Abstract
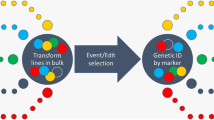
Gene-edited pigs for agricultural and biomedical applications are typically generated using somatic cell nuclear transfer (SCNT). However, SCNT requires the use of monoclonal cells as donors, and the time-consuming and laborious monoclonal selection process limits the production of large populations of gene-edited animals. Here, we developed a rapid and efficient method named RE-DSRNP (reporter RNA enriched dual-sgRNA/CRISPR-Cas9 ribonucleoproteins) for generating gene-edited donor cells. RE-DSRNP takes advantage of the precise and efficient editing features of dual-sgRNA and the high editing efficiency, low off-target effects, transgene-free nature, and low cytotoxic characteristics of reporter RNA enriched RNPs (CRISPR-Cas9 ribonucleoproteins), thus eliminating the need for the selection of monoclonal cells and thereby greatly reducing the generation time of donor cells from 3–4 weeks to 1 week, while also reducing the extent of apoptosis and chromosomal aneuploidy of donor cells. We applied RE-DSRNP to produce cloned pigs bearing a deletion edit of the wild-type p53-induced phosphatase 1 (WIP1) gene: among 32 weaned cloned pigs, 31 (97%) carried WIP1 edits, and 15 (47%) were homozygous for the designed fragment deletion, and no off-target event was detected. The WIP1 knockout (KO) pigs exhibited male reproductive disorders, illustrating the utility of RE-DSRNP for rapidly generating precisely edited animals for functional genomics and disease research. RE-DSRNP’s strong editing performance in a large animal and its marked reduction in the required time for producing SCNT donor cells support its application prospects for rapidly generating populations of transgene-free cloned animals.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Cho, S.J., Cha, B.S., Kwon, O.S., Lim, J., Shin, D.M., Han, D.W., Ishitani, T., Jho, E.H., Fornace, A.J., and Cha, H.J. (2017). Wip1 directly dephosphorylates NLK and increases Wnt activity during germ cell development. Biochim Biophys Acta 1863, 1013–1022.
Cong, L., Ran, F.A., Cox, D., Lin, S., Barretto, R., Habib, N., Hsu, P.D., Wu, X., Jiang, W., Marraffini, L.A., et al. (2013). Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823.
Du, X., Guo, Z., Fan, W., Hai, T., Gao, F., Li, P., Qin, Y., Chen, C., Han, Z., Ren, J., et al. (2021). Establishment of a humanized swine model for COVID-19. Cell Discov 7, 70.
Fan, Z., Liu, Z., Xu, K., Wu, T., Ruan, J., Zheng, X., Bao, S., Mu, Y., Sonstegard, T., and Li, K. (2021). Long-term, multidomain analyses to identify the breed and allelic effects in MSTN-edited pigs to overcome lameness and sustainably improve nutritional meat production. Sci China Life Sci doi: https://doi.org/10.1007/s11427-020-1927-9.
Filipponi, D., Muller, J., Emelyanov, A., and Bulavin, D.V. (2013). Wip1 controls global heterochromatin silencing via ATM/BRCA1-dependent DNA methylation. Cancer Cell 24, 528–541.
Hai, T., Teng, F., Guo, R., Li, W., and Zhou, Q. (2014). One-step generation of knockout pigs by zygote injection of CRISPR/Cas system. Cell Res 24, 372–375.
Kim, S., Kim, D., Cho, S.W., Kim, J., and Kim, J.S. (2014). Highly efficient RNA-guided genome editing in human cells via delivery of purified Cas9 ribonucleoproteins. Genome Res 24, 1012–1019.
Lamas-Toranzo, I., Guerrero-Sánchez, J., Miralles-Bover, H., Alegre-Cid, G., Pericuesta, E., and Bermejo-Álvarez, P. (2017). CRISPR is knocking on barn door. Reprod Dom Anim 52, 39–47.
Li, G., Liu, Y.G., and Chen, Y. (2019). Genome-editing technologies: the gap between application and policy. Sci China Life Sci 62, 1534–1538.
Li, X., Tremoleda, J.L., and Allen, W.R. (2003). Effect of the number of passages of fetal and adult fibroblasts on nuclear remodelling and first embryonic division in reconstructed horse oocytes after nuclear transfer. Reproduction 125, 535–542.
Li, X., Yang, Y., Bu, L., Guo, X., Tang, C., Song, J., Fan, N., Zhao, B., Ouyang, Z., Liu, Z., et al. (2014). Rosa26-targeted swine models for stable gene over-expression and Cre-mediated lineage tracing. Cell Res 24, 501–504.
Liang, Z., Chen, K., Li, T., Zhang, Y., Wang, Y., Zhao, Q., Liu, J., Zhang, H., Liu, C., Ran, Y., et al. (2017). Efficient DNA-free genome editing of bread wheat using CRISPR/Cas9 ribonucleoprotein complexes. Nat Commun 8, 14261.
Liang, Z., Chen, K., Zhang, Y., Liu, J., Yin, K., Qiu, J.L., and Gao, C. (2018). Genome editing of bread wheat using biolistic delivery of CRISPR/Cas9 in vitro transcripts or ribonucleoproteins. Nat Protoc 13, 413–430.
Liang, Z., Wu, Y., Guo, Y., Liu, Y., Ma, L., and Wu, Y. (2021). Bifunctional selection markers assist segregation of transgene-free, genome-edited mutants. Sci China Life Sci 64, 1567–1570.
Liu, Q., Jiao, X., Meng, X., Wang, C., Xu, C., Tian, Z., Xie, C., Li, G., Li, J., Yu, H., et al. (2021). FED: a web tool for foreign element detection of genome-edited organism. Sci China Life Sci 64, 167–170.
