Abstract
Thermophilic, thermotolerant and heat-resistant fungi developed different physiological traits, enabling them to sustain or even flourish under elevated temperatures, which are life-hostile for most other eukaryotes. With the growing demand of heat-stable molecules in biotechnology and industry, the awareness of heat-adapted fungi as a promising source of respective enzymes and biomolecules is still increasing. The aim of this study was to test two different strategies for the efficient isolation and identification of distinctly heat-adapted fungi from easily accessible substrates and locations. Eight compost piles and ten soil sites were sampled in combination with different culture-dependent approaches to describe suitable strategies for the isolation and selection of thermophilous fungi. Additionally, an approach with a heat-shock treatment, but without elevated temperature incubation led to the isolation of heat-resistant mesophilic species. The cultures were identified based on morphology, DNA barcodes, and microsatellite fingerprinting. In total, 191 obtained isolates were assigned to 31 fungal species, from which half are truly thermophilic or thermotolerant, while the other half are heat-resistant fungi. A numerous amount of heat-adapted fungi was isolated from both compost and soil samples, indicating the suitability of the used approaches and that the richness and availability of those organisms in such environments are substantially high.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Fungi can grow in a wide range of conditions and inhabit a multitude of environments. Adaptation to specific environmental conditions can provide a considerable ecological advantage in contrast to less adapted organisms. Ecological niches characterized by extreme abiotic factors can be occupied to avoid competition situations or to gain advantages in habitats with both stable and fluctuating environmental conditions. Based on their growth temperature ranges, fungi are classified as psychrophilic, mesophilic, thermophilic, or thermotolerant. Psychrophily and mesophily, as the abilities to grow at low (under 20 °C) and moderate (20–40 °C) temperatures, respectively, are widespread features among fungi. In contrast, the ability to grow at higher temperatures as thermophilous, i.e., as thermotolerant or as thermophile, is extremely rare. A consistent classification of heat-adaptations in the fungal kingdom is still missing. The most common definition by Cooney and Emerson (1964) classifies fungi with a minimum growth temperature above 20 °C and a maximum growth temperature above 50 °C as thermophilic, and those which can grow between temperatures lower than 20 °C and up to 50 °C as thermotolerant.
Among the 100,000 fungal species currently identified (Kirk et al. 2008), only 75 species are described to be thermophilic (Salar 2018). Additional temperature-adapted features include heat-resistance, which allows usually mesophilic fungi to survive extremely high temperatures (75–100 °C) for a limited amount of time (Samson et al. 2000). Heat-resistance is independent from thermotolerance and thermophily. Although some thermophilous fungi (e.g., Aspergillus fumigatus) are known to be heat-resistant, most heat-resistant fungi are actually mesophilic (e.g., Basidioascus undulatus) (Fergus and Amelung 1971; Ali et al. 2009; Nguyen et al. 2015).
Thermophilous and heat-resistant fungi possess a repertoire of physiological and molecular adaptations to high temperatures. These adaptations include thick sclerotia (Izzo et al. 2006; Samson and Dijksterhuis 2007), unique compositions of lipid membranes (Ianutsevich et al. 2016), synthesis of protective compounds enhancing molecular and cellular stability (Jepsen and Jensen 2004), possibly a rapid transcription and translation machinery (Caspeta et al. 2016), and proteins and enzymes with increased heat stability (McPhillips et al. 2014; Robledo et al. 2016). Moreover, the broad repertoire of secondary metabolite production and heat-stable enzymes makes thermophilous fungi biotechnologically valuable (Busk and Lange 2013; Sharma et al. 2016; Liu et al. 2017). Enzymes derived from thermophilous fungi can be used in paper processing (e.g., xylanases), cosmetics (e.g., lipases), and pharmaceutical (e.g., amidase) (Sevo et al. 2002; Ghatora et al. 2006; Maalej-Achouri et al. 2012; Sen et al. 2016; Rigoldi et al. 2018). Many heat-resistant fungi cause spoilage of heat-processed food products (Scaramuzza and Berni 2014; dos Santos et al. 2018) and produce toxic secondary metabolites (Piecková and Jesenská 1997; Puel et al. 2005). Therefore, besides the valuable aspects for biotechnology, heat-adapted fungi are also relevant for economy and medicine. Cultivation of thermophilic and thermotolerant fungi enables experimental approaches, which enhance the understanding of their physiology, evolutionary adaptations, and possible applications. The cultivation of most thermophilous fungi requires media supplemented with easily accessible carbon sources and temperatures between 35 and 50 °C (Maheshwari et al. 2000; Salar 2018). However, studying these organisms is challenging because their isolation and handling is troublesome.
The main difficulties include fast overgrowth of certain species (Tansey 1971), overall low isolation success (Langarica-Fuentes et al. 2014a), potential pathogenicity (De Gannes et al. 2013; Sandona et al. 2019), and demanding culture maintenance at high temperatures. Since some common fast-growing thermophilous fungi outperform others, the isolation of rare species can be difficult (Kumar and Aneja 1999; Morgenstern et al. 2012). Isolation techniques developed to overcome those problems and to isolate thermophilous fungi from various substrates (see Salar 2018) make use of humidity chambers (Buxton and Mellanby 1934), dilution plating (Apinis 1963), and paired petri dish plating (Cooney and Emerson 1964). Additionally, recent studies performed enrichment cultivation by using selective temperature ranges and distinct carbon sources (Anastasi et al. 2005; Nazir et al. 2007; Langarica-Fuentes et al. 2014a). However, different combinations of these techniques and factors might be necessary to cover the physiological requirements of differently heat-adapted species from various substrates.
Thermophilous fungi often colonize habitats, characterized by temporary or frequent heating or associated with dead plant material. For instance, plant compost, herbivore dung, municipal waste, bird nests, wood chip piles (Korniłłowicz-Kowalska and Kitowski 2013; Ahirwar et al. 2017; Noreen et al. 2019), as well as fermenting foods are known habitats for thermophilous fungi (Zhang et al. 2016). Furthermore, the detection of thermophilous fungi in various types of soil has been often reported (Eggins and Malik 1969; Powell et al. 2012; Ahirwar et al. 2017). As a result of the unstable temperatures, the occurrence of mesophilic fungi, including heat-resistant species, is common in these habitats. Due to their unique adaptation to temperature, heat-resistant fungi have advantages in colonizing habitats with periodically extremely high temperature disturbances, e.g., flare pits, forest fire regions, or areas with slash-and-burn agriculture. Additionally, heat-resistant fungi have been repeatedly detected in heat-sterilized food products, e.g., fruit juices or canned foods (Tournas and Traxler 1994; Kotzekidou 1997), leading to spoilage of such products and economic losses. However, habitats exclusive for one group of these heat-adapted fungi are extremely rare. In compost temperatures fluctuate between ambient temperature and over 70 °C during the different phases of composting (Lechner et al. 2005; Fischer and Glaser 2012; Langarica-Fuentes et al. 2014b). Thus, compost harbors a highly dynamic and rich diversity of heat-adapted and mesophilic saprotrophic species (Langarica-Fuentes et al. 2014b; Galitskaya et al. 2017). Therefore, compost is a promising source of fungal species with biotechnological potential due to the broad repertoire of heat-stable enzymes.
The aim of this study was to develop an approach suitable for the isolation of a broad range of thermotolerant, thermophilic, and heat-resistant fungal species with promising biotechnological application. For this, we sampled common garden composts and soil. For the isolation of thermophilous fungi, the incubations were performed at two distinct temperatures on five different media. Additionally, we applied a heat-shock treatment to select for heat-resistant thermophilous and heat-resistant mesophilic species. Variations of the sampling, substrate-processing and cultivation were applied in order to increase isolation efficiency of heat-adapted fungi.
Material and methods
Study design
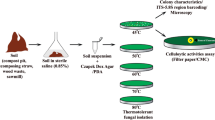
This study aimed for the efficient isolation and cultivation of differently heat-adapted fungi with different enzymatic repertoires and temperature ranges by using two strategies (Table 1, Fig. 1). The heat-adapted fungi were categorized as thermophilic (growing from 20 to > 50 °C) and thermotolerant (growing from < 20 to 50 °C) as defined by Cooney and Emerson (1964) and heat-resistant (resisting 75 °C for 30 min) according to Samson et al. (2000). We collected compost and soil samples from distinct locations and isolated fungi with variations of a flotation-based approach. The first strategy (strategies 1a and 1b), based on the combination of distinct temperature treatments (incubation at 45 °C or 55 °C) and standard (potato dextrose, dextrose) or enrichment media (starch, cellulose, xylan), was used for isolation of thermophilous fungi (i.e., thermophilic and thermotolerant) from compost material. Strategy 1 comprises two samplings, and strategy 1b represents an optimization of strategy 1a, with a reduced carbon source set. A second strategy (strategy 2) was used to isolate thermophilic fungi as well as heat-resistant mesophilic and heat-resistant thermophilic fungi from both compost and soil material. Therefore, an additional heat-shock treatment (30 min, 75 °C) was applied for a subset of samples, followed by an incubation at room temperature (RT) or 55 °C.
Sampling and cultivation strategies for the isolation of thermophilic, heat-tolerant, and heat-resistant fungi. Two different strategies were conducted to isolate heat-adapted fungi from soil and compost piles. Soil substrate was taken from the A-horizon. Compost substrate was sampled from both the upper (surface) and lower (core) layers of the pile. Material was mixed 1:9 with peptone solution, roughly mixed and incubated for 30 min with 300 rpm at 45 °C. Alternatively, for the isolation of heat-resistant species, the samples were treated with a heat-shock at 75 °C for 30 min. Several dilutions (1:10, 1:15, 1:25, 1:30, and undiluted) and five different media (starch, xylan, cellulose, potato dextrose, dextrose) were tested. For strategy 2, only potato dextrose medium was used. The plates were inoculated in triplicates with 150 μl and incubated at 45 °C, 55 °C, or room temperature (RT)
Sampling sites and collection
Eight compost and ten soil samples were collected between September 2018 and March 2019 from two different locations in Bochum, Germany (Online Resource 1). Six domestic compost piles of different age, containing various green waste (e.g., lawn cuttings, hay, vegetable waste, and fruit tree cuttings) were sampled at an allotment garden area (51° 28′ 16.4′′ N 7° 13′ 59.7′′ E). Located in the botanical garden of the Ruhr-University Bochum (51° 26′ 33.6′′ N 7° 16′ 04.3′′ E), two domestic compost piles consisted of botanical waste of both annual and persistent ornamental plants and the ten soil sites were associated to different plant communities (Online Resource 1).
