Abstract
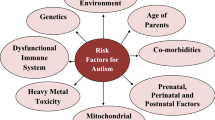
Autism spectrum disorder (ASD) is a spectral neurodevelopment disorder affecting approximately 1% of the population. ASD is characterized by impairments in reciprocal social interaction, communication deficits and restricted patterns of behavior. Multiple factors, including genetic/genomic, epigenetic/epigenomic and environmental, are thought to be necessary for autism development. Recent reviews have provided further insight into the genetic/genomic basis of ASD. It has long been suspected that epigenetic mechanisms, including DNA methylation, chromatin structures and long non-coding RNAs may play important roles in the pathology of ASD. In addition to genetic/genomic alterations and epigenetic/epigenomic influences, environmental exposures have been widely accepted as an important role in autism etiology, among which immune dysregulation and gastrointestinal microbiota are two prominent ones.
Article PDF
Similar content being viewed by others
References
Rosti RO, Sadek AA, Vaux KK, Gleeson JG. The genetic landscape of autism spectrum disorders. Dev Med Child Neurol, 2014: 56: 12–18
Devlin B, Scherer SW. Genetic architecture in autism spectrum disorder. Curr Opin Genet Dev, 2012, 22: 229–237
Huguet G, Ey E, Bourgeron T. The genetic landscapes of autism spectrum disorders. Annu Rev Genom Hum G, 2013, 14: 191–213
Shishido E, Aleksic B, Ozaki N. Copy-number variation in the pathogenesis of autism spectrum disorder. Psychiat Clin Neuros, 2014, 68: 85–95
Betancur C. Etiological heterogeneity in autism spectrum disorders: more than 100 genetic and genomic disorders and still counting. Brain Res, 2011, 1380: 42–77
Veltman JA, Brunner HG. De novo mutations in human genetic disease. Nat Rev Genet, 2012, 13: 565–575
Roberts JL, Hovanes K, Dasouki M, Manzardo AM, Butler MG. Chromosomal microarray analysis of consecutive individuals with autism spectrum disorders or learning disability presenting for genetic services. Gene, 2014, 535: 70–78
Levy D, Ronemus M, Yamrom B, Lee YH, Leotta A, Kendall J. Rare de novo and transmitted copy-number variation in autistic spectrum disorders. Neuron, 2011, 70: 886–897
Stobbe G, Liu YJ, Wu R, Hudgings LH, Thompson O, Hisama FM. Diagnostic yield of array comparative genomic hybridization in adults with autism spectrum disorders. Genet Med, 2014, 16: 70–77
Shen, Y, Dies, KA, Holm IA, Bridgemohan C, Sobeih MM, Caronna EB, Miller KJ, Frazier JA, Silverstein I, Picker J, Weissman L, Raffalli P, Jeste S, Demmer LA, Peters HK, Brewster SJ, Kowalczyk SJ, Rosen-Sheidley B, McGowan C, Duda III AW, Lincoln SA, Lowe KR, Schonwald A, Robbins M, Hisama F, Wolff R, Becker R, Nasir R, Urion DK, Milunsky JM, Rappaport L, Gusella JF, Walsh CA, Wu BL, Miller DT on behalf of the Autism Consortium Clinical Genetics/ DNA Diagnostics Collaboration. Clinical genetic testing for patients with autism spectrum disorders. Pediatrics, 2010, 125: 727–735
Girirajan S, Dennis Megan Y, Baker C, Malig M, Coe Bradley P, Campbell Catarina D. Refinement and discovery of new hotspots of copy-number variation associated with autism spectrum disorder. Am J Hum Genet, 2013, 92: 221–237
