Abstract
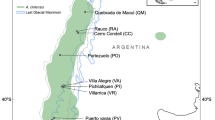
In-depth characterization of the genetic diversity and population structure of wild relatives is of paramount importance for genetic improvement and biodiversity conservation, and is particularly crucial when the wild relatives of crops are endangered. In this study, we sampled the Alpine plum (Briançon apricot) Prunus brigantina Vill. across its natural distribution in the French Alps, where its populations are severely fragmented and its population size strongly impacted by humans. We analysed 71 wild P. brigantina samples with 24 nuclear simple sequence repeat (microsatellite) markers and studied their genetic diversity and population structure, with the aim to inform in situ conservation measures and build a core collection for long-term ex situ conservation. We also examined the genetic relationships of P. brigantina with other species in the Prunophora subgenus, encompassing the Prunus (Eurasian plums), Prunocerasus (North American plums) and Armeniaca (apricots) sections, to check its current taxonomy. We detected a moderate genetic diversity in P. brigantina and a Bayesian model–based clustering approach revealed the existence of three genetically differentiated clusters, endemic to three geographical regions in the Alps, which will be important for in situ conservation measures. Based on genetic diversity and population structure analyses, a subset of 36 accessions were selected for ex situ conservation in a core collection that encompasses the whole detected P. brigantina allelic diversity. Using a dataset of cultivated apricots and wild cherry plums (P. cerasifera) genotyped with the same markers, we detected gene flow neither with European P. armeniaca cultivars nor with diploid plums. Similar to previous studies, dendrograms and networks placed P. brigantina closer to the Armeniaca section than to the Prunus section. Our results thus confirm the classification of P. brigantina within the Armeniaca section; it also illustrates the importance of the sampling size and design in phylogenetic studies.






Similar content being viewed by others
Data availability
The datasets generated by the current study, i.e. the SSR genotyping, are available at the INRAE data portal (https://data.inrae.fr/) where they can be freely retrieved under the link: https://doi.org/10.15454/4DKVVV
References
Allendorf FW, Luikart GH, Aitken SN (2012) Conservation and the genetics of populations, 2nd Edition edn. Wiley-Blackwell Ltd, Hoboken
Branca F, Donnini D (2011) Prunus brigantina. vol e.T172164A121228349. https://doi.org/10.2305/IUCN.UK.2011-1.RLTS.T172164A6840507.en
Chin S-W, Shaw J, Haberle R, Wen J, Potter D (2014) Diversification of almonds, peaches, plums and cherries – molecular systematics and biogeographic history of Prunus (Rosaceae). Mol Phylogenet Evol 76:34–48. https://doi.org/10.1016/j.ympev.2014.02.024
Cici S, Van Acker R (2010) Gene flow in Prunus species in the context of novel trait risk assessment. Environ Biosaf Res 9:75–85. https://doi.org/10.1051/ebr/2010011
Cornille A, Giraud T, Bellard C, Tellier A, le Cam B, Smulders MJM, Kleinschmit J, Roldan-Ruiz I, Gladieux P (2013a) Postglacial recolonization history of the European crabapple (Malus sylvestris Mill.), a wild contributor to the domesticated apple. Mol Ecol 22:2249–2263. https://doi.org/10.1111/mec.12231
Cornille A, Gladieux P, Giraud T (2013b) Crop-to-wild gene flow and spatial genetic structure in the closest wild relatives of the cultivated apple. Evol Appl 6:737–748. https://doi.org/10.1111/eva.12059
Couplan F (2009) Le régal végétal: plantes sauvages comestibles. Collection L’encyclopédie des plantes sauvages comestibles et toxiques de l'Europe.