Compost samples were collected from both the upper and the lower layer from a pile to obtain differently matured compost material. A volume of 300 to 400 ml was sieved (mesh size of 1 cm), roughly mixed and stored in plastic bags at RT for up to 24 h until processing. Prior to the sampling of soil material, the overlying surface layer (e.g., litter, grass) was removed. Using a steel cylinder (diameter of 5.5 cm and height of 4 cm) 200 to 300-ml bulk soil of the A-horizon were sampled. Soil in direct association with roots was avoided. Bigger solids and root residues were removed if present. The samples were stored at RT up to 24 h until processing.
Sample treatment and selective culture isolation
The first two sampling approaches applied similar strategies (strategies 1a and 1b), while the third sampling (strategy 2) differed considerably regarding sample types and treatments. For the first two samplings, a subsample of 4 g, while for the third sampling a subsample of 25 g was mixed 1:9 with sterile 0.1% proteose peptone solution (Fig. 1). Briefly, manual shaking was followed by an incubation in a water bath at 45 °C for 30 min to isolate either thermophilous fungi or at 75 °C for 30 min (at 300 rpm) to isolate heat-resistant fungi. Afterwards, the supernatant was combined with 0.1% proteose peptone solution to create differently diluted aliquots of the samples (Fig. 1). The aliquots were vortexed to avoid sedimentation of particles within the suspension and 150 μl each were evenly dispensed on petri dishes with solid growth media. To prevent bacterial growth and decelerate fungal growth speed, rose bengal (Ottow 1972) and chloramphenicol (Hunter et al. 1974) were added to the growth media.
The media contained various carbon-sources (Table 1) to promote growth of fungal species with different nutrition capabilities. Potato dextrose medium contained 39 g/l potato dextrose agar (Carl Roth) according to manufacturer instructions and pH adjusted to 6.2. Dextrose medium contained 10 g/l dextrose (Fisher Scientific), 5 g/l soy peptone, 1 g/l KH2PO4, 0.5 g/l MgSO4, and 15 g/l agar and pH adjusted to 6.2 (see Salar 2018). Starch medium (YpSs) contained 4 g/l yeast extract, 15 g/l starch (Carl Roth), 1 g/l KH2PO4, 0.5 g/l MgSO4*7H2O and 20 g/l agar, and pH adjusted to 6.2 (see Salar 2018). Medium with cellulose as carbon sources consisted of 20 g/l cellulose (Sigma-Aldrich), 1 g/l K3PO4, 0.5 g/l of (NH4)2SO4, 0.5 g/l L-Asparagin, 0.5 g/l KCl and yeast extract, 0.2 g/l MgSO4, 0.1 g/l CaCl2 and 15 g/l agar, and pH adjusted to 6.2 (adapted from the formula provided by Hardy Diagnostics). Xylan medium contained 3 g/l yeast extract, 1.5 g/l peptone, 3.5 g/l NaCl, 1 g/l NaNO3 and KH2PO4 each, 0.3 g/l MgSO4*7H2O, 10 g/l xylan (beechwood xylan, Sigma-Aldrich), and 20 g/l agar and pH adjusted to 6.2 (adapted from Kalim and Ali 2016). Triplicates of each medium were inoculated and incubated at different temperatures (strategies 1a and 1b: 45 °C, 55 °C; strategy 2: RT, 55 °C). To facilitate the growth of mainly thermophilic fungi, 55 °C was used as incubation temperature, while 45 °C was chosen to favor the growth of more thermotolerant species. The growth of mesophilic, but heat-resistant fungi was facilitated by incubation at RT, and the selection for heat-resistant and thermophilic species was achieved by incubating previously heat-shocked samples at 55 °C. The plates incubated at 45 °C or 55 °C were monitored daily for 7 days. Longer incubation of samples at increased temperatures was not feasible, due to desiccation of the agar or overgrowth of the plate, caused by single fungal colonies. Plates incubated at RT were monitored daily for 21 days, giving fungi, whose viability might be decreased due to the heat-shock treatment, enough time to germinate and grow. Colonies with distinct morphology (e.g., growth, colorization, mycelia branching) were sub-cultured and purified from the environmental plates of each sampling site, carbon source, and incubation temperature. Taxonomic assignment was achieved by rDNA barcode sequencing or microsatellite fingerprinting (Online Resource 2).
Identification of thermophilous fungal isolates via ITS-rDNA barcoding and microsatellite fingerprinting
After purification, isolates were categorized based on their morphology into morphotypes. One representative of each morphotype, each substrate, sample, and treatment was chosen for further analysis. Genomic DNA of representatives of each morphotype was extracted by a phenol-chloroform-based approach (Mülhardt 2013). Therefore, a few milligrams of freshly grown mycelia were used. DNA extracts were used for PCR to amplify the fungal barcode ITS-rDNA region (Schoch et al. 2012; Stielow et al. 2015). The fungal specific primer combination ITS1F (Gardes and Bruns 1993) and ITS4 (White et al. 1990) were used. Each PCR reaction had a volume of 12.5 μl and contained 6.25 μl GoTaq G2 Hot Start Colorless Master Mix (Promega), 5.25 μl ddH2O, 0.25 μl of each primer (10 μM), and 0.5 μl of DNA extract. The PCR program was the following: initial denaturation at 95 °C (3 min), 32 cycles of denaturation at 94 °C (27 s), annealing at 57 °C (60 s), and elongation at 72 °C (90 s), followed by final elongation at 72 °C (7 min). If the ITS amplification was not successful or if this barcode was not suitable for identification on species level, other barcodes (LSU, ß-tubulin or calmodulin) were additionally amplified. The primer combination LR0R (Cubeta et al. 1991) and LR6 (Vilgalys and Hester 1990) was used to amplify the adjacent LSU-region (Vu et al. 2019). The used PCR program was initial denaturation for 96 °C (2 min), 35 cycles of denaturation at 96 °C (20 s), annealing at 45 °C (40 s) and elongation at 72 °C (90 s), and a final elongation at 72 °C (10 min). For the Aspergillus species, the primers CF1 and CF4 (Peterson 2004) were used for amplification of the partial calmodulin barcode sequence. The PCR program contained the following steps: initial denaturation at 94 °C (5 min), 35 cycles of denaturation at 94 °C (45 s), annealing at 55 °C (45 s), and elongation at 72 °C (60 s), followed by final elongation at 72 °C (10 min) (Yin et al. 2017). For species identified as Penicillium, primers Bt2a and Bt2b (Glass and Donaldson 1995) were used to amplify the partial ß-tubulin barcode sequence. The PCR started with the initial denaturation at 95 °C (5 min), followed by 5 cycles of denaturation (94 °C, 60 s), annealing (68 °C decreasing 1 °C/cycle, 90 s), and elongation (72 °C, 2 min); afterwards, the PCR proceeded with 25 cycles of denaturation at 94 °C (60 s), annealing at 64 °C (90 s), and elongation at 72 °C (2 min), and a final elongation at 72 °C for 10 min (Frisvad et al. 2004). Amplicons were cleaned with Exonuclease I and Shrimp alkaline phosphatase (NEB) according to manufacturer conditions, but with 1:5 diluted enzyme concentration and sequenced by the sequencing facility of the Ruhr-University Bochum with a capillary sequencer (3130xl Genetic Analyzer, Applied Biosystems). The obtained sequences were checked against the GenBank database (https://blast.ncbi.nlm.nih.gov) with the BLASTn algorithm (Online Resource 2).
Besides sequencing the barcode region of representative isolates, extracted gDNA of the isolates was used for microsatellite-primed PCR (MSP-PCR). This enabled a differentiation of morphologically similar isolates by obtaining a genetic fingerprint for distinct species, without a barcode sequencing of each culture (Supplementary Table S2). Therefore, the (GTG)5 (Lieckfeldt et al. 1993; De Vuyst et al. 2008) primer was used in a PCR reaction containing 6.25 μl GoTaq G2 Hot Start Colorless Master Mix (Promega), 5.5 μl ddH2O, 0.25 μl (GTG)5 (10 μM), and 0.5 μl DNA extract in a total volume of 12.5 μl. The PCR program started with an initial denaturation at 95 °C (2 min), followed by 35 cycles of denaturation at 95 °C (30 s), annealing at 55 °C (40 s), and elongation at 72 °C (90 s), and a final elongation at 72 °C (7 min). The MSP-PCR products were tested by agarose gel electrophoresis with standardized conditions.
Availability of data and material
Sequences of taxonomically assigned isolates were deposited at the European Nucleotide Archive (ENA) database under accession numbers LR881294–LR881364, LR881366–LR881403, LR881444–LR881452, and LR993201–LR993205, LR993215–LR993238 (Online Resource 2). Representative isolates of each species were deposited at the DSMZ culture collection (Online Resource 2).
Results
Isolation efficiency of different isolation and cultivation strategies
For the first sampling (strategy 1a), three compost piles were sampled (Table 1). The processed substrate was incubated for 1 week at two temperatures on five media, on a total of 120 agar plates (Fig. 1). After 3 days of incubation, a variety of fungi was visible on most plates. Purification of morphologically distinct colonies led to 30 single cultures, which were taxonomically assigned to 13 different species (Table 1) based on molecular barcoding. For the second sampling (strategy 1b), 96 agar plates were inoculated with compost substrate from three different piles. For strategy 1b, similar conditions were applied as in strategy 1a, but only four media were used (Table 1). From this sampling, 46 morphologically different cultures were purified and assigned to 12 distinct species. In total, 16 different species were isolated by using the two complementary strategies (1a and 1b), with 9 species being isolated by both. Four and three species were only isolated with strategy 1a and strategy 1b, respectively. Strategy 2 (third sampling) included only one medium and an additional heat-shock treatment (Table 1, Fig. 1). From this, 115 cultures were purified from 216 agar plates. These cultures were taxonomically assigned to 19 different species (Table 1), of which 15 were exclusively isolated with this strategy. Using strategy 2, eight species were isolated from compost substrate, while 4 of those were only isolated in this strategy and by using the heat-shock treatment.
