Abstract
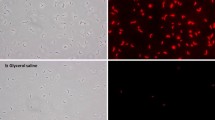
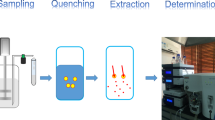
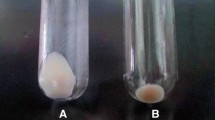
Effective and rapid inactivation of cellular metabolism is a prerequisite for accurate metabolome analysis. Cold methanol quenching is commonly applied to stop any metabolic activity and, at the same time remaining the cells’ integrity. However, it is reported that especially prokaryotic cells like Escherichia coli and Corynebacterium glutamicum tend to leak intracellular metabolites during cold methanol quenching. In this work leakage of adenylates is quantified for different quenching fluids. Further, a methanol/glycerol based quenching fluid is proposed, which reduces leakage drastically compared to the commonly applied methanol/water solution (16% ATP leakage compared to more than 70%).







Similar content being viewed by others
References
Al Zaid Siddiquee, K., Arauzo-Bravo, M. J., & Shimizu, K. (2004). Metabolic flux analysis of pykF gene knockout Escherichia coli based on 13C-labeling experiments together with measurements of enzyme activities and intracellular metabolite concentrations. Applied Microbiology and Biotechnology, 63, 407–417. doi:10.1007/s00253-003-1357-9.
Bolten, C. J., Kiefer, P., Letisse, F., Portais, J. C., & Wittmann, C. (2007). Sampling for metabolome analysis of microorganisms. Analytical Chemistry, 79, 3843–3849. doi:10.1021/ac0623888.
Buchholz, A., Takors, R., & Wandrey, C. (2001). Quantification of intracellular metabolites in Escherichia coli K12 using liquid chromatographic-electrospray ionization tandem mass spectrometric techniques. Analytical Biochemistry, 295, 129–137. doi:10.1006/abio.2001.5183.
Buziol, S., Bashir, I., Baumeister, A., Claassen, W., Noisommit-Rizzi, N., Mailinger, W., et al. (2002). New bioreactor-coupled rapid stopped-flow sampling technique for measurements of metabolite dynamics on a subsecond time scale. Biotechnology and Bioengineering, 80, 632–636. doi:10.1002/bit.10427.
Faijes, M., Mars, A. E., & Smid, E. J. (2007). Comparison of quenching and extraction methodologies for metabolome analysis of Lactobacillus plantarum. Microbial Cell Factories, 6, 27. doi:10.1186/1475-2859-6-27.
Fell, D. A. (1992). Metabolic control analysis: A survey of its theoretical and experimental development. The Biochemical Journal, 286, 313–330.
Hiller, J., Franco-Lara, E., Papaioannou, V., & Weuster-Botz, D. (2007). Fast sampling and quenching procedures for microbial metabolic profiling. Biotechnology Letters, 29, 1161–1167. doi:10.1007/s10529-007-9383-9.
Hoque, M. A., Ushiyama, H., Tomita, M., & Shimizu, K. (2005). Dynamic responses of the intracellular metabolite concentrations of the wild type and pykA mutant Escherichia coli against pulse addition of glucose or NH3 under those limiting continuous cultures. Biochemical Engineering Journal, 26, 38–49. doi:10.1016/j.bej.2005.05.012.
Jensen, N. B., Jokumsen, K. V., & Villadsen, J. (1999). Determination of the phosphorylated sugars of the Embden–Meyerhoff–Parnas pathway in Lactococcus lactis using a fast sampling technique and solid phase extraction. Biotechnology and Bioengineering, 63, 356–362. doi:10.1002/(SICI)1097-0290(19990505)63:3≤356::AID-BIT12≥3.0.CO;2-1.
Jenzsch, M., Gnoth, S., Beck, M., Kleinschmidt, M., Simutis, R., & Lübbert, A. (2006). Open-loop control of the biomass concentration within the growth phase of recombinant protein production processes. Journal of Biotechnology, 127, 84–94. doi:10.1016/j.jbiotec.2006.06.004.
Kacser, H., & Burns, J. A. (1973). The control of flux. Symposia of the Society for Experimental Biology, 27, 65–104.
Link, H., & Weuster-Botz, D. (2007). Steady state analysis of metabolic pathways: Comparing the double modulation method and the lin-log approach. Metabolic Engineering, 9, 433–441. doi:10.1016/j.ymben.2007.07.002.
Luo, B., Groenke, K., Takors, R., Wandrey, C., & Oldiges, M. (2007). Simultaneous determination of multiple intracellular metabolites in glycolysis, pentose phosphate pathway and tricarboxylic acid cycle by liquid chromatography-mass spectrometry. Journal of Chromatography A, 1147, 153–164.