Liao HM, Gau SS, Tsai WC, Fang JS, Su YC, Chou MC, Liu SK, Chou WJ, Wu YY, Chen CH. Chromosomal abnormalities in patients with autism spectrum disorders from Taiwan. Am J Med Genet B, 2013, 162B: 734–741
Girirajan S, Johnson RL, Tassone F, Balciuniene J, Katiyar N, Fox K, Baker C, Srikanth A, Yeoh KH, Khoo SJ, Nauth TB, Hansen R, Ritchie M, Hertz-Picciotto I, Eichler EE, Pessah IN, Selleck SB. Global increases in both common and rare copy number load associated with autism. Hum Mol Genet, 2013, 22: 2870–2880
Gilman SR, Iossifov I, Levy D, Ronemus M, Wigler M, Vitkup D. Rare de novo variants associated with autism implicate a large functional network of genes involved in formation and function of synapses. Neuron, 2011, 70: 898–907
O'Roak BJ, Vives L, Fu W, Egertson JD, Stanaway IB, Phelps IG, Carvill G, Kumar A, Lee C, Ankenman K, Munson J, Hiatt JB, Turner EH, Levy R, O'Day DR, Krumm N, Coe BP, Martin BK, Borenstein E, Nickerson DA, Mefford HC, Doherty D, Akey JM, Bernier R, Eichler EE, Shendure J. Multiplex targeted sequencing identifies recurrently mutated genes in autism spectrum disorders. Science, 2012, 338: 1619–1622
O'Roak BJ, Vives L, Girirajan S, Karakoc E, Krumm N, Coe BP, Levy R, Ko A, Lee C, Smith JD, Turner EH, Stanaway IB, Vernot B, Malig M, Baker C, Reilly B, Akey JM, Borenstein E, Rieder MJ, Nickerson DA, Bernier R, Shendure J, Eichler EE. Sporadic autism exomes reveal a highly interconnected protein network of de novo mutations. Nature, 2012, 485: 246–250
Kong A, Frigge ML, Masson G, Besenbacher S, Sulem P, Magnusson G, Gudjonsson SA, Sigurdsson A, Jonasdottir A, Jonasdottir A, Wong WS, Sigurdsson G, Walters GB, Steinberg S, Helgason H, Thorleifsson G, Gudbjartsson DF, Helgason A, Magnusson OT, Thorsteinsdottir U, Stefansson K. Rate of de novo mutations and the importance of father's age to disease risk. Nature, 2012, 488: 471–475
Sanders SJ, Murtha MT, Gupta AR, Murdoch JD, Raubeson MJ, Willsey AJ, Ercan-Sencicek AG, DiLullo NM, Parikshak NN, Stein JL, Walker MF, Ober GT, Teran NA, Song Y, El-Fishawy P, Murtha RC, Choi M, Overton JD, Bjornson RD, Carriero NJ, Meyer KA, Bilguvar K, Mane SM, Sestan N, Lifton RP, Günel M, Roeder K, Geschwind DH, Devlin B, State MW. De novo mutations revealed by whole-exome sequencing are strongly associated with autism. Nature, 2012, 485: 237–241
Iossifov I, Ronemus M, Levy D, Wang Z, Hakker I, Rosenbaum J, Yamrom B, Lee YH, Narzisi G, Leotta A, Kendall J, Grabowska E, Ma B, Marks S, Rodgers L, Stepansky A, Troge J, Andrews P, Bekritsky M, Pradhan K, Ghiban E, Kramer M, Parla J, Demeter R, Fulton LL, Fulton RS, Magrini VJ, Ye K, Darnell JC, Darnell RB, Mardis ER, Wilson RK, Schatz MC, McCombie WR, Wigler M. De novo gene disruptions in children on the autistic spectrum. Neuron, 2012, 74: 285–299
Krumm N, O'Roak BJ, Shendure J, Eichler EE. A de novo convergence of autism genetics and molecular neuroscience. Trends Neurosci, 2014, 37: 95–105
