Abstract
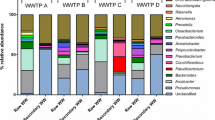
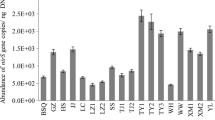
Bacteria play an important role in pollutant transformation in activated sludge-based wastewater treatment plants (WWTPs). Exploring the microbial community structure and diversity is essential to improving the performance of wastewater treatment processes. This study employed Illumina MiSeq high-throughput sequencing to investigate the microbial community composition and diversity in a cattle farm wastewater treatment plant (Cf-WWTP). The results showed that the dominant phyla in the whole process were Proteobacteria, Bacteroidetes, and Firmicutes. The principal coordinate analysis (PCoA) indicated that the different stages had a significant impact on the microbial community structure; Bacteroidetes was the dominant phylum in the anearobic stage and Proteobacteria was the dominant phylum in the anoxic-oxic stage. Redundancy analysis (RDA) revealed that total phosphorus (TP) was the most significant factor that regulated the microbial community composition, followed by chemical oxygen demand (COD), total nitrogen (TN), and pH. Proteobacteria, Patescibacteria, and Chloroflexi were simultaneously negatively correlated with TN, COD, and TP. Nitrogen metabolic pathway and transformation mechanism was elucidated by a complete denitrification function predicted with phylogenetic investigation of communities with reconstruction of unobserved states (PICRUSt), as well as detection of ammonia-oxidizing bacteria (AOB) and nitrite-oxidizing bacteria (NOB). These results provide new insights into our understanding of microbial community and metabolic functions of Cf-WWTP.






Similar content being viewed by others
Data availability
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
APHA (2017) Standard methods for the examination of water and wastewater, 23rd edn. American Public Health Association, Washington, DC
Bai F, Tian H, Ma J (2020) Landfill leachate treatment through the combination of genetically engineered bacteria Rhodococcus erythropolis expressing Nirs and AMO and membrane filtration processes. Environ Pollut 263:114061. https://doi.org/10.1016/j.envpol.2020.114061
Brailo M, Schreier HJ, McDonald R, Marsic-Lucic J, Gavrilovic A, Pecarevic M, Jug-Dujakovic J (2019) Bacterial community analysis of marine recirculating aquaculture system bioreactors for complete nitrogen removal established from a commercial inoculum. Aquaculture 503:198–206. https://doi.org/10.1016/j.aquaculture.2018.12.078
Chen JL, Xu YB, Li YX, Liao JS, Ling JY, Li JY, Xie GY (2019) Effective removal of nitrate by denitrification re-enforced with a two-stage anoxic/oxic (A/O) process from a digested piggery wastewater with a low C/N ratio. J Environ Manag 240:19–26. https://doi.org/10.1016/j.jenvman.2019.03.091
Fridrich B, Krcmar D, Dalmacija B, Molnar J, Pesic V, Kragulj M, Varga N (2014) Impact of wastewater from pig farm lagoons on the quality of local groundwater. Agric Water Manag 135:40–53. https://doi.org/10.1016/j.agwat.2013.12.014
Greay TL, Gofton AW, Zahedi A, Paparini A, Linge KL, Joll CA, Ryan UM (2019) Evaluation of 16S next-generation sequencing of hypervariable region 4 in wastewater samples: an unsuitable approach for bacterial enteric pathogen identification. Sci Total Environ 670:1111–1124. https://doi.org/10.1016/j.scitotenv.2019.03.278
Haller L, Tonolla M, Zopfi J, Peduzzi R, Wildi W, Pote J (2011) Composition of bacterial and archaeal communities in freshwater sediments with different contamination levels (Lake Geneva, Switzerland). Water Res 45:1213–1228. https://doi.org/10.1016/j.watres.2010.11.018
He LY, He LK, Liu YS, Zhang M, Zhao JL, Zhang QQ, Ying GG (2019) Microbial diversity and antibiotic resistome in swine farm environments. Sci Total Environ 685:197–207. https://doi.org/10.1016/j.scitotenv.2019.05.369
Huang F, Pan L, He Z, Zhang M, Zhang M (2020) Identification, interactions, nitrogen removal pathways and performances of culturable heterotrophic nitrification-aerobic denitrification bacteria from mariculture water by using cell culture and metagenomics. Sci Total Environ 732:139268. https://doi.org/10.1016/j.scitotenv.2020.139268
