Abstract
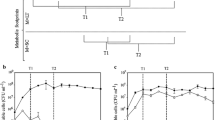
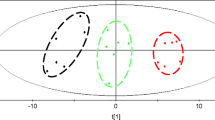
The intracellular metabolic profile characterization of Saccharomyces cerevisiae throughout industrial ethanol fermentation was investigated using gas chromatography coupled to time-of-flight mass spectrometry. A total of 143 and 128 intracellular metabolites in S. cerevisiae were detected and quantified in continuous and batch fermentations, respectively. The two fermentation processes were both clearly distinguished into three main phases by principal components analysis. Furthermore, the levels of some metabolites involved in central carbon metabolism varied significantly throughout both processes. Glycerol and phosphoric acid were principally responsible for discriminating seed, main and final phases of continuous fermentation, while lactic acid and glycerol contributed mostly to telling different phases of batch fermentation. In addition, the levels of some amino acids such as glycine varied significantly during both processes. These findings provide new insights into the metabolomic characteristics during industrial ethanol fermentation processes.







Similar content being viewed by others
References
Abrams, R., Hammarsten, E., & Shemin, D. (1948). Glycine as a precursor of purines in yeast. The Journal of Biological Chemistry, 173, 429.
Backhus, L. E., DeRisi, J. P. O., & Bisson, L. F. (2001). Functional genomic analysis of a commercial wine strain of Saccharomyces cerevisiae under differing nitrogen conditions. FEMS Yeast Research, 1, 111–125. doi:10.1111/j.1567-1364.2001.tb00022.x.
Blomberg, A., & Adler, L. (1989). Roles of glycerol and glycerol-3-phosphate dehydrogenase (NAD+) in acquired osmotolerance of Saccharomyces cerevisiae. Journal of Bacteriology, 171, 1087–1092.
Blomberg, A., & Adler, L. (1992). Physiology of osmotolerance in fungi. Advances in Microbial Physiology, 33, 145–212. doi:10.1016/S0065-2911(08)60217-9.
Brejning, J., Arneborg, N., & Jespersen, L. (2005). Identification of genes and proteins induced during the lag and early exponential phase of lager brewing yeasts. Journal of Applied Microbiology, 98, 261–271. doi:10.1111/j.1365-2672.2004.02472.x.
Brewster, J. L., De Valoir, T., Dwyer, N. D., Winter, E., & Gustin, M. C. (1993). An osmosensing signal transduction pathway in yeast. Science, 259, 1760–1763. doi:10.1126/science.7681220.
Cavalieri, D., Townsend, J. P., & Hartl, D. L. (2000). Manifold anomalies in gene expression in a vineyard isolate of Saccharomyces cerevisiae revealed by DNA microarray analysis. Proceedings of the National Academy of Sciences of the United States of America, 97, 12369–12374. doi:10.1073/pnas.210395297.
Cheng, J. S., Qiao, B., & Yuan, Y. J. (2008). Comparative proteome analysis of robust Saccharomyces cerevisiae insights into industrial continuous and batch fermentation. Applied Microbiology and Biotechnology, 81, 327–338. doi:10.1007/s00253-008-1733-6.
Davenport, K. R., Sohaskey, M., Kamada, Y., Levin, D. E., & Gustin, M. C. (1995). A second osmosensing signal transduction pathway in yeast: Hypotonic shock activates the PKC1 protein kinase-regulated cell integrity pathway. The Journal of Biological Chemistry, 270, 30157–30161. doi:10.1074/jbc.270.50.30157.
Devantier, R., Scheithauer, B., Villas-Bôas, S. G., Pedersen, S., & Olsson, L. (2005). Metabolite profiling for analysis of yeast stress response during very high gravity ethanol fermentations. Biotechnology and Bioengineering, 90, 703–714. doi:10.1002/bit.20457.
Dickinson, J. R., & Scheweizer, M. (1999). Stress responses in the metabolism and molecular physiology of Saccharomyces cerevisiae (p. 343). New York: Taylor and Francis.
Fiehn, O. (2002). Metabolomics—the link between genotypes and phenotypes. Plant Molecular Biology, 48, 155–171. doi:10.1023/A:1013713905833.