Yu TW, Chahrour MH, Coulter ME, Jiralerspong S, Okamura-Ikeda K, Ataman B, Schmitz-Abe K, Harmin DA, Adli M, Malik AN, D'Gama AM, Lim ET, Sanders SJ, Mochida GH, Partlow JN, Sunu CM, Felie JM, Rodriguez J, Nasir RH, Ware J, Joseph RM, Hill RS, Kwan BY, Al-Saffar M, Mukaddes NM, Hashmi A, Balkhy S, Gascon GG, Hisama FM, LeClair E, Poduri A, Oner O, Al-Saad S, Al-Awadi SA, Bastaki L, Ben-Omran T, Teebi AS, Al-Gazali L, Eapen V, Stevens CR, Rappaport L, Gabriel SB, Markianos K, State MW, Greenberg ME, Taniguchi H, Braverman NE, Morrow EM, Walsh CA. Using whole-exome sequencing to identify inherited causes of autism. Neuron, 2013, 77: 259–273
Lee H, Lin MC, Kornblum HI, Papazian DM, Nelson SF. Exome sequencing identifies de novo gain of function missense mutation in KCND2 in identical twins with autism and seizures that slows potassium channel inactivation. Hum Mol Genetics, 2014, 5: 5
Matsunami N, Hensel CH, Baird L, Stevens J, Otterud B, Leppert T, Varvil T, Hadley D, Glessner JT, Pellegrino R, Kim C, Thomas K, Wang F, Otieno FG, Ho K, Christensen GB, Li D, Prekeris R, Lambert CG, Hakonarson H, Leppert MF. Identification of rare DNA sequence variants in high-risk autism families and their prevalence in a large case/control population. Mol Autism, 2014, 5: 5
Cukier HN, Dueker ND, Slifer SH, Lee JM, Whitehead PL, Lalanne E, Leyva N, Konidari I, Gentry RC, Hulme WF, Booven DV, Mayo V, Hofmann NK, Schmidt MA, Martin ER, Haines JL, Cuccaro ML, Gilbert JR, Pericak-Vance MA. Exome sequencing of extended families with autism reveals genes shared across neurodevelopmental and neuropsychiatric disorders. Mol Autism, 2014, 5: 1
Shi LL, Zhang X, Golhar R, Otieno FG, He MZ, Hou CP, Kim C, Keating B, Lyon GJ, Wang K, Hakonarson H. Whole-genome sequencing in an autism multiplex family. Mol Autism, 2013, 4: 8
Xia K, Guo H, Hu Z, Xun G, Zuo L, Peng Y, Wang K, He Y, Xiong Z, Sun L, Pan Q, Long Z, Zou X, Li X, Li W, Xu X, Lu L, Liu Y, Hu Y, Tian D, Long L, Ou J, Liu Y, Li X, Zhang L, Pan Y, Chen J, Peng H, Liu Q, Luo X, Su W, Wu L, Liang D, Dai H, Yan X, Feng Y, Tang B, Li J, Miedzybrodzka Z, Xia J, Zhang Z, Luo X, Zhang X, St Clair D, Zhao J, Zhang F. Common genetic variants on 1p13.2 associate with risk of autism. Mol Psychiatry, 2014, 19: 1212–1219
Nijmeijer JS, Arias-Vásquez A, Rommelse NNJ, Altink ME, Buschgens CJM, Fliers EA, Franke B, Minderaa RB, Sergeant JA, Buitelaar JK, Hoekstra PJ, Hartman CA. Quantitative linkage for autism spectrum disorders symptoms in attention-deficit/ hyperactivity disorder: significant locus on chromosome 7q11. J Autism Dev Disord. 2014, 44: 1671–1680
Kovac J, Macedoni Lukšic M, Trebušak Podkrajšek K, Klancar G, Battelino T. Rare single nucleotide polymorphisms in the regulatory regions of the superoxide dismutase genes in autism spectrum disorder. Autism Res, 2014, 7: 138–144
Weiss LA, Arking DE, Gene Discovery Project of Johns H, the Autism C, Daly MJ, Chakravarti A. A genome-wide linkage and association scan reveals novel loci for autism. Nature, 2009, 461: 802–808
Glessner JT, Wang K, Cai G, Korvatska O, Kim CE, Wood S, Estes A, Brune CW, Bradfield JP, Imielinski M, Frackelton EC, Reichert J, Crawford EL, Munson J, Sleiman PM, Chiavacci R, Annaiah K, Thomas K, Hou C, Glaberson W, Flory J, Otieno F, Garris M, Soorya L, Klei L, Piven J, Meyer KJ, Anagnostou E, Sakurai T, Game RM, Rudd DS, Zurawiecki D, McDougle CJ, Davis LK, Miller J, Posey DJ, Michaels S, Kolevzon A, Silverman JM, Bernier R, Levy SE, Schultz RT, Dawson G, Owley T, McMahon WM, Wassink TH, Sweeney JA, Nurnberger JI, Coon H, Sutcliffe JS, Minshew NJ, Grant SF, Bucan M, Cook EH, Buxbaum JD, Devlin B, Schellenberg GD, Hakonarson H. Autism genome-wide copy number variation reveals ubiquitin and neuronal genes. Nature, 2009, 459: 569–573
