Abstract
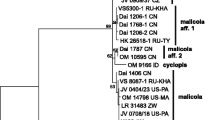
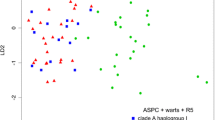
It has been previously shown that the Phlebia livida complex consists of two incompatible subspecies, viz. ssp. livida and ssp. tuberculata. Here, we explain that the two subspecies can be distinguished based on morphology, phylogenetic analyses of nuclear ITS sequences, genetic distance, and a haplotype network, as well as by substrate preference. Subspecies tuberculata is raised to specific rank and a thorough description is provided. According to the results of the network analysis, Phlebia tuberculata comb. nov. seems to be an older species than Phlebia livida, congruent with its wider distribution, and at least some of its present populations have survived in the Caucasus region during the last glaciation. A key to the species of the Phlebia livida complex is presented.


Similar content being viewed by others
References
Allmér J, Vasiliauskas R, Ihrmark K, Stenlid J, Dahlberg A (2006) Wood-inhabiting fungal communities in woody debris of Norway spruce (Picea abies (L.) Karst.), as reflected by sporocarps, mycelial isolations and T-RFLP identification. FEMS Microbiol Ecol 55:57–67
Atkinson RJ, Rokas A, Stone GN (2006) Longitudinal patterns in species richness and genetic diversity in European oaks and oak gallwasps. In: Weiss S, Ferrand N (ed) Phylogeography of Southern European Refugia. Evolutionary perspectives on the origins and conservation of European biodiversity. Springer, Netherlands, pp 127–151
Begerow D, Nilsson RH, Unterseher M, Maier W (2010) Current state and perspectives of fungal DNA barcoding and rapid identification procedures. Appl Microbiol Biotechnol 87:99–108
Clement M, Posada D, Crandall K (2000) TCS: a computer program to estimate gene genealogies. Mol Ecol 9:1657–1660
Crandall KA, Templeton AR (1993) Empirical tests of some predictions from coalescent theory with applications to intraspecific phylogeny reconstruction. Genetics 134:959–969
Damm S, Schierwater B, Hadrys H (2010) An integrative approach to species discovery in odonates: from character-based DNA barcoding to ecology. Mol Ecol 19:3881–3893
Dayrat B (2005) Towards integrative taxonomy. Biol J Linn Soc 85:407–415
Duhem (2008) À propos de plusieurs Phlebia leptocystidiés: Phlebia acanthocystis, P. caspica, P. chrysocreas, P. ochraceofulva et P. subochracea – Note sur Phlebia tristis. Bull Soc Mycol Fr 124:299–342
Eriksson J, Hjortstam K, Ryvarden L (1981) The Corticiaceae of North Europe, vol 6. Phlebia–Sarcodontia. Fungiflora, Oslo
Fries EM (1836, publ. 1838) Epicrisis Systematis Mycologici, seu Synopsis Hymenomycetum. Typographia Academica, Uppsala
Gadanho M, Sampaio JP, Spencer-Martins I (2001) Polyphasic taxonomy of the basidiomycetous yeast genus Rhodosporidium: R. azoricum sp. nov. Can J Microbiol 47:213–221
Gardes M, Bruns TD (1993) ITS primers with enhanced specificity for basidiomycetes – application to the identification of mycorrhizae and rusts. Mol Ecol 2:113–118
Ghobad-Nejhad M, Hallenberg N, Parmasto E, Kotiranta H (2009) A first annotated checklist of corticioid and polypore basidiomycetes of the Caucasus region. Mycol Balc 6:123–168
Ghobad-Nejhad M, Nilsson RH, Hallenberg N (2010) Phylogeny and taxonomy of the genus Vuilleminia (Basidiomycota) based on molecular and morphological evidence, with new insights into Corticiales. Taxon 59:1519–1534
