Abstract
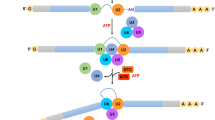
Photoperiod and temperature-sensitive genetic male sterility (PGMS and TGMS) plants have a number of desirable characteristics for hybrid production. Two-line hybrids developed using the PGMS/TGMS system now account for a large proportion of rice production in China. In this paper, we summarize recent advances on molecular regulation mechanisms and genetics of PGMS/TGMS in rice. We suggest that temperature-sensitive splicing, an important posttranscriptional regulatory mechanism in modulating gene expression and eventually development and differentiation, is also an important molecular regulation mechanism of TGMS in rice. We review those factors involved in temperature-sensitive splicing like cis splice site, snRNA, trans pre-mRNA splicing protein and SR proteins, and delineate that splicing could be regulated by a complex cell signaling pathway. These might shed light on other unknown molecular PGMS/TGMS mechanisms.
Similar content being viewed by others
References
Ma H. Molecular genetic analyses of microsporogenesis and microgametogenesis in flowering plants. Annu Rev Plant Biol, 2005, 56: 393–434
Shi M S. The discovery and preliminary studies of the photoperiodsensitive recessive male sterile rice (Oryza sativa L. ssp. japonica) (in Chinese). Sci Agric Sin, 1985, 2: 44–48
Zhang Z G. The influence of photoperiod on pollen fertility change of Hubei photoperiod-sensitive genic male sterile rice (in Chinese). Chin J Rice Sci, 1987, 1: 137–143
Yuan L P, Peng J M. Hybrid Rice and World Food Security (in Chinese). Beijing: China Science and Technology Press, 2005
Zhang Q F, Shen B Z, Dai X K, et al. Using bulked extremes and recessive class to map genes for photoperiod-sensitive genic male sterility in rice. Proc Natl Acad Sci USA, 1994, 91: 8675–8679
Mei M H, Dai X K, Xu C G, et al. Mapping and genetic analysis of the genes for photoperiod-sensitive genic sterility in rice using the original mutant Nongken 58S. Crop Sci, 1999, 39: 1711–1715
Mei M H, Chen L, Zhang Z H, et al. Pms3 is the locus causing the original photoperiod-sensitive genic male sterility mutation of Nongken 58S. Sci China Ser C-Life Sci, 1999, 42: 316–322
Lu Q, Li X H, Guo D, et al. Localization of pms3, a gene for photo period sensitive genic male sterility, to a 28.4-kb DNA fragment. Mol Genet Genomics, 2005, 273: 507–511
Liu N, Shan Y, Wang F P, et al. Identification of an 85-kb DNA fragment containing pms1, a locus for photoperiod-sensitive genic male sterility in rice. Mol Genet Genomics, 2001, 266: 271–275
Yu J S, Fan Y R, Liu N, et al. Rapid genome evolution in Pms1 region of rice revealed by comparative sequence analysis. Chinese Sci Bull, 2007, 52: 912–921
Peng H F, Zhang Z F, Wu B, et al. Molecular mapping of two reverse photoperiod-sensitive genic male sterility genes (rpms1 and rpms2) in rice (Oryza sativa L.). Theor Appl Genet, 2008, 118: 77–83
Wang B, Xu W W, Wang J Z, et al. Tagging and mapping the thermo-sensitive genic male-sterile gene in rice with molecular markers. Theor Appl Genet, 1995, 91: 1111–1114
Jia J H, Zhang D S, Li C Y, et al. Molecular mapping of the reverse thermo-sensitive genic male-sterile gene (rtms1) in rice. Theor Appl Genet, 2001, 103: 607–612
Yamaguchi Y, Ikeda R, Hirasawa H, et al. Linkage analysis of the thermo-sensitive genic male sterility gene Tms2 in rice (Oryza sativa L.). Breed Sci, 1997, 47: 371–377
Lopez M T, Toojinda T, Vanavichit A, et al. Microsatellite markers flanking the tms2 gene facilitated tropical TGMS rice line development. Crop Sci, 2003, 43: 2267–2271
Pitnjam K, Chakhonkaen S, Toojinda T, et al. Identification of a deletion in tms2 and development of gene-based markers for selection. Planta, 2008, 228: 813–822