Pinto D, Pagnamenta AT, Klei L, Anney R, Merico D, Regan R, Conroy J, Magalhaes TR, Correia C, Abrahams BS, Almeida J, Bacchelli E, Bader GD, Bailey AJ, Baird G, Battaglia A, Berney T, Bolshakova N, Bölte S, Bolton PF, Bourgeron T, Brennan S, Brian J, Bryson SE, Carson AR, Casallo G, Casey J, Chung BH, Cochrane L, Corsello C, Crawford EL, Crossett A, Cytrynbaum C, Dawson G, de Jonge M, Delorme R, Drmic I, Duketis E, Duque F, Estes A, Farrar P, Fernandez BA, Folstein SE, Fombonne E, Freitag CM, Gilbert J, Gillberg C, Glessner JT, Goldberg J, Green A, Green J, Guter SJ, Hakonarson H, Heron EA, Hill M, Holt R, Howe JL, Hughes G, Hus V, Igliozzi R, Kim C, Klauck SM, Kolevzon A, Korvatska O, Kustanovich V, Lajonchere CM, Lamb JA, Laskawiec M, Leboyer M, Le Couteur A, Leventhal BL, Lionel AC, Liu XQ, Lord C, Lotspeich L, Lund SC, Maestrini E, Mahoney W, Mantoulan C, Marshall CR, McConachie H, McDougle CJ, McGrath J, McMahon WM, Merikangas A, Migita O, Minshew NJ, Mirza GK, Munson J, Nelson SF, Noakes C, Noor A, Nygren G, Oliveira G, Papanikolaou K, Parr JR, Parrini B, Paton T, Pickles A, Pilorge M, Piven J, Ponting CP, Posey DJ, Poustka A, Poustka F, Prasad A, Ragoussis J, Renshaw K, Rickaby J, Roberts W, Roeder K, Roge B, Rutter ML, Bierut LJ, Rice JP, Salt J, Sansom K, Sato D, Segurado R, Sequeira AF, Senman L, Shah N, Sheffield VC, Soorya L, Sousa I, Stein O, Sykes N, Stoppioni V, Strawbridge C, Tancredi R, Tansey K, Thiruvahindrapduram B, Thompson AP, Thomson S, Tryfon A, Tsiantis J, van Engeland H, Vincent JB, Volkmar F, Wallace S, Wang K, Wang Z, Wassink TH, Webber C, Weksberg R, Wing K, Wittemeyer K, Wood S, Wu J, Yaspan BL, Zurawiecki D, Zwaigenbaum L, Buxbaum JD, Cantor RM, Cook EH, Coon H, Cuccaro ML, Devlin B, Ennis S, Gallagher L, Geschwind DH, Gill M, Haines JL, Hallmayer J, Miller J, Monaco AP, Nurnberger JI Jr, Paterson AD, Pericak-Vance MA, Schellenberg GD, Szatmari P, Vicente AM, Vieland VJ, Wijsman EM, Scherer SW, Sutcliffe JS, Betancur C. Functional impact of global rare copy number variation in autism spectrum disorders. Nature, 2010, 466: 368–372
Chow ML, Pramparo T, Winn ME, Barnes CC, Li HR, Weiss L, Fan JB, Murray S, April C, Belinson H, Fu XD, Wynshaw-Boris A, Schork NJ, Courchesne E. Age-dependent brain gene expression and copy number anomalies in autism suggest distinct pathological processes at young versus mature ages. PLoS Genet, 2012, 8: e1002592
Sakai Y, Shaw CA, Dawson BC, Dugas DV, Al-Mohtaseb Z, Hill DE, Zoghbi HY. Protein interactome reveals converging molecular pathways among autism disorders. Sci Transl Med, 2011, 3: 86ra49
Voineagu I, Wang X, Johnston P, Lowe JK, Tian Y, Horvath S, Mill J, Cantor RM, Blencowe BJ, Geschwind DH. Transcriptomic analysis of autistic brain reveals convergent molecular pathology. Nature, 2011, 474: 380–384