Goloboff P (1999) NONA (NO NAME) ver. 2. Published by the author, Tucumán, Argentina
Gross G (1972) Kernzahl und sporenvolumen bei einigen Hymenogasterarten. Zeit Pilzk 38:109–158
Hallenberg N (1980) New taxa of Corticiaceae from N Iran (basidiomycetes). Mycotaxon 11:447–475
Hallenberg N (1988) Species delimitation in Corticiaceae (basidiomycetes). Mycotaxon 31:445–465
Hallenberg N (1991) Pairing tests with species of Aphyllophorales (basidiomycetes) from two phytogeographically isolated areas. Mycotaxon 42:355–386
Hallenberg N, Larsson E (1991) Differences in cultural characters and electrophoretic patterns among sibling species in four different species complexes (Corticiaceae, basidiomycetes). Mycologia 83:131–141
Hallenberg N, Larsson E (1993) On taxonomy of Phlebia livida. Mycol Res 97:351–354
Hallenberg N, Ryberg M, Nilsson RH, Wood AR, Wu SH (2008) Pseudolagarobasidium (Basidiomycota): on the reinstatement of a genus of parasitic, saprophytic, and endophytic resupinate fungi. Botany 86:1319–1325
Harris DJ, Crandall KA (2000) Intragenomic variation within ITS1 and ITS2 of freshwater crayfishes (Decapoda: Cambaridae): Implications for phylogenetic and microsatellite studies. Mol Biol Evol 17:284–291
Hewitt GM (1999) Post-glacial re-colonization of European biota. Biol J Linn Soc 68:87–112
Hewitt GM (2004) The structure of biodiversity, insights from molecular biodiversity. Front Zool 1:4
Hinrikson HP, Hurst SF, Lott TJ, Warnock DW, Morrison CJ (2005) Assessment of ribosomal large-subunit D1-D2, internal transcribed region spacer 1, and internal transcribed spacer 2 regions as targets for molecular identification of medically important Aspergillus species. J Clin Microbiol 43:2092–2103
Hjortstam K (1989) Corticioid fungi described by M.J. Berkeley. Kew Bull 44:301–315
Huss VAR, Ciniglia C, Cennamo P, Cozzolino S, Pinto G, Pollio A (2002) Phylogenetic relationships and taxonomic position of Chlorella-like isolates from low pH environments (pH < 3.0). BMC Evol Biol 2:13
Katoh K, Toh H (2008) Recent developments in the MAFFT multiple sequence alignment program. Brief Bioinforma 9:286–298
Kotlík P, Marková S, Choleva L, Bogutskaya NG, Ekmekçi FG, Ivanova PP (2008) Divergence with gene flow between Ponto-Caspian refugia in an anadromous cyprinid Rutilus frisii revealed by multiple gene phylogeography. Mol Ecol 17:1076–1088
Larsen MJ (1974) Contribution to the taxonomy of the genus Tomentella. Mycol Mem 4:1–145
Liou GY, Chen SR, Wei YH, Lee FL, Fu HM, Yuan GF, Stalpers JA (2007) Polyphasic approach to the taxonomy of the Rhizopus stolonifer group. Mycol Res 111:196–203
Milne RI (2006) Northern Hemisphere plant disjunctions: a window on Tertiary land bridges and climate change? Ann Bot 98:465–472
Müller K, Quandt D, Müller J, Neinhuis C (2005) PhyDe: Phylogenetic Data Editor. v0.995. Available from: http://www.phyde.de
Nilsson RH, Kristiansson E, Ryberg M, Hallenberg N, Larsson KH (2008) Intraspecific variability in the kingdom fungi as expressed in the international sequence databases and its implications for molecular species identification. Evol Bioinforma Online 4:193–201
Nixon KC (2002) WinClada ver. 1.00.08 Published by the author, Ithaca, NY, USA
Nylander JAA (2004) MrModeltest v2. Evolutionary Biology Center, Uppsala University, Program distributed by the author
Parmasto E, Nilsson H, Larsson KH (2004) Cortbase v2 - extensive updates of a nomenclatural database for corticioid fungi (hymenomycetes). Phyloinformatics 5:1–7