Coyne JA, Orr HA (2004) Speciation. OUP USA,
Decroocq S, Cornille A, Tricon D, Babayeva S, Chague A, Eyquard JP, Karychev R, Dolgikh S, Kostritsyna T, Liu S, Liu W, Geng W, Liao K, Asma BM, Akparov Z, Giraud T, Decroocq V (2016) New insights into the history of domesticated and wild apricots and its contribution to Plum pox virus resistance. Mol Ecol 25:4712–4729. https://doi.org/10.1111/mec.13772
Doyle JJ (1992) Gene trees and species trees: molecular systematics as one-character taxonomy. Syst Bot 17:144–163. https://doi.org/10.2307/2419070
Dupouy J (1959) Le prunier de Briançon ou Marmottier (Prunus brigantiaca Vill.) et les huiles de marmotte. Bull SAJA 141:69–71
Earl DA, vonHoldt BM (2012) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour 4:359–361. https://doi.org/10.1007/s12686-011-9548-7
Escribano P, Viruel M, Hormaza J (2008) Comparison of different methods to construct a core germplasm collection in woody perennial species with simple sequence repeat markers. A case study in cherimoya (Annona cherimola, Annonaceae), an underutilised subtropical fruit tree species. Ann Appl Biol 153:25–32. https://doi.org/10.1111/j.1744-7348.2008.00232.x
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620. https://doi.org/10.1111/j.1365-294X.2005.02553.x
Fahrig L (2003) Effects of habitat fragmentation on biodiversity. Annu Rev Ecol Evol Syst 34:487–515. https://doi.org/10.1146/annurev.ecolsys.34.011802.132419
Frankel OH, Brown AHD (1983) Current Plant genetic resources today - a critical appraisal. In: Proceedings of the XV International Congress of Genetics. Crop Genetic Resources: Conservation and Evaluation, New Delhi, India. Oxford & IBH publishing CO, Oxford, pp 241–256
Glaszmann JC, Kilian B, Upadhyaya HD, Varshney RK (2010) Accessing genetic diversity for crop improvement. Curr Opin Plant Biol 13:167–173. https://doi.org/10.1016/j.pbi.2010.01.004
Govindaraj M, Vetriventhan M, Srinivasan M (2014) Importance of genetic diversity assessment in crop plants and its recent advances: an overview of its analytical perspectives. Genet Res Int 2015:1–14. https://doi.org/10.1155/2015/431487
Hagen L, Khadari B, Lambert P, Audergon J-M (2002) Genetic diversity in apricot revealed by AFLP markers: species and cultivar comparisons. Theor Appl Genet 105:298–305. https://doi.org/10.1007/s00122-002-0910-8
Heled J, Drummond AJ (2010) Bayesian Inference of species trees from multilocus data. Mol Biol Evol 27:570–580. https://doi.org/10.1093/molbev/msp274
Horvath A, Christmann H, Laigret F (2008) Genetic diversity and relationships among Prunus cerasifera (cherry plum) clones. Botany 86:1311–1318. https://doi.org/10.1139/b08-097
Jakobsson M, Rosenberg NA (2007) CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics 23:1801–1806. https://doi.org/10.1093/bioinformatics/btm233
Jombart T, Ahmed I (2011) Adegenet 1.3-1: new tools for the analysis of genome-wide SNP data. Bioinformatics 27:3070–3071. https://doi.org/10.1093/bioinformatics/btr521
Krüssmann G (1978) Manual of cultivated broad-leaved trees and shrubs, vol 3 Pru-Z. Timber Press, Portland
Layne REC, Sherman WB (1986) Interspecific hybridization of Prunus. HortScience 21:48–51
Lee-Yaw JA, Grassa CJ, Joly S, Andrew RL, Rieseberg LH (2019) An evaluation of alternative explanations for widespread cytonuclear discordance in annual sunflowers (Helianthus). New Phytol 221:515–526. https://doi.org/10.1111/nph.15386
Li DZ, Pritchard HW (2009) The science and economics of ex situ plant conservation. Trends Plant Sci 14:614–621
Ligges U, Mächler M (2003) Scatterplot3d - an R package for visualizing multivariate data. J Stat Softw 8:1–20. https://doi.org/10.17877/de290r-15194
Liu S, Cornille A, Decroocq S, Tricon D, Chague A, Eyquard JP, Liu WS, Giraud T, Decroocq V (2019) The complex evolutionary history of apricots: species divergence, gene flow and multiple domestication events. Mol Ecol 28:5299–5314. https://doi.org/10.1111/mec.15296
Meirmans PG, Van Tienderen PH (2004) Genotype and genodive: two programs for the analysis of genetic diversity of asexual organisms. Mol Ecol Notes 4:792–794. https://doi.org/10.1111/j.1471-8286.2004.00770.x
Noble V, Van Es J, Michaud H, Garraud L (2015) Liste Rouge de la flore vasculaire de Provence-Alpes-Côte d’Azur. Provence-Alpes-Côte d’Azur, France
Numaguchi K, Akagi T, Kitamura Y, Ishikawa R, Ishii T (2020) Interspecific introgression and natural selection in the evolution of Japanese apricot (Prunus mume). bioRxiv:2020.2006.2023.141200. https://doi.org/10.1101/2020.06.23.141200
Peakall R, Smouse P (2012) GenAIEx V5: Genetic analysis in Excel. Populations genetic software for teaching and research. Bioinformat 28:2537–2539. https://doi.org/10.1093/bioinformatics/bts460
Perrier X, Jacquemoud-Collet JP (2006) DARwin software. http://darwin.cirad.fr/.