Sweatt JD. The emerging field of neuroepigenetics. Neuron, 2013, 80: 624–632
LaSalle JM, Powell WT, Yasui DH. Epigenetic layers and players underlying neurodevelopment. Trends Neurosci, 2013, 36: 460–470
Lalande M, Calciano MA. Molecular epigenetics of Angelman syndrome. Cell Mol Life Sci, 2007, 64: 947–960
Graff J, Mansuy IM. Epigenetic codes in cognition and behaviour. Behav Brain Res, 2008, 192: 70–87
Badcock C. The imprinted brain: how genes set the balance between autism and psychosis. Epigenomics, 2011, 3: 345–359
Houston I, Peter CJ, Mitchell A, Straubhaar J, Rogaev E, Akbarian S. Epigenetics in the human brain. Neuropsychopharmacology, 2013, 38: 183–197
Nagarajan RP, Patzel KA, Martin M, Yasui DH, Swanberg SE, Hertz-Picciotto I, Hansen RL, van de Water J, Pessah IN, Jiang R, Robinson WP, LaSalle JM. MECP2 promoter methylation and X chromosome inactivation in autism. Autism Res, 2008, 1: 169–178
LaSalle JM, Yasui DH. Evolving role of MeCP2 in Rett syndrome and autism. Epigenomics, 2009, 1: 119–130
Harony-Nicolas H, Mamrut S, Brodsky L, Shahar-Gold H, Barki-Harrington L, Wagner S. Brain region-specific methylation in the promoter of the murine oxytocin receptor gene is involved in its expression regulation. Psychoneuroendocrinology, 2014, 39: 121–131
Zhu L, Wang X, Li XL, Towers A, Cao X, Wang P, Bowman R, Yang H, Goldstein J, Li YJ, Jiang YH. Epigenetic dysregulation of SHANK3 in brain tissues from individuals with autism spectrum disorders. Human Mol Genet, 2014, 23: 1563–1578
Vazna A, Musova Z, Vlckova M, Novotna D, Dvorakova L, Hrdlicka M, Havlovicova M, Sedlacek Z. FMR1 gene expansion, large deletion of Xp, and skewed X-inactivation in a girl with mental retardation and autism. Am J Med Genet A, 2010, 152a: 1273–1277
Nguyen A, Rauch TA, Pfeifer GP, Hu VW. Global methylation profiling of lymphoblastoid cell lines reveals epigenetic contributions to autism spectrum disorders and a novel autism candidate gene, RORA, whose protein product is reduced in autistic brain. FASEB J, 2010, 24: 3036–3051
Ladd-Acosta C, Hansen KD, Briem E, Fallin MD, Kaufmann WE, Feinberg AP. Common DNA methylation alterations in multiple brain regions in autism. Mol Psychiatry, 2014, 19: 862–871
Wong CC, Meaburn EL, Ronald A, Price TS, Jeffries AR, Schalkwyk LC, Plomin R, Mill J. Methylomic analysis of monozygotic twins discordant for autism spectrum disorder and related behavioural traits. Mol Psychiatry, 2014, 19: 495–503
Cheung I, Shulha HP, Jiang Y, Matevossian A, Wang J, Weng Z, Akbarian S. Developmental regulation and individual differences of neuronal H3K4me3 epigenomes in the prefrontal cortex. Proc Natl Acad Sci USA, 2010, 107: 8824–8829
de la Torre-Ubieta L, Bonni A. Combinatorial assembly of neurons: from chromatin to dendrites. Trends Cell Biol, 2008, 18: 48–51
Ronan JL, Wu W, Crabtree GR. From neural development to cognition: unexpected roles for chromatin. Nat Rev Genet, 2013, 14: 347–359
Sandner G, Host L, Angst MJ, Guiberteau T, Guignard B, Zwiller J. The HDAC inhibitor phenylbutyrate reverses effects of neonatal ventral hippocampal lesion in rats. Front Psychiatry, 2011, 1: 153