Magnani, L., Lee, K., Fodor, W.L., Machaty, Z., and Cabot, R.A. (2008). Developmental capacity of porcine nuclear transfer embryos correlate with levels of chromatin-remodeling transcripts in donor cells. Mol Reprod Dev 75, 766–776.
Mastromonaco, G.F., Perrault, S.D., Betts, D.H., and King, W.A. (2006). Role of chromosome stability and telomere length in the production of viable cell lines for somatic cell nuclear transfer. BMC Dev Biol 6, 41.
Norris, A.L., Lee, S.S., Greenlees, K.J., Tadesse, D.A., Miller, M.F., and Lombardi, H.A. (2020). Template plasmid integration in germline genome-edited cattle. Nat Biotechnol 38, 163–164.
Ramakrishna, S., Kwaku Dad, A.B., Beloor, J., Gopalappa, R., Lee, S.K., and Kim, H. (2014). Gene disruption by cell-penetrating peptide-mediated delivery of Cas9 protein and guide RNA. Genome Res 24, 1020–1027.
Ruan, D., Peng, J., Wang, X., Ouyang, Z., Zou, Q., Yang, Y., Chen, F., Ge, W., Wu, H., Liu, Z., et al. (2018). XIST derepression in active X chromosome hinders pig somatic cell nuclear transfer. Stem Cell Rep 10, 494–508.
Subburaj, S., Chung, S.J., Lee, C., Ryu, S.M., Kim, D.H., Kim, J.S., Bae, S., and Lee, G.J. (2016). Site-directed mutagenesis in Petunia×hybrida protoplast system using direct delivery of purified recombinant Cas9 ribonucleoproteins. Plant Cell Rep 35, 1535–1544.
Svitashev, S., Schwartz, C., Lenderts, B., Young, J.K., and Mark Cigan, A. (2016). Genome editing in maize directed by CRISPR-Cas9 ribonucleoprotein complexes. Nat Commun 7, 13274.
Wei, Y., Gao, Q., Niu, P., Xu, K., Qiu, Y., Hu, Y., Liu, S., Zhang, X., Yu, M., Liu, Z., et al. (2019). Integrative proteomic and phosphoproteomic profiling of testis from Wip1 phosphatase-knockout mice: insights into mechanisms of reduced fertility. Mol Cell Proteomics 18, 216–230.
Whitworth, K.M., Rowland, R.R.R., Ewen, C.L., Trible, B.R., Kerrigan, M. A., Cino-Ozuna, A.G., Samuel, M.S., Lightner, J.E., McLaren, D.G., Mileham, A.J., et al. (2016). Gene-edited pigs are protected from porcine reproductive and respiratory syndrome virus. Nat Biotechnol 34, 20–22.
Woo, J.W., Kim, J., Kwon, S.I., Corvalán, C., Cho, S.W., Kim, H., Kim, S. G., Kim, S.T., Choe, S., and Kim, J.S. (2015). DNA-free genome editing in plants with preassembled CRISPR-Cas9 ribonucleoproteins. Nat Biotechnol 33, 1162–1164.
Xu, K., Zhou, Y., Mu, Y., Liu, Z., Hou, S., Xiong, Y., Fang, L., Ge, C., Wei, Y., Zhang, X., et al. (2020). CD163 and pAPN double-knockout pigs are resistant to PRRSV and TGEV and exhibit decreased susceptibility to PDCoV while maintaining normal production performance. eLife 9, e57132.
Yan, S., Tu, Z., Liu, Z., Fan, N., Yang, H., Yang, S., Yang, W., Zhao, Y., Ouyang, Z., Lai, C., et al. (2018). A Huntingtin knockin pig model recapitulates features of selective neurodegeneration in Huntington’s disease. Cell 173, 989–1002.e13.
Yang, L., Güell, M., Niu, D., George, H., Lesha, E., Grishin, D., Aach, J., Shrock, E., Xu, W., Poci, J., et al. (2015). Genome-wide inactivation of porcine endogenous retroviruses (PERVs). Science 350, 1101–1104.
Zheng, Q., Lin, J., Huang, J., Zhang, H., Zhang, R., Zhang, X., Cao, C., Hambly, C., Qin, G., Yao, J., et al. (2017). Reconstitution of UCP1 using CRISPR/Cas9 in the white adipose tissue of pigs decreases fat deposition and improves thermogenic capacity. Proc Natl Acad Sci USA 114, E9474–E9482.
Zhou, X., Xin, J., Fan, N., Zou, Q., Huang, J., Ouyang, Z., Zhao, Y., Zhao, B., Liu, Z., Lai, S., et al. (2015). Generation of CRISPR/Cas9-mediated gene-targeted pigs via somatic cell nuclear transfer. Cell Mol Life Sci 72, 1175–1184.
Acknowledgements
This work was supported by the National Natural Science Foundation of China (32072690), the Major Scientific Research Tasks for Scientific and Technological Innovation Projects of the Chinese Academy of Agricultural Sciences (CAAS-ZDRW202006), the Science, Technology and Innovation Commission of Shenzhen Municipality (JCKYZDKY202009), the National Transgenic Breeding Project (2016ZX08010-004), the National Transgenic Breeding Project (2016ZX08006-001), and the Agricultural Science and Technology Innovation Program (ASTIP-IAS05).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Compliance and ethics The author(s) declare that they have no conflict of interest.
Electronic Supplementary Material
Rights and permissions
About this article
Cite this article
Xu, K., Zhang, X., Liu, Z. et al. A transgene-free method for rapid and efficient generation of precisely edited pigs without monoclonal selection. Sci. China Life Sci. 65, 1535–1546 (2022). https://doi.org/10.1007/s11427-021-2058-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11427-021-2058-2