Jenkins D, Richard MG, Daigger GT (2003) Manual on the causes and control of activated sludge bulking, foaming, and other solids separation problems. CRC Press, Boca Raton. https://doi.org/10.1201/9780203503157
Jia LP, Jiang BH, Huang F, Hu XM (2019) Nitrogen removal mechanism and microbial community changes of bioaugmentation subsurface wastewater infiltration system. Bioresour Technol 294:122140. https://doi.org/10.1016/j.biortech.2019.122140
Jin T, Zhang T, Yan QM (2010) Characterization and quantification of ammonia-oxidizing archaea (AOA) and bacteria (AOB) in a nitrogen-removing reactor using T-RFLP and qPCR. Appl Microbiol Biotechnol 87:1167–1176. https://doi.org/10.1007/s00253-010-2595-2
Keshri J, Ram AP, Sime-Ngando T (2018) Distinctive patterns in the taxonomical resolution of bacterioplankton in the sediment and pore waters of contrasted freshwater lakes. Microb Ecol 75:662–673. https://doi.org/10.1007/s00248-017-1074-z
Kim DJ, Kim SH (2006) Effect of nitrite concentration on the distribution and competition of nitrite-oxidizing bacteria in nitratation reactor systems and their kinetic characteristics. Water Res 40:887–894. https://doi.org/10.1016/j.watres.2005.12.023
Li M, Cao HL, Hong YG, Gu JD (2011) Spatial distribution and abundances of ammonia-oxidizing archaea (AOA) and ammonia-oxidizing bacteria (AOB) in mangrove sediments. Appl Microbiol Biotechnol 89:1243–1254. https://doi.org/10.1007/s00253-010-2929-0
Lie AAY, Liu Z, Hu SK, Jones AC, Kim DY, Countway PD, Amaral-Zettler LA, Cary SC, Sherr EB, Sherr BF, Gast RJ, Caron DA (2014) Investigating microbial eukaryotic diversity from a global census: insights from a comparison of pyrotag and full-length sequences of 18S rRNA genes. Appl Environ Microbiol 80:4363–4373. https://doi.org/10.1128/AEM.00057-14
Liu N, Xie X, Yang F, Shen L, Yu C, Liu J (2016) Bioaugmentation with microorganisms for hydrolysis acidification treatment of dyeing wastewater. Chinese J Environ Eng 10:2245–2251
Ma Q, Qu Y, Shen W, Zhang Z, Wang J, Liu Z, Li D, Li H, Zhou J (2015) Bacterial community compositions of coking wastewater treatment plants in steel industry revealed by Illumina high-throughput sequencing. Bioresour Technol 179:436–443. https://doi.org/10.1016/j.biortech.2014.12.041
McAteer PG, Christine Trego A, Thorn C, Mahony T, Abram F, O'Flaherty V (2020) Reactor configuration influences microbial community structure during high-rate, low-temperature anaerobic treatment of dairy wastewater. Bioresour Technol 307:123221. https://doi.org/10.1016/j.biortech.2020.123221
Miura Y, Hiraiwa MN, Ito T, Itonaga T, Watanabe Y, Okabe S (2007) Bacterial community structures in MBRs treating municipal wastewater: relationship between community stability and reactor performance. Water Res 41:627–637. https://doi.org/10.1016/j.watres.2006.11.005
Moura RB, Santos CED, Okada DY, Martins TH, Ferraz Junior ADN, Damianovic M, Foresti E (2018) Carbon-nitrogen removal in a structured-bed reactor (SBRRIA) treating sewage: operating conditions and metabolic perspectives. J Environ Manag 224:19–28. https://doi.org/10.1016/j.jenvman.2018.07.014
Mulkerrins D, O'Connor E, Lawlee B, Barton P, Dobson A (2004) Assessing the feasibility of achieving biological nutrient removal from wastewater at an Irish food processing factory. Bioresour Technol 91:207–214. https://doi.org/10.1016/s0960-8524(03)00173-1
Nakasaki K, Nguyen KK, Ballesteros FC, Maekawa T, Koyama M (2020) Characterizing the microbial community involved in anaerobic digestion of lipid-rich wastewater to produce methane gas. Anaerobe 61:102082. https://doi.org/10.1016/j.anaerobe.2019.102082
Pan KL, Gao JF, Li HY, Fan XY, Li DC, Jiang H (2018) Ammonia-oxidizing bacteria dominate ammonia oxidation in a full-scale wastewater treatment plant revealed by DNA-based stable isotope probing. Bioresour Technol 256:152–159. https://doi.org/10.1016/j.biortech.2018.02.012
Reddy MV, Mohan SV (2012) Effect of substrate load and nutrients concentration on the polyhydroxyalkanoates (PHA) production using mixed consortia through wastewater treatment. Bioresour Technol 114:573–582. https://doi.org/10.1016/j.biortech.2012.02.127
Soares-Castro P, Yadav TC, Viggor S, Kivisaar M, Kapley A, Santos PM (2019) Seasonal bacterial community dynamics in a crude oil refinery wastewater treatment plant. Appl Microbiol Biotechnol 103:9131–9141. https://doi.org/10.1007/s00253-019-10130-8