Adegbola A, Gao H, Sommer S, Browning M. A novel mutation in JARID1C/SMCX in a patient with autism spectrum disorder (ASD). Am J Med Genet A, 2008, 146A: 505–511
Goncalves TF, Goncalves AP, Fintelman Rodrigues N, Dos Santos JM, Pimentel MM, Santos-Reboucas CB. KDM5C mutational screening among males with intellectual disability suggestive of X-Linked inheritance and review of the literature. Eur J Med Genet, 2014, 57: 138–144
Halgren C, Kjaergaard S, Bak M, Hansen C, El-Schich Z, Anderson CM, Henriksen KF, Hjalgrim H, Kirchhoff M, Bijlsma EK, Nielsen M, den Hollander NS, Ruivenkamp CA, Isidor B, Le Caignec C, Zannolli R, Mucciolo M, Renieri A, Mari F, Anderlid BM, Andrieux J, Dieux A, Tommerup N, Bache I. Corpus callosum abnormalities, intellectual disability, speech impairment, and autism in patients with haploinsufficiency of ARID1B. Clin Genet, 2012, 82: 248–255
Neale BM, Kou Y, Liu L, Ma'ayan A, Samocha KE, Sabo A, Lin CF, Stevens C, Wang LS, Makarov V, Polak P, Yoon S, Maguire J, Crawford EL, Campbell NG, Geller ET, Valladares O, Schafer C, Liu H, Zhao T, Cai G, Lihm J, Dannenfelser R, Jabado O, Peralta Z, Nagaswamy U, Muzny D, Reid JG, Newsham I, Wu Y, Lewis L, Han Y, Voight BF, Lim E, Rossin E, Kirby A, Flannick J, Fromer M, Shakir K, Fennell T, Garimella K, Banks E, Poplin R, Gabriel S, DePristo M, Wimbish JR, Boone BE, Levy SE, Betancur C, Sunyaev S, Boerwinkle E, Buxbaum JD, Cook EH Jr, Devlin B, Gibbs RA, Roeder K, Schellenberg GD, Sutcliffe JS, Daly MJ. Patterns and rates of exonic de novo mutations in autism spectrum disorders. Nature, 2012, 485: 242–245
Talkowski ME, Rosenfeld JA, Blumenthal I, Pillalamarri V, Chiang C, Heilbut A, Ernst C, Hanscom C, Rossin E, Lindgren AM, Pereira S, Ruderfer D, Kirby A, Ripke S, Harris DJ, Lee JH, Ha K, Kim HG, Solomon BD, Gropman AL, Lucente D, Sims K, Ohsumi TK, Borowsky ML, Loranger S, Quade B, Lage K, Miles J, Wu BL, Shen Y, Neale B, Shaffer LG, Daly MJ, Morton CC, Gusella JF. Sequencing chromosomal abnormalities reveals neurodevelopmental loci that confer risk across diagnostic boundaries. Cell, 2012, 149: 525–537
Jiang YH, Yuen RK, Jin X, Wang M, Chen N, Wu X, Ju J, Mei J, Shi Y, He M, Wang G, Liang J, Wang Z, Cao D, Carter MT, Chrysler C, Drmic IE, Howe JL, Lau L, Marshall CR, Merico D, Nalpathamkalam T, Thiruvahindrapuram B, Thompson A, Uddin M, Walker S, Luo J, Anagnostou E, Zwaigenbaum L, Ring RH, Wang J, Lajonchere C, Wang J, Shih A, Szatmari P, Yang H, Dawson G, Li Y, Scherer SW. Detection of clinically relevant genetic variants in autism spectrum disorder by whole-genome sequencing. Am J Hum Genet, 2013, 93: 249–263
Tahiliani M, Mei P, Fang R, Leonor T, Rutenberg M, Shimizu F, Li J, Rao A, Shi Y. The histone H3K4 demethylase SMCX links REST target genes to X-linked mental retardation. Nature, 2007, 447: 601–605
Ben-David E, Shifman S. Combined analysis of exome sequencing points toward a major role for transcription regulation during brain development in autism. Mol Psychiatry, 2013, 18: 1054–1056
Wang KC, Yang YW, Liu B, Sanyal A, Corces-Zimmerman R, Chen Y, Lajoie BR, Protacio A, Flynn RA,Gupta RA, Wysocka J, Lei M, Dekker J, Helms JA, Chang HY. A long noncoding RNA maintains active chromatin to coordinate homeotic gene expression. Nature, 2011, 472: 120–124