The molecular identification of the isolates based on barcode sequences resulted in a taxonomic assignment on species level for 179 isolates. The identification of 12 isolates was only possible on the genus level (Fibulochlamys sp., Lichtheimia sp., and Triangularia sp.) due to low similarity with reference material (Online resource 2), which might, therefore, represent novel-undescribed species. Besides that, intraspecific diversity has also been indicated by microsatellite fingerprinting. For 9 species or genera (e.g., R. emersonii, T. aurantiacus), the microsatellite fingerprints of several isolates displayed different patterns intraspecifically, which could indicate the isolation of multiple strains or lineages of the same species. However, isolates assigned to ten species or genera showed identical microsatellite fingerprints (Online resource 2).
Diversity of thermophilous and heat-resistant fungi in compost and soil
Within this study, we obtained 191 fungal isolates from compost (92 isolates) and soil (99 isolates) samples, belonging to 31 different species (Table 2). The main proportion of isolates was assigned to Ascomycota with 23 species belonging mainly to the orders Eurotiales and Sordariales. The Mucoromycetes was the second most common phylum, represented by seven species of the orders Mucorales and Mortierellales, followed by Basidiomycota with one species of the order Agaricales. Aspergillus lacinosus, A. fumigatus, Chaetomium thermophilum, Rasamsonia emersonii, Thermoascus aurantiacus, Triangularia sp., and Trichocladium pyriforme were isolated from both substrates. In contrast, 14 species were detected only in one sample, e.g., Malbranchea cinnamomea, Thermomyces lanuginosus, Fibulochlamys sp., Rasamsonia byssochlamydoides, and R. composticola.
Diversity in compost
From compost, 20 species were isolated of which 13 were assigned to Ascomycota and 7 species to Mucoromycota. On average, six species were isolated from each compost pile. A. fumigatus (87.5%), C. thermophilum (62.5%), and T. aurantiacus, Rhizomucor pusillus, Rhizopus microsporus, and A. spinulosporus (all 50%) showed the highest occurrence in compost piles. Ten species were isolated with an incubation temperature of 55 °C and can therefore be considered as thermophilic fungi, while 8 species were only isolated at 45 °C (strategy 1) and are rather thermotolerant. Two thermophilic, three mesophilic fungi, and A. fumigatus were isolated in combination with the heat-shock approach (strategy 2), resulting in the isolation of 6 heat-resistant species from compost.
Diversity in soil
Isolates obtained from soil substrate, were assigned to 18 species, of which 17 belonged to Ascomycota and one species to Basidiomycota. On average, five species were isolated from each soil sample. The majority of species (14) were isolated with a heat-shock treatment in combination with incubation at room temperature (strategy 2) and can therefore be considered as mesophilic heat-resistant fungi. Although an incubation temperature of 55 °C was used as in previous samplings, only 4 species were isolated from soil. The highest occurrence in soil samples was observed for the thermophilic R. emersonii and T. aurantiacus (both 80%) and for the mesophilic heat-resistant A. nishimurae (60%).
Isolation efficiency of different carbon sources
The isolation success of species differed among different media. Less complex carbon sources (potato dextrose, dextrose medium) showed increased fungal growth. This resulted in quick overgrowth, hampering the purification of species. However, 29 species were isolated using potato dextrose medium, while seven and six species were isolated from either starch and xylan media and seven species from cellulose-containing medium. Although, ten species were obtained from multiple media, 22 species were isolated from only one carbon source (Table 2). Aspergillus fumigatus was the only species isolated from all media. Notably, the application of the non-standard media containing xylan, cellulose, and starch led to the isolation of biotechnologically relevant species, e.g., Malbranchea cinnamomea, Thermothelomyces heterothallicus, and Rasamsonia emersonii.
Effect of temperature for the isolation of heat-adapted fungi
Application of different incubation temperatures resulted in diverse fungal colonies with different growth rates. Plates incubated at 45 °C showed quick fungal overgrowth, which was a limiting factor for the isolation success. Incubations at 55 °C displayed no overgrowth, and plates were colonized by a reduced number of fungal colonies.
Different fungal species were obtained from each incubation temperature. Chaetomium thermophilum, R. pusillus, and A. fumigatus are the only species obtained from both 45 °C and 55 °C. Incubation at 45 °C resulted in nine isolated species (Table 2), including most of the isolated Mucoromycota (e.g., Lichtheimia spp., Rhizopus microsporus, Mortierella wolfii, Rhizomucor pusillus). Thirteen species were isolated at 55 °C (Table 2), including the biotechnologically important species Malbranchea cinnamomea, Thermothelomyces heterothallicus, Rasamsonia emersonii, Thermoascus aurantiacus, and Thermomyces lanuginosus.
The combination of a heat-shock treatment at 75 °C with incubation at 55 °C resulted in the isolation of Rasamsonia byssochlamydoides, R. composticola, R. emersonii, and Thermoascus aurantiacus. From the combination of the heat-shock with incubation at RT, 14 fungal species were obtained (Table 2), which were not isolated from the previous approaches (e.g., various members of the genus Aspergillus).
Discussion
Isolation of thermophilous species from compost and soil
A total amount of 31 fungal species were isolated from compost (20 isolates) and soil (18 isolates) samples, from which 18 can be considered as thermophilous (i.e., thermotolerant, or thermophilic) and 13 as mesophilic heat-resistant species (Table 2). The number of isolated thermophilous fungal species from compost or soil was higher than in recent studies (Chadha et al. 2004; Lee et al. 2014; Langarica-Fuentes et al. 2014a, 2015). Distinct temperature treatments, the use of two different substrates and multiple sampling sites, as well as the variation of carbon sources (for compost samples) was shown to be an efficient combination of factors to isolate heat-adapted fungi. For instance, five species were only obtained from one carbon source within the first two complementary samplings of compost (strategies 1a and 1b) when multiple carbon sources were applied (Table 2). In contrast, sampling 3 (strategy 2) used only one medium (PD), but both compost and soil substrates were sampled to investigate the potential of soil as a substrate for isolating thermophilic fungi instead of the influence of used cultivation media. The additional heat-shock treatment in strategy 2 seemed to have influenced the outcome of the isolation of thermophiles. R. byssochlamydoides and R. composticola were only isolated from samples heat-shocked before incubation at increased temperatures, while C. thermophilum (9 isolates from 4 different locations) was solely isolated without the heat-shock. Moreover, in this study, six and five species were only isolated at 45 °C and 55 °C, respectively (Online Resource 2). This is in accordance with other studies, in which different temperatures (strategy 1 and strategy 2) and carbon sources (strategies 1a and 1b) led to the isolation of different fungal species (Tansey 1971; Anastasi et al. 2005; Langarica-Fuentes et al. 2014a; Birajdar et al. 2020). Furthermore, we isolated thirteen species only from a single sampling site (7 species from soil and 6 species from compost), indicating the suitability of distinct sampling locations. Strategy 1 led to the isolation of five of those species, while eight were obtained with strategy 2 (Online Resource 2).
In our study, each variation of the sampling and isolation process seemed to affect the outcome of the applied strategy. Therefore, we consider the combination of strategies including different parameters to be suitable to isolate a high diversity of heat-adapted fungi. The usage of multiple media (strategies 1a and 1b) is assumed to help isolating fungi with most probably different nutritional requirements. However, the isolation success was influenced more by temperature variations (strategy 1: 45 °C, 55 °C; strategy 2: 55 °C, heat-shock, RT) and the sampling of different locations. The inclusion of an additional substrate in strategy 2 also slightly increased the isolation success of thermophilic fungi, since one species (R. byssochlamydoides) was only isolated from soil.
Thermophilous fungi have specific physiological and ecological requirements; however, different species in this group display different adaptations to temperature ranges and carbon sources (Powell et al. 2012; Morgenstern et al. 2012; Thanh et al. 2019). This diversity allows thermophilous fungi to colonize several niches and sub-habitats (Ahirwar et al. 2017; Salar 2018). Compost provides very heterogenous sub-habitats that are dependent on the feedstock plant material (Neher et al. 2013) and the composting phase and process (Galitskaya et al. 2017). These factors shape fungal community composition and influence community succession (Langarica-Fuentes et al. 2014b; Meng et al. 2019; Jiang et al. 2020). Therefore, for the isolation of differently heat-adapted fungal species, it is necessary to consider and mimic this environmental niche diversity through variations in the sample collection, processing, and cultivation. For the isolation of thermotolerant and thermophilic species, the use of distinct temperature regimes (e.g., 45 °C, 55 °C) is advantageous. Cultivation between 40 and 45 °C rather promotes growth of thermotolerant fungi and prevents growth of mesophiles (Dix and Webster 1995; Houbraken et al. 2012). In order to promote the growth of thermophiles, incubation temperatures above 50 °C are recommended, since many thermotolerant species show minor or no growth at these temperatures (Maheshwari et al. 2000; Morgenstern et al. 2012). An incubation temperature of 50 °C could provide suitable temperature conditions for either very thermotolerant or less thermophilic fungi. However, incubation temperatures at 45 °C and 55 °C facilitate isolation of thermotolerant and thermophilic fungi, respectively. The thermophilic species Chaetomium thermophilum, Malbranchea cinnamomea, as well as Thermoascus aurantiacus grow at 55 °C (Morgenstern et al. 2012, Table 2), but might display different fitness on distinct carbon sources. These differences in the efficiency to breakdown different carbon sources, were especially useful to prevent the overgrowth of Aspergillus fumigatus, which is a fast-growing fungus with a broad range of growth conditions (Jensen 1931; Kozakiewicz and Smith 1994; Tekaia and Latgé 2005; Rhodes 2006). In this study, Aspergillus fumigatus was isolated from all used carbon sources (strategies 1a, 1b, and 2), but showed reduced growth on cellulose and xylan. Additionally, the usage of cellulose led to the isolation of a novel Lichtheimia species (Table 2, Online resource 2). This demonstrates that the combination of different temperatures with different carbon sources, further facilitates the selection of different species.