Pignatti S (1982) Flora d'Italia, vol 2. Edagricole, Bologna
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Reales A, Sargent DJ, Tobutt KR, Rivera D (2010) Phylogenetics of Eurasian plums, Prunus L. section Prunus (Rosaceae), according to coding and non-coding chloroplast DNA sequences. Tree Genet Genomes 6:37–45. https://doi.org/10.1007/s11295-009-0226-9
Rehder A (1940) Manual of cultivated trees and shrubs hardy in North America, Biosystematics, floristic & phylogeny series, vol 1. Collier Macmillan Ltd, New York
Rosenberg NA (2004) DISTRUCT: a program for the graphical display of population structure. Mol Ecol Notes 4:137–138. https://doi.org/10.1046/j.1471-8286.2003.00566.x
Schoen DJ, Brown AHD (1993) Conservation of allelic richness in wild crop relatives is aided by assessment of genetic markers. Proc Natl Acad Sci 90:10623–10627. https://doi.org/10.1073/pnas.90.22.10623
Shannon CE (1948) A mathematical theory of communication. Bell Syst Tech J 27:379–423. https://doi.org/10.1002/j.1538-7305.1948.tb01338.x
Shi S, Li J, Sun J, Yu J, Zhou S (2013) Phylogeny and classification of Prunus sensu lato (Rosaceae). J Integr Plant Biol 55:1069–1079. https://doi.org/10.1111/jipb.12095
Simon CJ, Hannan RM (1995) Development and use of core subsets of cool-season food legume germplasm collections. HortScience 30:907C. https://doi.org/10.21273/hortsci.30.4.907c
Soltis DE, Kuzoff RK (1995) Discordance between nuclear and chloroplast phylogenies in the Heuchera group (Saxifragaceae). Evolution 49:727–742. https://doi.org/10.1111/j.1558-5646.1995.tb02309.x
Szpiech ZA, Jakobsson M, Rosenberg NA (2008) ADZE: a rarefaction approach for counting alleles private to combinations of populations. Bioinformatics 24:2498–2504. https://doi.org/10.1093/bioinformatics/btn478
Takeda T, Shimada T, Nomura K, Ozaki T, Haji T, Yamaguchi M, Yoshida M (1998) Classification of apricot varieties by RAPD analysis. Engei Gakkai Zasshi 67:21–27. https://doi.org/10.2503/jjshs.67.21
Takezaki N, Nei M, Tamura K (2014) POPTREEW: web version of POPTREE for constructing population trees from allele frequency data and computing some other quantities. Mol Biol Evol 31:1622–1624. https://doi.org/10.1093/molbev/msu093
Tison J-M, De Foucault B (2014) Flora Gallica - Flore de France. Biotope, Mèze
Villars D (1786) Histoire des plantes de Dauphiné: Contenant une Préface Historique, un Dictionnaire des Termes de Botanique, les Classes, les Familles, les Genres, & les Herborisations des Environs de Grenoble, de la Grande Chartreuse, de Briançon, de Gap & de Montelimar, vol 1. Prevost, Paris
Welk E, de Rigo D, Caudullo G (2016) Prunus avium in Europe: distribution, habitat, usage and threats. In: San-Miguel-Ayanz J, de Rigo D, Caudullo G, Houston Durrant TAM (eds) European atlas of forest tree species. Publication Office of the European Union, Luxembourg
Wiens JJ, Servedio MR (1997) Accuracy of phylogenetic analysis including and excluding polymorphic characters. Syst Biol 46:332–345. https://doi.org/10.1093/sysbio/46.2.332
Zhebentyayeva T et al (2019) Genetic characterization of worldwide Prunus domestica (plum) germplasm using sequence-based genotyping. Horticult Res 6:12. https://doi.org/10.1038/s41438-018-0090-6
Acknowledgements