Magnus, J. B., Hollwedel, D., Oldiges, M., & Takors, R. (2006). Monitoring and modeling of the reaction dynamics in the valine/leucine synthesis pathway in Corynebacterium glutamicum. Biotechnology Progress, 22, 1071–1083. doi:10.1021/bp060072f.
Mashego, M. R., Rumbold, K., De Mey, M., Vandamme, E., Soetaert, W., & Heijnen, J. J. (2007). Microbial metabolomics: Past, present and future methodologies. Biotechnology Letters, 29, 1–16. doi:10.1007/s10529-006-9218-0.
Nasution, U., van Gulik, W. M., Kleijn, R. J., van Winden, W. A., Proell, A., & Heijnen, J. J. (2006). Measurement of intracellular metabolites of primary metabolism and adenine nucleotides in chemostat cultivated Penicillium chrysogenum. Biotechnology and Bioengineering, 94, 159–166. doi:10.1002/bit.20842.
Oldiges, M., Lütz, S., Pflug, S., Schroer, K., Stein, N., & Wiendahl, C. (2007). Metabolomics: Current state and evolving methodologies and tools. Applied Microbiology and Biotechnology, 76, 495–511. doi:10.1007/s00253-007-1029-2.
Savageau, M. (1969). Biochemical system analysis, I. Some mathematical properties of the rate law for the component enzymatic reactions. Journal of Theoretical Biology, 25, 365–369. doi:10.1016/S0022-5193(69)80026-3.
Schaefer, U., Boos, W., Takors, R., & Weuster-Botz, D. (1999). Automated sampling device for monitoring intracellular metabolite dynamics. Analytical Biochemistry, 270, 88–96. doi:10.1006/abio.1999.4048.
Schaub, J., Schiesling, C., Reuss, M., & Dauner, M. (2006). Integrated sampling procedure for metabolome analysis. Biotechnology Progress, 22, 1434–1442. doi:10.1021/bp050381q.
Takors, R., Bathe, B., Rieping, M., Hans, S., Kelle, R., & Huthmacher, K. (2007). Systems biology for industrial strains and fermentation processes–example: amino acids. Journal of Biotechnology, 129, 181–190. doi:10.1016/j.jbiotec.2007.01.031.
Vaidyanathan, S. (2005). Profiling microbial metabolomes: What do we stand to gain? Metabolomics, 1, 17–28. doi:10.1007/s11306-005-1104-6.
Vallino, J. J., & Stephanopoulos, G. (1993). Metabolic flux distributions in Corynebacterium glutamicum during growth and lysine overproduction. Biotechnology and Bioengineering, 41, 633–646. doi:10.1002/bit.260410606.
Villas-Bôas, S. G., & Bruheim, P. (2007). Cold glycerol-saline: The promising quenching solution for accurate intracellular metabolite analysis of microbial cells. Analytical Biochemistry, 370, 87–97. doi:10.1016/j.ab.2007.06.028.
Visser, D., van Zuylen, G. A., van Dam, J. C., Oudshoorn, A., Eman, M. R., Ras, C., et al. (2002). Rapid sampling for analysis of in vivo kinetics using the BioScope: A system for continuous-pulse experiments. Biotechnology and Bioengineering, 79, 674–681. doi:10.1002/bit.10328.
Wang, L., & Hatzimanikatis, V. (2006). Metabolic engineering under uncertainty. I: Framework development. Metabolic Engineering, 8, 133–141. doi:10.1016/j.ymben.2005.11.003.
Weuster-Botz, D. (1997). Sampling tube device for monitoring intracellular metabolite dynamics. Analytical Biochemistry, 246, 225–233. doi:10.1006/abio.1997.2009.
Wittmann, C., Krömer, J. O., Kiefer, P., Binz, T., & Heinzle, E. (2004). Impact of the cold shock phenomenon on quantification of intracellular metabolites in bacteria. Analytical Biochemistry, 327, 135–139. doi:10.1016/j.ab.2004.01.002.
Wu, L., Wang, W., van Winden, W. A., van Gulik, W. M., & Heijnen, J. J. (2004). A new framework for the estimation of control parameters in metabolic pathways using lin-log kinetics. European Journal of Biochemistry, 271, 3348–3359. doi:10.1111/j.0014-2956.2004.04269.x.
Acknowledgement
This study is based on the work supported by the Deutsche Forschungsgemeinschaft DFG under Grant No. WE 2715/10-1.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Link, H., Anselment, B. & Weuster-Botz, D. Leakage of adenylates during cold methanol/glycerol quenching of Escherichia coli . Metabolomics 4, 240–247 (2008). https://doi.org/10.1007/s11306-008-0114-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11306-008-0114-6