Pattengale ND, Alipour M, Bininda-Emonds ORP, Moret BME, Stamatakis A (2009) How many bootstrap replicates are necessary? LNCS 5541:184–200
Persoon CH (1799, publ. 1800) Observationes Mycologicae 2. Gesnerus, Usterius & Wolfius, Germany, Leipzig
Petersen JH (1996) Farvekort. The Danish Mycological Society´s colour-chart. Foreningen til Svampekundskabens Fremme, Greve
Posada D, Crandall KA (2001) Intraspecific gene genealogies: Trees grafting into networks. Trends Ecol Evol 16:37–45
Randi E (2006) Phylogeography of South European mammals. In: Weiss S, Ferrand N (ed) Phylogeography of Southern European Refugia. Evolutionary perspectives on the origins and conservation of European biodiversity. Springer, Netherlands, pp 101–126
Ronquist F, Huelsenbeck JP (2003) MRBAYES 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19:1572–1574
Samson RA, Varga J (2009) What is a species in Aspergillus? Med Mycol 47 (Supplement 1):S13–S20
Samson RA, Houbraken J, Varga J, Frisvad JC (2009) Polyphasic taxonomy of the heat resistant ascomycete genus Byssochlamys and its Paecilomyces anamorphs. Persoonia 22:14–27
Silvestro D, Michalak I (2010) raxmlGUI: a graphical front-end for RAxML. http://sourceforge.net/projects/raxmlgui/. Accessed August 2010
Slepecky RA, Starmer WT (2009) Phenotypic plasticity in fungi: a review with observations on Aureobasidium pullulans. Mycologia 101:823–832
Stamatakis A (2006) RAxML-VI-HPC: maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics 22:2688–2690
Taberlet P, Fumagalli L, Wust-Saucy AG, Cosson JF (1998) Comparative phylogeography and postglacial colonization routes in Europe. Mol Ecol 7:453–464
Tarasov PE, Volkova VS, Webb T III, Guiot J, Andreev AA, Bezusko LG, Bezusko TV, Bykova GV, Dorofeyuk NI, Kvavadze EV, Osipova IM, Panova NK, Sevastyanov DV (2000) Last glacial maximum biomes reconstructed from pollen and plant macrofossil data from northern Eurasia. J Biogeogr 27:609–620
Volodicheva N (2002) The Caucasus. In: Shahgedanova M (ed) The Physical Geography of Northern Eurasia. Oxford University Press, Oxford, pp 350–376
Wu SH, Nilsson RH, Chen CT, Yu SY, Hallenberg N (2010) The white-rotting genus Phanerochaete is polyphyletic and distributed throughout the phleboid clade of the Polyporales (Basidiomycota). Fungal Divers 42:107–118
Acknowledgements
The curators of herbaria BPI, FH, GB, H, IRAN, K, and UPS are warmly thanked. We thank Bernard Duhem (Paris) for kindly sending us some material of Phlebia tuberculata, and two anonymous reviewers and Neil Bell (Helsinki) for advice on the text. Support from Societas pro Fauna et Flora Fennica is gratefully acknowledged.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Fig. 1
Fruiting body configuration in Phlebia tuberculata (a–f) compared with Phlebia livida (g–l). a, b Ghobad-Nejhad 634 c, d Ghobad-Nejhad 796 e, f Hallenberg 9015 g, h Miettinen 101 i, j Hallenberg 3876 k, l Kotiranta 22433. Scale bars a–d and g–l = 3 mm, scale bars e–f = 1 mm (PDF 188 kb)
Rights and permissions
About this article
Cite this article
Ghobad-Nejhad, M., Hallenberg, N. Multiple evidence for recognition of Phlebia tuberculata, a more widespread segregate of Phlebia livida (Polyporales, Basidiomycota). Mycol Progress 11, 27–35 (2012). https://doi.org/10.1007/s11557-010-0722-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11557-010-0722-1