Sorokin DY, Vejmelkova D, Lucker S, Streshinskaya GM, Rijpstra WIC, Damste JSS, Kleerbezem R, van Loosdrecht M, Muyzer G, Daims H (2014) Nitrolancea hollandica gen. nov., sp nov., a chemolithoautotrophic nitrite-oxidizing bacterium isolated from a bioreactor belonging to the phylum Chloroflexi. Int J Syst Evol Microbiol 64:1859–1865. https://doi.org/10.1099/ijs.0.062232-0
Stazi V, Tomei MC (2018) Enhancing anaerobic treatment of domestic wastewater: State of the art, innovative technologies and future perspectives. Sci Total Environ 635:78–91. https://doi.org/10.1016/j.scitotenv.2018.04.071
Wang P, Wu D, You X, Su Y, Xie B (2021) Antibiotic and metal resistance genes are closely linked with nitrogen-processing functions in municipal solid waste landfills. J Hazard Mater 403:123689. https://doi.org/10.1016/j.jhazmat.2020.123689
Wu D, Chen G, Zhang X, Yang K, Xie B (2017) Change in microbial community in landfill refuse contaminated with antibiotics facilitates denitrification more than the increase in ARG over long-term. Sci Rep 7:41230. https://doi.org/10.1038/srep41230
Xu HZ, Pei HY, Jin Y, Ma CX, Wang YT, Sun JM, Li HM (2018a) High-throughput sequencing reveals microbial communities in drinking water treatment sludge from six geographically distributed plants, including potentially toxic cyanobacteria and pathogens. Sci Total Environ 634:769–779. https://doi.org/10.1016/j.scitotenv.2018.04.008
Xu S, Yao JQ, Ainiwaer M, Hong Y, Zhang YJ (2018b) Analysis of bacterial community structure of activated sludge from wastewater treatment plants in winter. Biomed Res Int 2018:8278970–8278978. https://doi.org/10.1155/2018/8278970
Xu P, Xiao E, He F, Xu D, Zhang Y, Wang Y, Wu Z (2019) High performance of integrated vertical-flow constructed wetland for polishing low C/N ratio river based on a pilot-scale study in Hangzhou, China. Environ Sci Pollut Res 26:22431–22449. https://doi.org/10.1007/s11356-019-05508-0
Yuan K, Li SQ, Zhong F (2020) Treatment of coking wastewater in biofilm-based bioaugmentation process: biofilm formation and microbial community analysis. J Hazard Mater 400:123117. https://doi.org/10.1016/j.jhazmat.2020.123117
Zhang B, Yu QW, Yan GQ, Zhu HB, Xu XY, Zhu L (2018) Seasonal bacterial community succession in four typical wastewater treatment plants: correlations between core microbes and process performance. Sci Rep 8:4566. https://doi.org/10.1038/s41598-018-22683-1
Zhang L, Shen Z, Fang WK, Gao G (2019) Composition of bacterial communities in municipal wastewater treatment plant. Sci Total Environ 689:1181–1191. https://doi.org/10.1016/j.scitotenv.2019.06.432
Zhao WH, Peng YZ, Wang MX, Huang Y, Li XY (2019) Nutrient removal and microbial community structure variation in the two-sludge system treating low carbon/nitrogen domestic wastewater. Bioresour Technol 294:122161. https://doi.org/10.1016/j.biortech.2019.122161
Zheng DC, Gu WZ, Zhou QM, Zhang LX, Wei CC, Yang QZM, Li DP (2020) Ammonia oxidation and denitrification in a bio-anode single -chambered microbial electrolysis cell. Bioresour Technol 310:123466. https://doi.org/10.1016/j.biortech.2020.123466
Funding
This work was financially supported by the National Key Research and Development Program of China (2018YFC1901000), Fujian Province and Chinese Academy of Sciences STS Project (2019T3027), and Shanghai Engineering Research Center of Biotransformation of Organic Solid Waste Program (SERC2020A03).
Author information
Authors and Affiliations
Contributions
Weizhi Yan: writing, formal analysis; Na Wang: investigation; Dong Wei: investigation; Chengyu Liang: investigation; Xiaomiao Chen: visualization; Li Liu: conceptualization, funding acquisition; Jiping Shi: supervision, funding acquisition.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Responsible Editor: Diane Purchase
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
ESM 1
(DOCX 63210 kb)
Rights and permissions
About this article
Cite this article
Yan, W., Wang, N., Wei, D. et al. Bacterial community compositions and nitrogen metabolism function in a cattle farm wastewater treatment plant revealed by Illumina high-throughput sequencing. Environ Sci Pollut Res 28, 40895–40907 (2021). https://doi.org/10.1007/s11356-021-13570-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-021-13570-w