Fiehn, O., Kopka, J., Dormann, P., Altmann, T., Trethewey, R. N., & Willmitzer, L. (2000). Metabolite profiling for plant functional genomics. Nature Biotechnology, 18, 1157–1161. doi:10.1038/81137.
Franzen, C. J. (2003). Metabolic flux analysis of RQ-controlled microaerobic ethanol production by Saccharomyces cerevisiae. Yeast (Chichester, England), 20, 117–132. doi:10.1002/yea.956.
Gancedo, C., Gancedo, J. M., & Sols, A. (1968). Glycerol metabolism in yeasts Pathways of utilization and production. European Journal of Biochemistry, 5, 165–172. doi:10.1111/j.1432-1033.1968.tb00353.x.
Gika, H. G., Theodoridis, G. A., & Wilson, I. D. (2008). Liquid chromatography and ultra-performance liquid chromatography–mass spectrometry fingerprinting of human urine: Sample stability under different handling and storage conditions for metabonomics studies. Journal of Chromatography A, 1189, 314–322. doi:10.1016/j.chroma.2007.10.066.
Hansen, R., Pearson, S. Y., Brosnan, J. M., Meaden, P. G., & Jamieson, D. J. (2006). Proteomic analysis of a distilling strain of Saccharomyces cerevisiae during industrial grain fermentation. Applied Microbiology and Biotechnology, 72, 116–125. doi:10.1007/s00253-006-0508-1.
Hohl, L. A., & Joslyn, M. A. (1941). Lactic acid formation in alcoholic fermentation by yeast. Plant Physiology, 16, 343–360.
Hohmann, S. (2002). Osmotic stress signaling and osmoadaptation in yeasts. Microbiology and Molecular Biology Reviews, 66, 300–372. doi:10.1128/MMBR.66.2.300-372.2002.
James, T. C., Campbell, S., Donnelly, D., & Bond, U. (2003). Transcription profile of brewery yeast under fermentation conditions. Journal of Applied Microbiology, 94, 432–448. doi:10.1046/j.1365-2672.2003.01849.x.
Kamada, Y., Jung, U. S., Piotrowski, J., & Levin, D. E. (1995). The protein kinase C-activated MAP kinase pathway in Saccharomyces cerevisiae mediates a novel aspect of the heat shock response. Genes & Development, 9, 1559–1571. doi:10.1101/gad.9.13.1559.
Kell, B., & Westerhoff, H. V. (1986). Metabolic control theory: Its role in microbiology and biotechnology. FEMS Microbiology Letters, 39, 305–320. doi:10.1111/j.1574-6968.1986.tb01863.x.
Kobi, D., Zugmeyer, S., Potier, S., & Jaquet-Gutfreund, L. (2004). Two dimensional protein map of an “ale”-brewing yeast strain: Proteome dynamics during fermentation. FEMS Yeast Research, 5, 213–230. doi:10.1016/j.femsyr.2004.07.004.
Koning, W., & Van Dam, K. (1992). A method for the determination of changes of glycolytic metabolites in yeast on a subsecond time scale using extraction at neutral pH. Analytical Biochemistry, 204, 118–123. doi:10.1016/0003-2697(92)90149-2.
Krebs, H. A., Lowenstein, J. M., & Greenberg, D. M. (1960). Metabolic pathways (Vol. 1, p. 129). New York/London: Academic Press.
Lenz, E., & Wilson, I. D. (2007). Analytical strategies in metabonomics. Journal of Proteome Research, 6, 443. doi:10.1021/pr0605217.
Major, H. J., Williams, R., Wilson, A. J., & Wilson, I. D. (2006). A metabonomic analysis of plasma from Zucker rat strains using gas chromatography/mass spectrometry and pattern recognition. Rapid Communications in Mass Spectrometry, 20, 3295–3302. doi:10.1002/rcm.2732.
Marks, V. D., Ho Sui, S. J., Erasmus, D., Van der Merwe, G. K., et al. (2008). Dynamics of the yeast transcriptome during wine fermentation reveals a novel fermentation stress response. FEMS Yeast Research, 8, 35–52.