Magistri M, Faghihi MA, St Laurent G, 3rd, Wahlestedt C. Regulation of chromatin structure by long noncoding RNAs: focus on natural antisense transcripts. Trends Genet, 2012, 28: 389–396
Ziats MN, Rennert OM. Aberrant expression of long noncoding RNAs in autistic brain. J Mol Neurosci, 2013, 49: 589–593
Boulanger LM. Immune proteins in brain development and synaptic plasticity. Neuron, 2009, 64: 93–109
Huh GS, Boulanger LM, Du H, Riquelme PA, Brotz TM, Shatz CJ. Functional requirement for class I MHC in CNS development and plasticity. Science, 2000, 290: 2155–2159
Glynn MW, Elmer BM, Garay PA, Liu XB, Needleman LA, El-Sabeawy F, McAllister AK. MHCI negatively regulates synapse density during the establishment of cortical connections. Nat Neurosci, 2011, 14: 442–451
Elmer BM, Estes ML, Barrow SL, McAllister AK. MHCI requires MEF2 transcription factors to negatively regulate synapse density during development and in disease. J Neurosci, 2013, 33: 13791–13804
Stevens B, Allen NJ, Vazquez LE, Howell GR, Christopherson KS, Nouri N, Micheva KD, Mehalow AK, Huberman AD, Stafford B, Sher A, Litke AM, Lambris JD, Smith SJ, John SW, Barres BA. The classical complement cascade mediates CNS synapse elimination. Cell, 2007, 131: 1164–1178
Kaneko M, Stellwagen D, Malenka RC, Stryker MP. Tumor necrosis factor-alpha mediates one component of competitive, experiencedependent plasticity in developing visual cortex. Neuron, 2008, 58: 673–680
Onore C, Careaga M, Ashwood P. The role of immune dysfunction in the pathophysiology of autism. Brain Behav Immun, 2012, 26: 383–392
Croen LA, Braunschweig D, Haapanen L, Yoshida CK, Fireman B, Grether JK, Kharrazi M, Hansen RL, Ashwood P, van de Water J. Maternal mid-pregnancy autoantibodies to fetal brain protein: the early markers for autism study. Biol Psychiatry, 2008, 64: 583–588
Patterson PH. Immune involvement in schizophrenia and autism: etiology, pathology and animal models. Behav Brain Res, 2009, 204: 313–321
Diamond B, Huerta PT, Mina-Osorio P, Kowal C, Volpe BT. Losing your nerves? Maybe it's the antibodies. Nat Rev Immunol, 2009, 9: 449–456
Braunschweig D, Ashwood P, Krakowiak P, Hertz-Picciotto I, Hansen R, Croen LA, Pessah IN, van de Water J. Autism: maternally derived antibodies specific for fetal brain proteins. Neurotoxicology, 2008, 29: 226–231
Bauman MD, Iosif AM, Ashwood P, Braunschweig D, Lee A, Schumann CM, van de Water J, Amaral DG. Maternal antibodies from mothers of children with autism alter brain growth and social behavior development in the rhesus monkey. Transl Psychiatry, 2013, 3: e278
Li X, Chauhan A, Sheikh AM, Patil S, Chauhan V, Li XM, Brown T, Malik M. Elevated immune response in the brain of autistic patients. J Neuroimmunol, 2009, 207: 111–116
Morgan JT, Chana G, Pardo CA, Achim C, Semendeferi K, Buckwalter J, Courchesne E, Everall IP. Microglial activation and increased microglial density observed in the dorsolateral prefrontal cortex in autism. Biol Psychiatry, 2010, 68: 368–376
Tambuyzer BR, Ponsaerts P, Nouwen EJ. Microglia: gatekeepers of central nervous system immunology. J Leukoc Biol, 2009, 85: 352–370