Previous studies attempted to isolate thermophilous fungi by performing increased sampling depth of several substrates and intensive cultivation approaches (Nazir et al. 2007; Rajavaram et al. 2010; Ahirwar et al. 2017). For instance, Ahirwar et al. (2017) isolated 19 thermophilous fungal species from 79 samples of eight different habitats. However, only seven different species were isolated from 12 compost samples. In our study, by combining different cultivation conditions (Table 1), we were able to obtain a higher yield from a smaller sampling size (eight compost samples). Seventeen thermophilic or thermotolerant and additionally three heat-resistant fungal species were isolated from compost. Sixteen of those thermophilous species were isolated with strategy 1 (1a and 1b) from six different composts, while eight species (4 thermophilic, 3 heat-resistant, 1 heat-resistant thermophilic) were isolated with strategy 2 from only two composts. From soil substrates, previous studies isolated 12 thermophilous fungal species from 46 soil samples from three geothermal sites in China (Pan et al. 2010), while 10 thermotolerant and six thermophilic species were isolated from 40 soil samples from four different sites in India (Salar and Aneja 2006). By applying a restrictive approach (strategy 2, Fig. 1), we were able to isolate five thermophilic species from 10 soil samples (Table 2). Despite the small sampling size and the restrictive cultivation approach, we were able to isolate thermophilic fungi with a high rate of success.
It is assumed that thermophilous fungi growing in suitable conditions, disperse by aerosols to reach new habitats (Le Goff et al. 2010). After sedimentation in a new environment (e.g., soil), they remain in a dormant stage until favorable conditions are present. This explains their occurrence in non-favorable habitats without elevated temperatures, which are mainly dominated by mesophilic fungi. The growth of thermophilous fungi in soil is assumed to be due to temporary sun-heating (Tansey and Jack 1976). However, molecular data is still missing to verify their activity in soil. The application of cultivation-independent approaches based on RNA might be used to provide insights into the ecology of thermophilous fungi.
Selective isolation of heat-resistant species
Most studies isolate heat-resistant species from soil substrates and heat-processed food products (Frąc et al. 2015). So far, only few studies tested for heat-resistance of fungi isolated from compost (e.g., Fergus and Amelung 1971). To our knowledge, our study is the first in attempting to selectively isolate heat-resistant fungi from both soil and compost with a heat-shock approach. Previous studies attempting to isolate heat-resistant fungi from soil were able to obtain between six and 16 species (Okagbue 1989; Jesenská et al. 1992; Horie et al. 2003; Ali et al. 2009; Sezen and Demirel 2019). In this study, we isolated 18 heat-resistant fungal species, of which four species (Rasamsonia byssochlamydoides, R. composticola, Curvularia buchloes, and Trichocladium pyriforme) and two novel taxa (Fibulochlamys sp. and Triangularia sp.) are here reported for the first time as heat-resistant. Most of the heat-resistant species (11) were isolated exclusively from soil, while only Rasamsonia composticola was exclusively found in compost. Notably, Rasamsonia composticola was here found for the first time since its original isolation from compost (Su and Cai 2013). Furthermore, a third member of the undersampled Fibulochlamys genus (Mahajan et al. 2016) was isolated. The most frequently isolated heat-resistant species belong to the genera Aspergillus and Thermoascus, which is supported by previous studies. Members of the genus Aspergillus are among the most frequently isolated heat-resistant species from soil (Jesenská et al. 1993; Jesenská and Piecková 1995; Ali et al. 2009), while the genus Thermoascus comprises mainly soil fungi, which produce heat-resistant spores able to survive up to 90 °C (King et al. 1969; Hosoya et al. 2014; Scaramuzza and Berni 2014). Furthermore, we successfully isolated Rasamsonia emersonii, Devriesia thermodurans, Paecilomyces niveus, and Penicillium species, which were previously obtained from soil using a heat-treatment (Jesenská et al. 1992; Jesenská et al. 1993; Ali et al. 2009; Seifert et al. 2004). Usually, heat-resistance of fungi underlies the production of heat-resistant ascospores, which are described for many species isolated in this study, e.g., many Aspergillus species, T. aurantiacus, R. emersonii, R. byssochlamydoides, R. composticola, and Paecilomyces niveus (Houbraken et al. 2012; Su and Cai 2013; Berni et al. 2017; Biango-Daniels et al. 2019). Notably, several of the genera isolated in this study include species known to contaminate various heat-processed food products with their heat-resistant ascospores. For instance, species belonging to the genus Thermoascus are detected in processed tea and fruit juices (Hosoya et al. 2014). Members of Byssochlamys (anamorph: Paecilomyces), Neosartorya (anamorph: Aspergillus), and Talaromyces are among the commonly isolated molds causing spoilage of raw foods, juices, heat-processed cheese, canned fruits and pasteurized foods (Tournas 1994; Kotzekidou 1997; Pitt and Hocking 2009; Frąc et al. 2015; Tranquillini et al. 2017; dos Santos et al. 2018). Therefore, several studies investigated heat-resistance of ascospores under conditions, used in processing and sterilization in food industries (Piecková and Samson 2000; Salomão et al. 2014). However, ascospores are not reported for all species isolated with the heat-shock approach in this study, but heat-resistant structures other than ascospores have already been described for several fungi (Samson and Dijksterhuis 2007). For instance, Devriesia thermodurans produces heat-resistant chlamydospores, requiring a heat activation prior germination (Seifert et al. 2004). A Fibulochlamys species is known to produce thick-walled conidia which might cause a certain heat-resistance (Madrid et al. 2010) and although for T. pyriforme, no information about ascospores is available, for several Trichocladium species, heat-resistant ascospores or chlamydospores have been reported (Wang et al. 2019). Our results suggest that both compost and soil harbor a promising diversity of heat-resistant fungi. Since differently adapted fungal species were detected in these habitats, we hypothesize that a higher diversity of heat-resistant fungi is present in habitats not considered so far. Therefore, further studies should analyze the ecology and diversity of heat-resistant fungi across habitats.
Strategies to increase isolation success
The adaptation to distinct temperature regimes by specific species is useable as an initial and powerful tool for selective approaches to isolate differently heat-adapted species. High temperatures (50–60 °C) facilitate the isolation success of truly thermophilic fungi, while moderately increased temperatures (40–45 °C) lead to the isolation of mainly thermotolerant species (Maheshwari et al. 2000). Additionally, in combination with a heat-shock treatment (strategy 2), the isolation success can be selective towards heat-resistant species. The overlapping growth temperatures of many thermophilic and thermotolerant species (Morgenstern et al. 2012) make additional levels of selection necessary to avoid overgrowth of species and increase the isolation success. The usage of multiple carbon sources, with varying complexity, in combination with different temperature conditions (strategy 1), diversifies the cultivation, which influences the isolation success based on the different enzymatic properties of the fungal species. Most heat-adapted species will grow on rich media with easily accessible carbon sources (e.g., potato dextrose). Aiming for the isolation of a high diversity of species, usage of such media might be sufficient, when enough technical replicates are prepared. However, the isolation strategy can be adapted to the aim of the respective study. Aiming for the isolation of rare or slow growing species, or specialists regarding the utilization of different carbon sources, a diversified approach with multiple media might be preferred (strategies 1a and 1b). Sampling of various habitats and substrates (strategy 2) can increase the number of isolated species, if suitable cultivation conditions are applied (Ahirwar et al. 2017). Furthermore, dilution plating and addition of growth inhibitors (strategies 1 and 2) into the cultivation medium is useful to reduce overgrowth of single species. Categorization into morphotypes serves as an additional level of selection, and the application of microsatellite fingerprinting simplifies the identification and differentiation of isolates on a genetic level, allowing detection of intraspecific diversity. The combination of these screening approaches reduces workload, handling time, and associated costs. The presented effective and time-efficient approaches to isolate differently heat-adapted fungi benefit from the combination of parallelized enrichment cultivation using different incubation temperatures and carbon sources.
In this study, we provide with strategy 1 (1a and 1b) an example for an efficient and convenient approach to isolate both thermophilic and thermotolerant fungi, occupying different ecological niches, from the same substrate. With strategy 2, we were able to create a complementary approach, aiming for both the selective isolation of truly thermophilic fungi and additionally the isolation of heat-resistant thermophilic and mesophilic fungi from two substrates (Fig. 1). These two strategies allowed us to isolate 31 heat-adapted fungal species, including three novel undescribed fungal species. Thus, we exploited differences among and within at least three ecophysiological groups to efficiently isolate differently heat-adapted fungi (Table 2), by applying two distinct approaches. Such approaches can consist of multiple, parallelized selective parameters. This facilitates growth of a broader range of species and reduces overgrowth and loss of potentially slow growing isolates, which leads to the isolation of a higher diversity of heat-adapted fungi, including rare species. The high demand for thermostable enzymes and biomolecules in biotechnology is increasing in the last decades. Although enzyme engineering is expanding quickly (Chowdhury and Maranas 2019), a highly diverse repertoire of natural enzymes is still required. Since fungi are able to breakdown a broad range of recalcitrant compounds, thermophilic fungi are a promising source of enzymes. Currently, only around 75 thermophilic fungal species are known. Therefore, isolation of thermophiles and heat-resistant fungi provides a great opportunity to identify and characterize species with biotechnological potential. Cultivation-independent approaches are useful to characterize microbial community structure, composition, and diversity in the environment. However, for experimental approaches and downstream applications, the isolation of heat-adapted fungi from environmental samples is essential.