Subudhi P K, Borkakati R K, Virmani S S, et al. Molecular mapping of a thermosensitive genetic male-sterility gene in rice using bulked segregant analysis. Genome, 1997, 40: 188–194
Wang Y G, Xing Q H, Deng Q Y, et al. Fine mapping of the rice thermo-sensitive genic male-sterile gene tms5. Theor Appl Genet, 2003, 107: 917–921
Yang Q K, Liang C Y, Zhuang W, et al. Characterization and identification of the candidate gene of rice thermo-sensitive genic male sterile gene tms5 by mapping. Planta, 2007, 225: 321–330
Dong N V, Subudhi P K, Luong P N, et al. Molecular mapping of a rice gene conditioning thermosensitive genic male sterility using AFLP, RFLP and SSR techniques. Theor Appl Genet, 2000, 100: 727–734
Reddy O, Siddiq E, Ali N, et al. Genetic analysis of temperature-sensitive male sterility in rice. Theor Appl Genet, 2000, 100: 794–801
Wang C H, Zhang P, Ma Z R, et al. Development of a genetic marker linked to a new thermo-sensitive male sterile gene in rice (Oryza sativa L.). Euphytica, 2004, 140: 217–222
Lee D S, Chen L J, Suh H S. Genetic characterization and fine mapping of a novel thermo-sensitive genic male-sterile gene tms6 in rice (Oryza sativa L.). Theor Appl Genet, 2005, 111: 1271–1277
Koh H J, Heu M H. Agronomic characteristics of a mutant for genic male sterility-chalky endosperm and its utilization on F1 hybrid breeding system in rice. Korean J Crop Sci, 1995, 40: 684–696
Koh H J, Son Y H, Heu M H, et al. Molecular mapping of a new genic male-sterility gene causing chalky endosperm in rice (Oryza sativa L.). Euphytica, 1999, 106: 57–62
Woo M O, Ham T H, Ji H S, et al. Inactivation of the UGPase1 gene causes genic male sterility and endosperm chalkiness in rice (Oryza sativa L.). Plant J, 2008, 54: 190–204
Chen R Z, Zhao X, Shao Z, et al. Rice UDP-glucose pyrophosphorylase1 is essential for pollen callose deposition and its cosuppression results in a new type of thermosensitive genic male sterility. Plant Cell, 2007, 19: 847–861
Christensen A H, Sharrock R A, Quail P H. Maize polyubiquitin genes: Structure, thermal perturbation of expression and transcript splicing, and promoter activity following transfer to protoplasts by electroporation. Plant Mol Biol, 1992, 18: 675–689
Bowman J L, Smyth D R, Meyerowitz E M. Genes directing flower development in Arabidopsis. Plant Cell, 1989, 1: 37–52
Zachgo S, Silva E A, Motte P, et al. Functional analysis of the Antirrhinum floral homeotic DEFICIENS gene in vivo and in vitro by using a temperature-sensitive mutant. Development, 1995, 121: 2861–2875
Sablowski R W, Meyerowitz E M. Temperature-sensitive splicing in the floral homeotic mutant apetala3-1. Plant Cell, 1998, 10: 1453–1463
Yi Y, Jack T. An intragenic suppressor of the Arabidopsis floral organ identity mutant apetala3-1 functions by suppressing defects in splicing. Plant Cell, 1998, 10: 1465–1477
Majercak J, Sidote D, Hardin P E, et al. How a circadian clock adapts to seasonal decreases in temperature and day length. Neuron, 1999, 24: 219–230
Colot H V, Loros J J, Dunlap J. Temperature-modulated alternative splicing and promoter use in the circadian clock gene frequency. Mol Biol Cell, 2005, 16: 5563–5571
Diernfellner A C R, Schafmeier T, Merrow M W, et al. Molecular mechanism of temperature-sensing by the circadian clock of Neurospora crassa. Genes Dev, 2005, 19: 1968–1973
Majercak J, Chen W F, Edery I. Splicing of the period gene 3′-terminal intron is regulated by light, circadian clock factors, and phospholipase C. Mol Cell Biol, 2004, 24: 3359–3372