Tremblay ME, Stevens B, Sierra A, Wake H, Bessis A, Nimmerjahn A. The role of microglia in the healthy brain. J Neurosci, 2011, 31: 16064–16069
Suzuki K, Sugihara G, Ouchi Y, Nakamura K, Futatsubashi M, Takebayashi K, Yoshihara Y, Omata K, Matsumoto K, Tsuchiya KJ, Iwata Y, Tsujii M, Sugiyama T, Mori N. Microglial activation in young adults with autism spectrum disorder. JAMA Psychiatry, 2013, 70: 49–58
Ehninger D, Sano Y, de Vries PJ, Dies K, Franz D, Geschwind DH, Kaur M,Lee YS,Li W,Lowe JK,Nakagawa JA,Sahin M,Smith K,Whittemore V, Silva AJ. Gestational immune activation and Tsc2 haploinsufficiency cooperate to disrupt fetal survival and may perturb social behavior in adult mice. Mol Psychiatry, 2012, 17: 62–70
de Theije CG, Wopereis H, Ramadan M, van Eijndthoven T, Lambert J, Knol J, Garssen J, Kraneveld AD, Oozeer R. Altered gut microbiota and activity in a murine model of autism spectrum disorders. Brain Behav Immun, 2014, 37: 197–206
Coury DL, Ashwood P, Fasano A, Fuchs G, Geraghty M, Kaul A, Mawe G, Patterson P, Jones NE. Gastrointestinal conditions in children with autism spectrum disorder: developing a research agenda. Pediatrics, 2012, 130 Suppl 2: S160–S168
Chaidez V, Hansen RL, Hertz-Picciotto I. Gastrointestinal problems in children with autism, developmental delays or typical development. J Autism Dev Disord, 2014, 44: 1117–1127
Williams BL, Hornig M, Buie T, Bauman ML, Cho Paik M, Wick I, Bennett A, Jabado O, Hirschberg DL, Lipkin WI. Impaired carbohydrate digestion and transport and mucosal dysbiosis in the intestines of children with autism and gastrointestinal disturbances. PLoS One, 2011, 6: e24585
Walker SJ, Fortunato J, Gonzalez LG, Krigsman A. Identification of unique gene expression profile in children with regressive autism spectrum disorder (ASD) and ileocolitis. PLoS One, 2013, 8: e58058
Wang Y, Kasper LH. The role of microbiome in central nervous system disorders. Brain Behav Immun, 2014, 38C: 1–12
Cryan JF, Dinan TG. Mind-altering microorganisms: the impact of the gut microbiota on brain and behaviour. Nat Rev Neurosci, 2012, 13: 701–712
Desbonnet L, Clarke G, Shanahan F, Dinan TG, Cryan JF. Microbiota is essential for social development in the mouse. Mol Psychiatry, 2014, 19: 146–148
Hsiao EY, McBride SW, Hsien S, Sharon G, Hyde ER, McCue T, Codelli JA, Chow J, Reisman SE, Petrosino JF, Patterson PH, Mazmanian SK. Microbiota modulate behavioral and physiological abnormalities associated with neurodevelopmental disorders. Cell, 2013, 155: 1451–1463
Author information
Authors and Affiliations
Corresponding author
Additional information
Contributed equally to this work
This article is published with open access at link.springer.com
Electronic supplementary material
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (https://creativecommons.org/licenses/by/4.0), which permits use, duplication, adaptation, distribution, and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Yu, L., Wu, Y. & Wu, BL. Genetic architecture, epigenetic influence and environment exposure in the pathogenesis of Autism. Sci. China Life Sci. 58, 958–967 (2015). https://doi.org/10.1007/s11427-015-4941-1
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11427-015-4941-1