Data availability and materials availability
All data and materials generated during this study are included in this manuscript, on its supplementary information or publicly available on databases, or microbial culture collections.
Code availability
Not applicable.
References
Ahirwar S, Soni H, Prajapati BP, Kango N (2017) Isolation and screening of thermophilic and thermotolerant fungi for production of hemicellulases from heated environments. Mycol 8:125–134. https://doi.org/10.1080/21501203.2017.1337657
Ali M, Moghazy S, Shaban G, El-Sababty Z (2009) Heat-resistant fungi isolated from soil in Minia Governorate. Assiut Univ J Bot 38:93–106
Anastasi A, Varese GC, Filipello Marchisio V (2005) Isolation and identification of fungal communities in compost and vermicompost. Mycologia 97:33–44. https://doi.org/10.1080/15572536.2006.118328362006.11832836
Apinis AE (1963) Thermophilous fungi of coastal grasslands. In: Doeksen J, Van der Drift J (eds) Soil Organisms, North-Holland Publishing Co, Amsterdam, pp 427–438
Balajee SA, Gribskov J, Brandt M, Ito J, Fothergill A, Marr KA (2005) Mistaken identity. Neosartorya pseudofischeri and its anamorph masquerading as Aspergillus fumigatus. J Clin Microbiol 43:5996–5999. https://doi.org/10.1128/JCM.43.12.5996-5999.2005
Basotra N, Dhiman SS, Agrawal D, Sani RK, Tsang A, Chadha BS (2019) Characterization of a novel lytic polysaccharide monooxygenase from Malbranchea cinnamomea exhibiting dual catalytic behavior. Carbohydr Res 478:46–53. https://doi.org/10.1016/j.carres.2019.04.006
Berni E, Tranquillini R, Scaramuzza N, Brutti A, Bernini V (2017) Aspergilli with Neosartorya-type ascospores. Heat resistance and effect of sugar concentration on growth and spoilage incidence in berry products. Int J Food Microbiol 258:81–88. https://doi.org/10.1016/j.ijfoodmicro.2017.07.008
Biango-Daniels MN, Snyder AB, Worobo RW, Hodge KT (2019) Fruit infected with Paecilomyces niveus. A source of spoilage inoculum and patulin in apple juice concentrate. Food Control 97:81–86. https://doi.org/10.1016/j.foodcont.2018.10.020
Birajdar GM, Kumbhar VR, Kadam KK, Bhale UN (2020) Occurrence of thermophilic fungal communities and its growth rate on different media and temperatures from available natural substrates. Plant Sci Today 7:172–177. https://doi.org/10.14719/pst.2020.7.2.719
Busk PK, Lange L (2013) Cellulolytic potential of thermophilic species from four fungal orders. AMB Express 3:47. https://doi.org/10.1186/2191-0855-3-47
Buxton PA, Mellanby K (1934) The measurement and control of humidity. Bull Entomol Res 25:171–175. https://doi.org/10.1017/S0007485300012608
Caspeta L, Chen Y, Nielsen J (2016) Thermotolerant yeasts selected by adaptive evolution express heat stress response at 30 °C. Sci Rep 6:27003. https://doi.org/10.1038/srep27003
Chadha BS, Harmeet G, Mandeep M, Saini HS, Singh N (2004) Phytase production by the thermophilic fungus Rhizomucor pusillus. World J Microbiol Biotechnol 20:105–109. https://doi.org/10.1023/B:WIBI.0000013319.13348.0a
Cheng VCC, Chan JFW, Ngan AHY, To KKW, Leung SY, Tsoi HW, Yam WC, Tai JWM, Wong SSY, Tse H, Li IWS, Lau SKP, Woo PCY, Leung AYH, Lie AKW, Liang RHS, Que TL, Ho PL, Yuen KY (2009) Outbreak of intestinal infection due to Rhizopus microsporus. J Clin Microbiol 47:2834–2843. https://doi.org/10.1128/JCM.00908-09
Chowdhury R, Maranas CD (2019) From directed evolution to computational enzyme engineering - A review. AIChE J 66(3):e16847. https://doi.org/10.1002/aic.16847
Cooney DG, Emerson R (1964) Thermophilic fungi (Vol. 27). WH Freeman, San Francisco
Corbel MJ, Eades SM (1991) Observations on the experimental pathogenicity and toxigenicity of Mortierella wolfii strains of bovine origin. Br Vet J 147:504–516. https://doi.org/10.1016/0007-1935(91)90020-N
Cubeta MA, Echandi E, Abernethy T, Vilgalys R (1991) Characterization of anastomosis groups of binucleate rhizoctonia species using restriction analysis of an amplified ribosomal RNA gene. Phytopathol 81:1395. https://doi.org/10.1094/Phyto-81-1395
Damásio ARL, Maller A, Silva TM, Jorge JA, Terenzi HF, Polizeli MLTM (2011) Biotechnological potential of alternative carbon sources for production of pectinases by Rhizopus microsporus var. rhizopodiformis. Braz Arch Biol Technol 54:141–148. https://doi.org/10.1590/S1516-89132011000100019
de Gannes V, Eudoxie G, Hickey WJ (2013) Insights into fungal communities in composts revealed by 454-pyrosequencing. Implications for human health and safety. Front Microbiol 4:164
de Oliveira APA, Silvestre MA, Garcia NFL, Alves-Prado HF, Rodrigues A, da Paz MF, Fonseca GG, Leite RSR (2016) Production and catalytic properties of amylases from Lichtheimia ramosa and Thermoascus aurantiacus by solid-state fermentation. Sci World J 2016:7323875. https://doi.org/10.1155/2016/7323875
de Vuyst L, Camu N, de Winter T, Vandemeulebroecke K, van de Perre V, Vancanneyt M, de Vos P, Cleenwerck I (2008) Validation of the (GTG)5-rep-PCR fingerprinting technique for rapid classification and identification of acetic acid bacteria, with a focus on isolates from Ghanaian fermented cocoa beans. Int J Food Microbiol 125:79–90. https://doi.org/10.1016/j.ijfoodmicro.2007.02.030
Dien Bard J, Mangahis A, Hofstra TC, Bender JM (2014) First case report of bloodstream infection by Rhizomucor pusillus in a child with hemophagocytic lymphohistiocytosis. Med Mycol Case Rep 5:20–23. https://doi.org/10.1016/j.mmcr.2014.05.002
Dix NJ, Webster J (1995) Fungal Ecology. Springer Netherlands, Dordrecht
dos Santos JLP, Samapundo S, Biyikli A, van Impe J, Biyikli A, Akkermans S, Höfte M, Abatih EN, Sant'Ana AS, Devlieghere F (2018) Occurrence, distribution and contamination levels of heat-resistant moulds throughout the processing of pasteurized high-acid fruit products. Int J Food Microbiol 281:72–81. https://doi.org/10.1016/j.ijfoodmicro.2018.05.019
Dwyer K (2019) A study of selected xylanolytic and chitinolytic enzymes from Rasamsonia emersonii, and their potential application in the valorisation of mushroom-production waste streams. Dissertation, University of Limerick
Eggins HOW, Malik KA (1969) The occurrence of thermophilic cellulolytic fungi in a pasture land soil. Antonie van Leeuwenhoek 35:178–184. https://doi.org/10.1007/BF02219128
Engel G, Teuber M (1991) Heat resistance of ascospores of Byssochlamys nivea in milk and cream. Int J Food Microbiol 12:225–233. https://doi.org/10.1016/0168-1605(91)90073-X
Fergus CL, Amelung RM (1971) The heat resistance of some thermophilic fungi on mushroom compost. Mycologia 63:675–679. https://doi.org/10.2307/3757572
Ferreira EHR, Rosenthal A, Calado V, Saraiva J, Mendo S (2009) Byssochlamys nivea inactivation in pineapple juice and nectar using high pressure cycles. J Food Eng 95:664–669. https://doi.org/10.1016/j.jfoodeng.2009.06.053
Fischer D, Glaser B (2012) Synergisms between compost and biochar for sustainable soil amelioration. In: Chindo PS, Bello LY, Kumar N (eds) Utilization of organic wastes for the management of phyto-parasitic nematodes in developing economies. INTECH Open Access Publisher. https://doi.org/10.5772/31200
Frąc M, Jezierska-Tys S, Yaguchi T (2015) Occurrence, detection, and molecular and metabolic characterization of heat-resistant fungi in soils and plants and their risk to human health. Adv Agron 132: 161–204. https://doi.org/10.1016/bs.agron.2015.02.003
Frisvad JC, Frank JM, Houbraken JAMP, Kuijpers AFA, Samson RA (2004) New ochratoxin A producing species of Aspergillus section Circumdati. Stud Mycol:23–43
Galitskaya P, Biktasheva L, Saveliev A, Grigoryeva T, Boulygina E, Selivanovskaya S (2017) Fungal and bacterial successions in the process of co-composting of organic wastes as revealed by 454 pyrosequencing. PloS one 12:e0186051. https://doi.org/10.1371/journal.pone.0186051
Gardes M, Bruns TD (1993) ITS primers with enhanced specificity for basidiomycetes - application to the identification of mycorrhizae and rusts. Mol Ecol 2:113–118. https://doi.org/10.1111/j.1365-294X.1993.tb00005.x
Ghatora SK, Chadha BS, Badhan AK, Saini HS, Bhat MK (2006) Identification and characterization of diverse xylanases from thermophilic and thermotolerant fungi. BioResour 1(1):18–33
Glass NL, Donaldson GC (1995) Development of primer sets designed for use with the PCR to amplify conserved genes from filamentous ascomycetes. Appl Environ Microbiol 61:1323–1330
Gomes MZR, Lewis RE, Kontoyiannis DP (2011) Mucormycosis caused by unusual mucormycetes, non-Rhizopus, -Mucor, and -Lichtheimia species. Clin Microbiol Rev 24:411–445. https://doi.org/10.1128/CMR.00056-10