Collins B H, Rosato E, Kyriacou C P. Seasonal behavior in Drosophila melanogaster requires the photoreceptors, the circadian clock, and phospholipase C. Proc Natl Acad Sci USA, 2004, 101: 1945–1950
Chen W F, Low K H, Lim C, et al. Thermosensitive splicing of a clock gene and seasonal adaptation. Cold Spring Harb Symp Quant Biol, 2007, 72: 599–606
Brunner M, Schafmeier T. Transcriptional and post-transcriptional regulation of the circadian clock of cyanobacteria and Neurospora. Genes Dev, 2006, 20: 1061–1074
Reddy A S N. Alternative splicing of pre-messenger RNAs in plants in the genomic era. Annu Rev Plant Biol, 2007, 58: 267–294
Rappsilber J, Ryder U, Lamond A I, et al. Large scale proteomic analysis of the human spliceosome. Genome Res, 2002, 12: 1231–1245
Zhou Z, Licklider L J, Gygi S P, et al. Comprehensive analysis of the human spliceosome. Nature, 2002, 419: 182–185
Chen Y I G, Moore R E, Ge H Y, et al. Proteomic analysis of in vivo-assembled pre-mRNA splicing complexes expands the catalog of participating factors. Nucleic Acids Res, 2007, 35: 3928–3944
Lin R J, Lustig A, Abelson J. Splicing of yeast nuclear pre-mRNA in vitro requires a functional 40S spliceosome and several extrinsic factors. Genes Dev, 1987, 1: 7–18
Grainger R J, Beggs J D. Prp8 protein: At the heart of the spliceosome. RNA, 2005, 11: 533–557
Schwer B, Guthrie C. A conformational rearrangement in the spliceosome is dependent on PRP16 and ATP hydrolysis. EMBO J, 1992, 11: 5033–5039
Wang Y, Wagner J D O, Guthrie C. The DEAH-box splicing factor Prp16 unwinds RNA duplexes in vitro. Curr Biol, 1998, 8: 441–451
Raghunathan P L, Guthrie C. RNA unwinding in U4/U6 snRNPs requires ATP hydrolysis and the DEIH-box splicing factor Brr2. Curr Biol, 1998, 8: 847–855
Kim S H, Lin R J. Spliceosome activation by PRP2 ATPase prior to the first transesterification reaction of pre-mRNA splicing. Mol Cell Biol, 1996, 16: 6810–6819
Wagner J D, Jankowsky E, Company M, et al. The DEAH-box protein PRP22 is an ATPase that mediates ATP-dependent mRNA release from the spliceosome and unwinds RNA duplexes. EMBO J. 1998, 17: 2926–2937
Arenas J E, Abelson J N. Prp43: An RNA helicase-like factor involved in spliceosome disassembly. Proc Natl Acad Sci USA, 1997, 94: 11798–11802
Blanton S, Srinivasan A, Rymond B C. PRP38 encodes a yeast protein required for pre-mrna splicing and maintenance of stable U6 small nuclear RNA levels. Mol Cell Biol, 1992, 12: 3939–3947
Chan S P, Kao D I, Tsai W Y, et al. The Prp19p-associated complex in spliceosome activation. Science, 2003, 302: 279–282
Crotti L B, Bacikova D, Horowitz D S. The Prp18 protein stabilizes the interaction of both exons with the U5 snRNA during the second step of pre-mRNA splicing. Genes Dev, 2007, 21: 1204–1216
Valadkhan S. The spliceosome: Caught in a web of shifting interactions. Curr Opin Struct Biol, 2007, 17: 310–315
Madhani H D, Bordonne R, Guthrie C. Multiple roles for U6 snRNA in the splicing pathway. Genes Dev, 1990, 4: 2264–2277
Alvarez C J, Romfo C M, Vanhoy R W, et al. Mutational analysis of U1 function in Schizosaccharomyces pombe: Pre-mRNAs differ in the extent and nature of their requirements for this snRNA in vivo. RNA, 1996, 2: 404–418
Maddock J R, Roy J, Woolford J L J. Six novel genes necessary for pre-mRNA splicing in Saccharomyces cerevisiae. Nucleic Acids Res, 1996, 24: 1037–1044
Xiao S H, Manley J L. Phosphorylation of the ASF/SF2 RS domain affects both protein-protein and protein-RNA interactions and is necessary for splicing. Genes Dev, 1997, 11: 334–344
Xiao S H, Manley J L. Phosphorylation-dephosphorylation differentially affects activities of splicing factor ASF/SF2. EMBO J, 1998, 17: 6359–6367
Gui J F, Lane W S, Fu X D. A serine kinase regulates intracellular localization of splicing factors in the cell cycle. Nature, 1994, 369: 678–682
