi HT, Li RJ, Lu YT (2011) Overexpression of a cytosolic glyceraldehyde-3-phosphate dehydrogenase gene OsGAPC3 confers salt tolerance in rice. Plant Cell Tissue Org Cult 107:1–11
Conflict of interest
The authors state that they have no conflict of interest.
Open Access
This article is distributed under the terms of the Creative Commons Attribution License which permits any use, distribution, and reproduction in any medium, provided the original author(s) and the source are credited.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by S. Abe.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 2.0 International License (https://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
About this article
Cite this article
Badowiec, A., Świgońska, S., Szypulska, E. et al. Influence of exogenous abscisic acid on alterations in protein expression in the proteome of Triticosecale seedlings. Acta Physiol Plant 34, 2359–2368 (2012). https://doi.org/10.1007/s11738-012-1040-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11738-012-1040-9