Gonçalves FA, Leite RSR, Rodrigues A, Argandoña EJS, Fonseca GG (2013) Isolation, identification and characterization of a novel high level β-glucosidase-producing Lichtheimia ramosa strain. Biocatal Agric Biotechnol 2:377–384. https://doi.org/10.1016/j.bcab.2013.06.006
Hasanin MS, Hashem AH, Abd El-Sayed ES, El-Saied H (2020) Green ecofriendly bio-deinking of mixed office waste paper using various enzymes from Rhizopus microsporus AH3. Efficiency and characteristics. Cellul 27:4443–4453. https://doi.org/10.1007/s10570-020-03071-3
Hogaboam CM, Blease K, Mehrad B, Steinhauser ML, Standiford TJ, Kunkel SL, Lukacs NW (2000) Chronic airway hyperreactivity, goblet cell hyperplasia, and peribronchial fibrosis during allergic airway disease induced by Aspergillus fumigatus. Am J Pathol 156(2):723–732. https://doi.org/10.1016/S0002-9440(10)64775-X
Horie Y, Abliz P, Fukushima K, Okada K, Takaki GC (2003) Two new species of Neosartorya from Amazonian soil, Brazil. Mycosci 44:397–402. https://doi.org/10.1007/S10267-003-0132-1
Hosoe T, Mori N, Kamano K, Itabashi T, Yaguchi T, Kawai K (2011) A new antifungal yellow pigment from Aspergillus nishimurae. J Antibiot 64:211–212. https://doi.org/10.1038/ja.2010.132
Hosoya K, Nakayama M, Tomiyama D, Matsuzawa T, Imanishi Y, Ueda S, Yaguchi T (2014) Risk analysis and rapid detection of the genus Thermoascus, food spoilage fungi. Food Control 41:7–12. https://doi.org/10.1016/j.foodcont.2013.12.021
Houbraken J, Spierenburg H, Frisvad JC (2012) Rasamsonia, a new genus comprising thermotolerant and thermophilic Talaromyces and Geosmithia species. Antonie van Leeuwenhoek 101:403–421. https://doi.org/10.1007/s10482-011-9647-1
Hua C, Li W, Han W, Wang Q, Bi P, Han C, Zhu L (2018) Characterization of a novel thermostable GH7 endoglucanase from Chaetomium thermophilum capable of xylan hydrolysis. Int J Biol Macromol 117:342–349. https://doi.org/10.1016/j.ijbiomac.2018.05.189
Hunter BB, Hoyes JV, Barnett HL (1974) The additions of aureomycin and chloramphenicol to various fungal agar media to prevent bacterial contamination. Proc Pa Acad Sci 48:88–92
Hüttner S, Granchi Z, Nguyen TT, van Pelt S, Larsbrink J, Thanh VN, Olsson L (2018) Genome sequence of Rhizomucor pusillus FCH 5.7, a thermophilic zygomycete involved in plant biomass degradation harbouring putative GH9 endoglucanases. Biotechnol Rep 20:e00279. https://doi.org/10.1016/j.btre.2018.e00279
Ianutsevich EA, Danilova OA, Groza NV, Kotlova ER, Tereshina VM (2016) Heat shock response of thermophilic fungi. Membrane lipids and soluble carbohydrates under elevated temperatures. Microbiol 162:989–999. https://doi.org/10.1099/mic.0.000279
Izzo A, Nguyen DT, Bruns TD (2006) Spatial structure and richness of ectomycorrhizal fungi colonizing bioassay seedlings from resistant propagules in a Sierra Nevada forest. Comparisons using two hosts that exhibit different seedling establishment patterns. Mycol 98:374–383. https://doi.org/10.1080/15572536.2006.11832672
Jayasuriya H, Zink D, Basilio A, Vicente F, Collado J, Bills G, Goldman ML, Motyl M, Huber J, Dezeny G, Byrne K, Singh SB (2009) Discovery and antibacterial activity of glabramycin A-C from Neosartorya glabra by an antisense strategy. J Antibio 62:265–269. https://doi.org/10.1038/ja.2009.26
Jensen HL (1931) The fungus flora of the soil. Soil Science 31(2):123
Jepsen HF, Jensen B (2004) Accumulation of trehalose in the thermophilic fungus Chaetomium thermophilum var. coprophilum in response to heat or salt stress. Soil Biol Biochem 36:1669–1674. https://doi.org/10.1016/j.soilbio.2004.07.010
Jesenská Z, Piecková E (1995) Heat-resistant fungi. Czech Mycol. 48:73–76. https://doi.org/10.33585/cmy.48110
Jesenská Z, Piecková E, Bernát D (1992) Heat-resistant fungi in the soil. Int J Food Microbiol 16:209–214. https://doi.org/10.1016/0168-1605(92)90081-D
Jesenská Z, Piecková E, Bernát D (1993) Heat resistance of fungi from soil. Int J Food Microbiol 19:187–192. https://doi.org/10.1016/0168-1605(93)90076-S
Jiang X, Deng L, Meng Q, Sun Y, Han Y, Wu X, Sheng S, Zhu H, Ayodeji B, Egbeagu UU, Xu X (2020) Fungal community succession under influence of biochar in cow manure composting. Env Sci Pollut Res Int 27:9658–9668. https://doi.org/10.1007/s11356-019-07529-1
Kalim B, Ali NM (2016) Optimization of fermentation media and growth conditions for microbial xylanase production. 3 Biotech 6:122. https://doi.org/10.1007/s13205-016-0445-3
Khan FI, Bisetty K, Singh S, Permaul K, Hassan MI (2015) Chitinase from Thermomyces lanuginosus SSBP and its biotechnological applications. Extrem: Life Extrem Cond 19:1055–1066. https://doi.org/10.1007/s00792-015-0792-8
Kimura M, Udagawa S, Makimura K, Satoh K, Toyazaki N, Ito H (2009) Isolation and identification of Rhizomucor pusillus from pleural zygomycosis in an immunocompetent patient. Med Mycol 47(8):869–873. https://doi.org/10.3109/13693780903059485
King AD, Michener HD, Ito KA (1969) Control of Byssochlamys and related heat-resistant fungi in grape products. Appl Env Microbiol 18(2):166–173
Kirk PM, Cannon PF, Minter DW, Stalpers JA (2008) Dictionary of the fungi. CAB Int. UK, Wallingford
Korniłłowicz-Kowalska T, Kitowski I (2013) Aspergillus fumigatus and Other thermophilic fungi in nests of wetland birds. Mycopath 175:43–56. https://doi.org/10.1007/s11046-012-9582-3
Kotzekidou P (1997) Heat resistance of Byssochlamys nivea, Byssochlamys fulva and Neosartorya fischeri isolated from canned tomato paste. J Food Sci 62:410–412. https://doi.org/10.1111/j.1365-2621.1997.tb04014.x
Kozakiewicz Z, Smith D (1994) Physiology of Aspergillus. In: Smith JE (ed) Aspergillus. Springer, Boston, MA, pp 23–40. https://doi.org/10.1007/978-1-4615-2411-3_2
Kumar R, Aneja KR (1999) Influence of incubation temperature on growth rates of fifteen thermophilous fungi. J Mycopathol Res 37:5–8
Langarica-Fuentes A, Handley PS, Houlden A, Fox G, Robson GD (2014a) An investigation of the biodiversity of thermophilic and thermotolerant fungal species in composts using culture-based and molecular techniques. Fungal Ecol 11:132–144. https://doi.org/10.1016/j.funeco.2014.05.007
Langarica-Fuentes A, Zafar U, Heyworth A, Brown T, Fox G, Robson GD (2014b) Fungal succession in an in-vessel composting system characterized using 454 pyrosequencing. FEMS Microbiol Ecol 88:296–308. https://doi.org/10.1111/1574-6941.12293
Langarica-Fuentes A, Fox G, Robson GD (2015) Metabarcoding analysis of home composts reveals distinctive fungal communities with a high number of unassigned sequences. Microbiol 161:1921–1932. https://doi.org/10.1099/mic.0.000153
Latgé JP (1999) Aspergillus fumigatus and aspergillosis. Clin Microbiol Rev 12(2):310–350. https://doi.org/10.1128/CMR.12.2.310
Le Goff O, Bru-Adan V, Bacheley H, Godon JJ, Wéry N (2010) The microbial signature of aerosols produced during the thermophilic phase of composting. J Appl Microbiol 108:325–340. https://doi.org/10.1111/j.1365-2672.2009.04427.x
Le Naourès C, Bonhomme J, Terzi N, Duhamel C, Galateau-Sallé F (2011) A fatal case with disseminated Myceliophthora thermophila infection in a lymphoma patient. Diagn Microbiol Infect Dis 70:267–269. https://doi.org/10.1016/j.diagmicrobio.2011.01.003
Lechner P, Linzer R, Mostbauer P, Binner E, Smidt E (2005) Klimarelevanz der Kompostierung unter Berücksichtigung der Verfahrenstechnik und Kompost-anwendung (KliKo). University of Natural Resources and Life Sciences of Vienna
Lee H, Lee YM, Jang Y, Lee S, Lee H, Ahn BJ, Kim GH, Kim JJ (2014) Isolation and analysis of the enzymatic properties of thermophilic fungi from compost. Mycobiol 42:181–184. https://doi.org/10.5941/MYCO.2014.42.2.181
Lieckfeldt E, Meyer W, Börner T (1993) Rapid identification and differentiation of yeasts by DNA and PCR fingerprinting. J Basic Microbiol 33:413–425. https://doi.org/10.1002/jobm.3620330609
Liu Q, Gao R, Li J, Lin L, Zhao J, Sun W, Tian C (2017) Development of a genome-editing CRISPR/Cas9 system in thermophilic fungal Myceliophthora species and its application to hyper-cellulase production strain engineering. Biotechnol Biofuels 10:1. https://doi.org/10.1186/s13068-016-0693-9
Maalej-Achouri I, Guerfali M, Romdhane IB-B, Gargouri A, Belghith H (2012) The effect of Talaromyces thermophilus cellulase-free xylanase and commercial laccase on lignocellulosic components during the bleaching of kraft pulp. Int Biodeterior Biodegrad 75:43–48. https://doi.org/10.1016/j.ibiod.2012.04.015
Madrid H, Cano J, Stchigel AJ, Gené J, Guarro J (2010) Ramophialophora humicola and Fibulochlamys chilensis, two new microfungi from soil. Mycol 102:605–612. https://doi.org/10.3852/09-128