Mohler, R. E., Tu, B. P., Dombek, K. M., Hoggard, J. C., Young, E. T., & Synovec, R. E. (2008). Identification and evaluation of cycling yeast metabolites in two-dimensional comprehensive gas chromatography-time-of-flight-mass spectrometry data. Journal of Chromatography A, 1186, 401–411. doi:10.1016/j.chroma.2007.10.063.
Nissen, T. L., Schulze, U., Nielsen, J., & Villadsen, J. (1997). Flux distributions in anaerobic, glucose-limited continuous cultures of Saccharomyces cerevisiae. Microbiology, 143, 203–218.
Ossipov, V., Ossipova, S., Bykov, V., Oksanen, E., Koricheva, J., & Haukioja, E. (2008). Application of metabolomics to genotype and phenotype discrimination of birch trees grown in a long-term open-field experiment. Metabolomics, 4, 39–51. doi:10.1007/s11306-007-0097-8.
Pham, T. K., Chong, P. K., Gan, C. S., & Wright, P. C. (2006). Proteomic analysis of Saccharomyces cerevisiae under high gravity fermentation conditions. Journal of Proteome Research, 5, 3411–3419. doi:10.1021/pr060377p.
Roessner, U., Luedemann, A., Brust, D., et al. (2001). Metabolic profiling allows comprehensive phenotyping of genetically or environmentally modified plant systems. The Plant Cell, 13, 11–29.
Roessner, U., Wagner, C., Kopka, J., Trethewey, R. N., & Willmitzer, L. (2000). Simultaneous analysis of metabolites in potato tuber by gas chromatography–mass spectrometry. The Plant Journal, 23, 131–142. doi:10.1046/j.1365-313x.2000.00774.x.
Trabalzini, L., Paffetti, A., Scaloni, A., et al. (2003). Proteomic response to physiological fermentation stresses in a wild-type wine strain of Saccharomyces cerevisiae. The Biochemical Journal, 370, 35–46. doi:10.1042/BJ20020140.
Van Dijken, J. P., & Scheffers, W. A. (1986). Redox balances in the metabolism of sugars by yeast. FEMS Microbiology Reviews, 32, 199–224.
Villas-Bôas, S. G., Delicado, D. G., Akesson, M., & Nielsen, J. (2003). Simultaneous analysis of amino and nonamino organic acids as methyl chloroformate derivatives using gas chromatography–mass spectrometry. Analytical Biochemistry, 322, 134–138. doi:10.1016/j.ab.2003.07.018.
Villas-Bôas, S. G., Moxley, J. F., Akesson, M., Stephanopoulos, G., & Nielsen, J. (2005). High-throughput metabolic state analysis: The missing link in integrated functional genomics of yeasts. The Biochemical Journal, 388, 669–677. doi:10.1042/BJ20041162.
Wagner, C., Sefkow, M., & Kopka, J. (2003). Construction and application of a mass spectral and retention time index database generated from GC/EI-TOF-MS metabolite profiles. Phytochemistry, 62, 887–900. doi:10.1016/S0031-9422(02)00703-3.
Wiebe, M. G., Rintala, E., Tamminen, A., et al. (2008). Central carbon metabolism of Saccharomyces cerevisiae in anaerobic, oxygen-limited and fully aerobic steady-state conditions and following a shift to anaerobic conditions. FEMS Yeast Research, 8, 140–154.
Yang, W. L., & Bernards, M. A. (2007). Metabolite profiling of potato (Solanum tuberosum L.) tubers during wound-induced suberization. Metabolomics, 3, 147–159. doi:10.1007/s11306-007-0053-7.
Acknowledgments
The authors are grateful for the financial support from the National Natural Science Foundation of China (Key Program Grant No.20736006), the National Basic Research Program of China (Program: 2007CB714301), and Key Projects in the National Science & Technology Pillar Program (No.2007BAD42B02), and the National Natural Science Foundation of China (No. 20706044).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ding, MZ., Cheng, JS., Xiao, WH. et al. Comparative metabolomic analysis on industrial continuous and batch ethanol fermentation processes by GC-TOF-MS. Metabolomics 5, 229–238 (2009). https://doi.org/10.1007/s11306-008-0145-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11306-008-0145-z