Colwill K, Pawson T, Andrews B, et al. The Clk/Sty protein kinase phosphorylates SR splicing factors and regulates their intranuclear distribution. EMBO J, 1996, 15: 265–275
Misteli T, Spector D L. Serine/threonine phosphatase 1 modulates the subnuclear distribution of pre-mRNA splicing factors. Mol Biol Cell, 1996, 7: 1559–1572
Misteli T. RNA splicing: what has phosphorylation got to do with it? Curr Biol, 1999, 9: 198–200
Shi Y, Reddy B, Manley J L. PP1/PP2A phosphatases are required for the second step of pre-mRNA splicing and target specific snRNP proteins. Mol Cell, 2006, 23: 819–829
Huang Y, Steitz J A. Splicing factors SRp20 and 9G8 promote the nucleocytoplasmic export of mRNA. Mol Cell, 2001, 7: 899–905
Lemaire R, Prasad J, Kashima T, et al. Stability of a PKCI-1-related mRNA is controlled by the splicing factor ASF/SF2: A novel function for SR proteins. Genes Dev, 2002, 16: 594–607
Sanford J R, Gray N K, Beckmann K, et al. A novel role for shuttling SR proteins in mRNA translation. Genes Dev, 2004, 18: 755–768
Graveley B R. Sorting out the complexity of SR protein functions. RNA, 2000, 6: 1197–1211
Iida K, Go M. Survey of conserved alternative splicing events of mRNAs encoding SR proteins in land plants. Mol Biol Evol, 2006, 23: 1085–1094
Isshiki M, Tsumoto A, Shimamoto K. The serine/arginine-rich protein family in rice plays important roles in constitutive and alternative splicing of pre-mRNA. Plant Cell, 2006, 18: 146–158
Reddy A S N. Plant serine/arginine-rich proteins and their role in pre-mRNA splicing. Trends Plant Sci, 2004, 9: 541–547
Hopf N, Plesofsky-Vig N, Brambl R. The heat shock response of pollen and other tissues of maize. Plant Mol Biol, 1992, 19: 623–630
Larkin P D, Park W D. Transcript accumulation and utilization of alternate and nonconsensus splice sites in rice granule-bound starch synthase are temperature-sensitive and controlled by a single-nucleotide polymorphism. Plant Mol Biol, 1999, 40: 719–727
Palusa S G, Ali G S, Reddy A S N. Alternative splicing of premRNAs of Arabidopsis serine/arginine-rich proteins: regulation by hormones and stresses. Plant J, 2007, 49: 1091–1107
Yost H J, Petersen R B, Lindquist S. RNA metabolism: Strategies for regulation in the heat shock response. Trends Genet, 1990, 6: 223–227
Shin C, Manley J L. The SR protein SRp38 represses splicing in M phase cells. Cell, 2002, 111: 407–417
Shin C, Feng Y, Manley J L. Dephosphorylated SRp38 acts as a splicing repressor in response to heat shock. Nature, 2002, 427: 553–558
Shi Y S, Manley J L. A complex signaling pathway regulates SRp38 phosphorylation and pre-mRNA splicing in response to heat shock. Mol Cell, 2007, 28: 79–90
Feng Y, Chen M, Manley J L. Phosphorylation switches the general splicing repressor SRp38 to a sequence-specific activator. Nat Struct Mol Biol, 2008, 15: 1040–1048
Jiang S Y, Cai M, Ramachandran S. ORYZA SATIVA MYOSIN XI B controls pollen development by photoperiod-sensitive protein localizations. Dev Biol, 2007, 304: 579–592
Author information
Authors and Affiliations
Corresponding author
Additional information
Supported by the National Key Basic Research and Development Program of China (Grant No. 2007CB108705) and the National Natural Science Foundation of China (Grant No. 30700448)
About this article
Cite this article
Chen, R., Pan, Y., Wang, Y. et al. Temperature-sensitive splicing is an important molecular regulation mechanism of thermosensitive genic male sterility in rice. Chin. Sci. Bull. 54, 2354–2362 (2009). https://doi.org/10.1007/s11434-009-0349-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11434-009-0349-2