Mahajan C, Basotra N, Singh S, Di Falco M, Tsang A, Chadha BS (2016) Malbranchea cinnamomea. A thermophilic fungal source of catalytically efficient lignocellulolytic glycosyl hydrolases and metal dependent enzymes. Bioresour Technol 200:55–63. https://doi.org/10.1016/j.biortech.2015.09.113
Maheshwari R, Bharadwaj G, Bhat MK (2000) Thermophilic fungi. Their physiology and enzymes. Microbiol Mol Biol Rev 64:461–488. https://doi.org/10.1128/MMBR.64.3.461-488.2000
Martin N, Guez MAU, Sette LD, Da Silva R, Gomes E (2010) Pectinase production by a Brazilian thermophilic fungus Thermomucor indicae-seudaticae N31 in solid-state and submerged fermentation. Microbiol 79:306–313. https://doi.org/10.1134/S0026261710030057
McPhillips K, Waters DM, Parlet C, Walsh DJ, Arendt EK, Murray PG (2014) Purification and characterisation of a β-1,4-xylanase from Remersonia thermophila CBS 540.69 and its application in bread making. Appl Biochem Biotech 172:1747–1762. https://doi.org/10.1007/s12010-013-0640-1
Meng Q, Yang W, Men M, Bello A, Xu X, Xu B, Deng L, Jiang X, Sheng S, Wu X, Han Y, Zhu H (2019) Microbial community succession and response to environmental variables during cow manure and corn straw composting. Front Microbiol 10:529. https://doi.org/10.3389/fmicb.2019.00529
Merheb-Dini C, Garcia GAC, Penna ALB, Gomes E, da Silva R (2012) Use of a new milk-clotting protease from Thermomucor indicae-seudaticae N31 as coagulant and changes during ripening of Prato cheese. Food Chem 130:859–865. https://doi.org/10.1016/j.foodchem.2011.07.105
Morgenstern I, Powlowski J, Ishmael N, Darmond C, Marqueteau S, Moisan MC, Quenneville G, Tsang A (2012) A molecular phylogeny of thermophilic fungi. Fungal Biol 116. https://doi.org/10.1016/j.funbio.2012.01.010
Mouronte-Roibás C, Leiro-Fernández V, Botana-Rial M, Ramos-Hernández C, Lago-Preciado G, Fiaño-Valverde C, Fernández-Villar A (2016) Lichtheimia ramosa. A fatal case of mucormycosis. Can Respir J 2016:2178218. https://doi.org/10.1155/2016/2178218
Mülhardt C (2013) Der Experimentator Molekularbiologie/Genomics, 7th edn. Springer Berlin Heidelberg, Berlin s.l.
Munday JS, Laven RA, Orbell GMB, Pandey SK (2006) Meningoencephalitis in an adult cow due to Mortierella Wolfii. J Vet Diagn Investig 18(6):619–622. https://doi.org/10.1177/104063870601800620
Nazir N, Mirza H, Akhtar N, Bajwa R, Nasin G (2007) Some studies of thermophilic and thermotolerant fungi from Lahore, Pakistan. Mycopath 5:95–100
Neher DA, Weicht TR, Bates ST, Leff JW, Fierer N (2013) Changes in bacterial and fungal communities across compost recipes, preparation methods, and composting times. PloS one 8:e79512. https://doi.org/10.1371/journal.pone.0079512
Nguyen HDT, Chabot D, Hirooka Y, Roberson RW, Seifert KA (2015) Basidioascus undulatus. Genome, origins, and sexuality. IMA Fungus 6:215–231. https://doi.org/10.5598/imafungus.2015.06.01.14
Noreen N, Ramzan N, Parveen Z, Shahzad S (2019) A comparative study of cow dung compost, goat pellets, poultry waste manure and plant debris for thermophilic, thermotolerant and mesophilic microflora with some new reports from Pakistan. Pak J Bot 51. https://doi.org/10.30848/PJB2019-3(42)
Nourrisson C, Garcia-Hermoso D, Morio F, Kauffmann-Lacroix C, Berrette N, Bonhomme J, Poirier P, Lortholary O (2017) Thermothelomyces thermophila human infections. Clin Microbiol Infect 23:338–341. https://doi.org/10.1016/j.cmi.2016.10.025
Okagbue RN (1989) Heat-resistant fungi in soil samples from northern nigeria. J Food Prot 52:59–61. https://doi.org/10.4315/0362-028X-52.1.59
Oliveira DS, Telis-Romero J, Da-Silva R, Franco CML (2014) Effect of a Thermoascus aurantiacus thermostable enzyme cocktail on wheat bread qualitiy. Food Chem 143:139–146. https://doi.org/10.1016/j.foodchem.2013.07.103
Ottow JCG (1972) Rose bengal as a selective aid in the isolation of fungi and actinomycetes from natural sources. Mycol 64:304–315. https://doi.org/10.1080/00275514.1972.12019265
Pan WZ, Huang XW, Wei KB, Zhang CM, Yang DM, Ding JM, Zhang KQ (2010) Diversity of thermophilic fungi in Tengchong Rehai National Park revealed by ITS nucleotide sequence analyses. J Micro (Seoul, Korea) 48:146–152. https://doi.org/10.1007/s12275-010-9157-2
Peterson SW (2004) Multilocus DNA sequence analysis shows that Penicillium biourgeianum is a distinct species closely related to P. brevicompactum and P. olsonii. Mycol Res 108:434–440. https://doi.org/10.1017/S0953756204009761
Peterson SW, Jurjević Z (2013) Talaromyces columbinus sp. nov., and genealogical concordance analysis in Talaromyces clade 2a. PloS one 8:e78084. https://doi.org/10.1371/journal.pone.0078084
Piecková E, Jesenská Z (1997) Toxinogenicity of heat-resistant fungi detected by a bio-assay. Int J Food Microbiol 36:227–229. https://doi.org/10.1016/S0168-1605(97)01239-7
Piecková E, Samson RA (2000) Heat resistance of Paecilomyces variotii in sauce and juice. J Ind Microbiol Biotech 24:227–230. https://doi.org/10.1038/sj.jim.2900794
Pitt JI, Hocking AD (2009) Spoilage of stored, processed and preserved foods. In: Pitt JI, Hocking AD (eds) Fungi and food spoilage. Springer Intern Publishing, Cham, pp 489–507
Powell AJ, Parchert KJ, Bustamante JM, Ricken JB, Hutchinson MI, Natvig DO (2012) Thermophilic fungi in an aridland ecosystem. Mycol 104:813–825. https://doi.org/10.3852/11-298
Puel O, Tadrist S, Galtier P, Oswald IP, Delaforge M (2005) Byssochlamys nivea as a source of mycophenolic acid. Appl Env Microbiol 71:550–553. https://doi.org/10.1128/AEM.71.1.550-553.2005
Raheja Y, Kaur B, Falco M, Tsang A, Chadha BS (2020) Secretome analysis of Talaromyces emersonii reveals distinct CAZymes profile and enhanced cellulase production through response surface methodology. Indust Crop Prod 152:112554. https://doi.org/10.1016/j.indcrop.2020.112554
Rajavaram RK, Bathini S, Girisham S, Reddy SM (2010) Incidence of thermophilic fungi from different substrates in Andhra Pradesh (India). Int J Pharm Bio Sci 3:BS33
Rhodes JC (2006) Aspergillus fumigatus. Growth and virulence. Med Mycol 44(Suppl 1):S77–S81. https://doi.org/10.1080/13693780600779419
Rigoldi F, Donini S, Redaelli A, Parisini E, Gautieri A (2018) Review. Engineering of thermostable enzymes for industrial applications. APL bioengineering 2:11501. https://doi.org/10.1063/1.4997367
Robledo A, Aguilar CN, Belmares-Cerda RE, Flores-Gallegos AC, Contreras-Esquivel JC, Montañez JC, Mussatto SI (2016) Production of thermostable xylanase by thermophilic fungal strains isolated from maize silage. CyTA - J Food 14:302–308. https://doi.org/10.1080/19476337.2015.1105298
Rodrigues RC, Fernandez-Lafuente R (2010) Lipase from Rhizomucor miehei as an industrial biocatalyst in chemical process. J Mol Catal B: Enzym 64:1–22. https://doi.org/10.1016/j.molcatb.2010.02.003
Rodriguez E, Mullaney EJ, Lei XG (2000) Expression of the Aspergillus fumigatus phytase gene in Pichia pastoris and characterization of the recombinant enzyme. Biochem Biophys Res Commun 268(2):373–378. https://doi.org/10.1006/bbrc.2000.2121
Salar RK (2018) Thermophilic fungi. Basic concepts and biotechnological applications. CRC PRESS, Boca Raton. https://doi.org/10.1201/9781351118187
Salar RK, Aneja KR (2006) Thermophilous fungi from temperate soils of northern India. J Agric Technol 2(1):49–58
Salomão BCM, Muller C, Amparo HC, Aragão GMF (2014) Survey of molds, yeast and Alicyclobacillus spp. from a concentrated apple juice productive process. Braz J Microbiol 45(2014):49–58. https://doi.org/10.1590/s1517-83822014000100008
Samson RA, Dijksterhuis J (2007) Food Mycology. A multifaceted approach to fungi and food. CRC PRESS, Boca Raton. https://doi.org/10.1201/9781420020984
Samson RA, Hoekstra ES, Lund F, Filtenborg O, Frisvad JC (2000) Methods for the detection, isolation and characterization of food-borne fungi. In: Samson RA, Hoekstra ES, Frisvad JC, Filtenborg O (eds) Introduction to food- and airborne fungi. Centraalbureau voor Schimmelcultures, Utrecht, pp 283–297
Sandona K, Billingsley Tobias TL, Hutchinson MI, Natvig DO, Porras-Alfaro A (2019) Diversity of thermophilic and thermotolerant fungi in corn grain. Mycol 111:719–729. https://doi.org/10.1080/00275514.2019.1631137
Scaramuzza N, Berni E (2014) Heat-resistance of Hamigera avellanea and Thermoascus crustaceus isolated from pasteurized acid products. Int J Food Microbiol 168-169:63–68. https://doi.org/10.1016/j.ijfoodmicro.2013.10.007
Schiano-di-Cola C, Kołaczkowski B, Sørensen TH, Christensen SJ, Cavaleiro AM, Windahl MS, Borch K, Morth JP, Westh P (2020) Structural and biochemical characterization of a family 7 highly thermostable endoglucanase from the fungus Rasamsonia emersonii. FEBS J 287:2577–2596. https://doi.org/10.1111/febs.15151
Schoch CL, Seifert KA, Huhndorf S et al (2012) Nuclear ribosomal internal transcribed spacer (ITS) region as a universal DNA barcode marker for fungi. Proc Nat Acad Sci 109:6241–6246. https://doi.org/10.1073/pnas.1117018109
Schuerg T, Gabriel R, Baecker N, Baker SE, Singer SW (2017) Thermoascus aurantiacus is an intriguing host for the industrial production of cellulases. CBIOT 6:89–97. https://doi.org/10.2174/2211550105666160520123504
Seifert KA, Nickerson NL, Corlett M, Jackson ED, Louis-Seize G, Davies RJ (2004) Devriesia, a new hyphomycete genus to accommodate heat-resistant, cladosporium-like fungi. Can J Bot. 82:914–926. https://doi.org/10.1139/b04-070
Sen SK, Jana A, Bandyopadhyay P, Das Mohapatra PK, Raut S (2016) Thermostable amylase production from hot spring isolate Exiguobacterium sp. A promising agent for natural detergents. Sustain Chem Pharm 3:59–68. https://doi.org/10.1016/j.scp.2016.04.002
Sevo M, Degrassi G, Skoko N, Venturi V, Ljubijankić G (2002) Production of glycosylated thermostable Providencia rettgeri penicillin G amidase in Pichia pastoris. FEMS Yeast Res 1:271–277. https://doi.org/10.1111/j.1567-1364.2002.tb00045.x
Sezen S, Demirel R (2019) Preliminary study on the determination of heat resistant fungi in agricultural soils. Mantar Dergisi 10:60–66. https://doi.org/10.30708/mantar.632181
Sharma A, Tewari R, Rana SS, Soni R, Soni SK (2016) Cellulases. Classification, methods of determination and industrial applications. Appl Biochem Biotechnol 179:1346–1380. https://doi.org/10.1007/s12010-016-2070-3
Singh B (2016) Myceliophthora thermophila syn. Sporotrichum thermophile. A thermophilic mould of biotechnological potential. Crit Rev Biotechnol 36:59–69. https://doi.org/10.3109/07388551.2014.923985
Sivagnanam S, Chen SC-A, Halliday C, Packham D (2013) Thermomyces lanuginosus infective endocarditis. Case report and a review of endocarditis due to uncommon moulds. Med Mycol Case Rep 2:152–155. https://doi.org/10.1016/j.mmcr.2013.09.001
Stielow JB, Lévesque CA, Seifert KA et al (2015) One fungus, which genes? Development and assessment of universal primers for potential secondary fungal DNA barcodes. Persoonia 35:242–263. https://doi.org/10.3767/003158515X689135
Su YY, Cai L (2013) Rasamsonia composticola, a new thermophilic species isolated from compost in Yunnan, China. Mycol Prog 12:213–221. https://doi.org/10.1007/s11557-012-0827-9
Tansey MR (1971) Isolation of thermophilic fungi from self-heated, industrial wood chip piles. Mycol 63:537. https://doi.org/10.2307/3757550
Tansey MR, Jack MA (1976) Thermophilic fungi in sun-heated soils. Mycol 68:1061–1075. https://doi.org/10.1080/00275514.1976.12019988
Tekaia F, Latgé JP (2005) Aspergillus fumigatus. Saprophyte or pathogen. Curr Opin Microbiol 8:385–392. https://doi.org/10.1016/j.mib.2005.06.017
Thanh VN, Thuy NT, Huong HTT, Hien DD, Hang DTM, Anh DTK, Hüttner S, Larsbrink J, Olsson L (2019) Surveying of acid-tolerant thermophilic lignocellulolytic fungi in Vietnam reveals surprisingly high genetic diversity. Sci Rep 9:3674. https://doi.org/10.1038/s41598-019-40213-5
Tong X, Lange L, Grell MN, Busk PK (2015) Hydrolysis of wheat arabinoxylan by two acetyl xylan esterases from Chaetomium thermophilum. Appl Biochem Biotechnol 175:1139–1152. https://doi.org/10.1007/s12010-014-1348-6
Tournas V (1994) Heat-resistant fungi of importance to the food and beverage industry. Crit Rev Biotechnol 20:243–263. https://doi.org/10.3109/10408419409113558
Tournas V, Traxler RW (1994) Heat resistance of a Neosartorya fischeri strain isolated from pineapple juice frozen concentrate. J Food Prot 57:814–816. https://doi.org/10.4315/0362-028x-57.9.814
Tranquillini R, Scaramuzza N, Berni E (2017) Occurrence and ecological distribution of heat resistant moulds spores (HRMS) in raw materials used by food industry and thermal characterization of two Talaromyces isolates. Int J Food Microbiol 242:116–123. https://doi.org/10.1016/j.ijfoodmicro.2016.11.023
Vilgalys R, Hester M (1990) Rapid genetic identification and mapping of enzymatically amplified ribosomal DNA from several Cryptococcus species. J Bacteriol 172:4238–4246. https://doi.org/10.1128/JB.172.8.4238-4246.1990
Vu D, Groenewald M, de Vries M, Gehrmann T, Stielow B, Eberhardt U, Al-Hatmi A, Groenewald JZ, Cardinali G, Houbraken J, Boekhout T, Crous PW, Robert V, Verkley GJM (2019) Large-scale generation and analysis of filamentous fungal DNA barcodes boosts coverage for kingdom fungi and reveals thresholds for fungal species and higher taxon delimitation. Stud Mycol 92:135–154. https://doi.org/10.1016/j.simyco.2018.05.001
Wang XW, Yang FY, Meijer M, Kraak B, Sun BD, Jiang YL, Wu YM, Bai FY, Seifert KA, Crous PW, Samson RA, Houbraken J (2019) Redefining Humicola sensu stricto and related genera in the Chaetomiaceae. Stud Mycol 93:65–153. https://doi.org/10.1016/j.simyco.2018.07.001
White TJ, Burns T, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis M, Gelfand D, Sninsky J, White T (eds) PCR protocols: a guide to methods and applications. Academic Press, Cambridge, pp 315–322
Yin G, Zhang Y, Pennerman KK, Wu G, Hua SST, Yu J, Jurick WM, Guo A, Bennett JW (2017) Characterization of blue mold Penicillium species isolated from stored fruits using multiple highly conserved loci. J fungi 3:12. https://doi.org/10.3390/jof3010012
Zhang W, Yang R, Fang W, Yan L, Lu J, Sheng J, Lv J (2016) Characterization of thermophilic fungal community associated with pile fermentation of Pu-erh tea. Int J Food Microbiol 227:29–33. https://doi.org/10.1016/j.ijfoodmicro.2016.03.025
Acknowledgements
We thank Sebastian Diamantidis for helping during sampling and isolation of specimens. Furthermore, we thank Sabine Adler, the Kleingärtnerverein Friederika 1932 e.V. (Bochum, Germany) and the botanical garden of the Ruhr-University Bochum for granting access to compost and soil material. We are also grateful to the department of applied microbiology (Ruhr-University Bochum) and Prof. Dr. Julia Bandow for access to their facilities and equipment. We acknowledge the constructive comments of the two anonymous reviewers, which helped us improving the manuscript.
Funding
This project was funded by Stiftung Mercator (MERCUR: Pr 2017-0020).
Author information
Authors and Affiliations
Contributions
FW and DB designed the sampling campaign, FW conducted the isolation, cultivation, and molecular work with the help of students. FW and MG evaluated the results and performed the taxonomic assignment. FW wrote the manuscript. MG and DB critically reviewed the manuscript. MG edited the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Consent to participate
Not applicable
Consent for publication
Not applicable
Conflict of interest
The authors declare no competing interests.
Additional information
Section Editor: Marc Stadler
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Online Resource 1
Sampled plots and collection dates. Botanical garden samples originated from the Ruhr-University Bochum (RUB) and allotment gardens belonged to the Kleingärtnerverein Friederika 1932 e.V. in Bochum, Germany. The coordinates of each plot are indicated, as well as the associated plant species and the type of compost. (DOCX 24 kb)
Online Resource 2
Fungal isolates obtained from compost and soil, using different sampling strategies (see Table 1), were collected from different plots (see Online resource 1). Isolates were obtained using different media, in combination with incubation at 45 °C, 55 °C or room temperature (RT), and a heat-shock (HS) treatment. Molecular identification of the isolates was performed based on MSP-PCR patterns and/or barcode(s) (ITS and/or LSU, ß-tubulin (tub2), calmodulin (calM)). The newly generated barcodes were deposited at ENA database and the respective accession numbers are provided. The accession number of the best hit and the sequence similarity from the BLAST analyses are indicated. When feasible, at least one isolate of each species was deposited in the DSMZ culture collection. (XLSX 28 kb)
ESM 3
(FASTA 101 kb)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Witfeld, F., Begerow, D. & Guerreiro, M.A. Improved strategies to efficiently isolate thermophilic, thermotolerant, and heat-resistant fungi from compost and soil. Mycol Progress 20, 325–339 (2021). https://doi.org/10.1007/s11557-021-01674-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11557-021-01674-z