Molecular analysis was performed at the GenoToul Get-PlaGe (INRAE Center of Toulouse) and GENTYANE (INRAE Center of Clermont-Ferrand) platforms. The authors wish to acknowledge all the people who helped in collecting the samples, in particular, collaborators from Le Plantivore at Château Ville-vieille, Histoire de Confiture at Plampinet, Nevache, C. Gatineau from Cervières and the managers of the Queyras and Mercantour natural parks. We thank the curators of the French Genetic Resources Centre (Marine Delmas) and of the USDA-ARS National Clonal Germplasm Repository (Davis, California, John Preece). We acknowledge the good and efficient care of the plants at the UMR BFP (Unité Mixte de Recherches Biologie du Fruit et Pathogènes, INRAE) by Jean-Philippe Eyquard and Pascal Briard. Appropriate permissions from responsible authorities for collecting and using Prunus samples from Central Asia and Caucasia were obtained by the local collaborators. The official authorization for the survey and sampling of P. brigantina genetic resources is registered and accessible through the following link: https://absch.cbd.int/database/ABSCH-IRCC-FR-246978. The rest of the samples were kindly provided, with due authorizations, by the curators of the French INRAE Genetic Resources Centre (GRC) and the USDA-ARS National Clonal Germplasm Repository (Davis, California), further details are available on their respective databases.
Funding
S.L. is a recipient of a Chinese Scholarship Council PhD grant. Part of the cost for genotyping and E.H. internship were supported by the PRIMA FREECLIMB project (ANR-18-PRIM-0001-10).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Z. Kaya
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Figure S1.
DeltaK plot as a function of K for the Prunus brigantina (A) and Prunophora (B) dataset. (PNG 120 kb)
Figure S2.
Bayesian clustering on Prunus brigantina samples in the French Alps. Prunus brigantina dataset that included 71 individuals sampled from the French Alps and two samples from the French CRB repository (PNG 1621 kb)
Figure S3.
Bayesian analysis on Armeniaca and wild Prunus brigantina accessions. Genetic subdivision among Armeniaca species, P. brigantina included, was inferred with STRUCTURE with 24 microsatellite markers listed in Table S2. The 648 samples belong to the five Armeniaca species as described by Rehder (1940) and correspond to European, Central Asian and Chinese P. armeniaca (N = 474), P. mume (N = 9), P. sibirica (N = 84), P. mandshurica (N = 8) and P. brigantina (N = 73). More details on those samples, except for P. brigantina (this study), are provided by Liu et al. (2019). (PNG 635 kb)
Figure S4.
Bayesian analysis on the Prunus brigantina dataset together with an extended Prunophora dataset. Genetic subdivision among Armeniaca, Prunus and Prunocerasus species was inferred with STRUCTURE with 23 microsatellite markers after removal of UDP96_018 due to poor amplification in cherry plums (supplemental information for the list of markers). The 226 samples belong to three different Prunophora species including P. brigantina (N = 73), P. cerasifera (N = 66), P. armeniaca (N = 87), P. salicina (N = 10), P. mume (N = 9), P. mexicana (N = 1), P. munsoniana (N = 1), P. maritima (N = 1), P. americana (N = 1) and P. subcordata (N = 1). The blue stars (*), at the bottom of the bar plots, correspond to Japanese plums (P. salicina) admixed with P. cerasifera. (PNG 1776 kb)
ESM 1
(XLSX 77 kb)
ESM 2
(XLSX 72 kb)
ESM 3
(XLSX 77 kb)
ESM 4
(XLSX 10 kb)
ESM 5
(XLSX 69 kb)
ESM 6
(DOCX 2545 kb)
Rights and permissions
About this article
Cite this article
Liu, S., Decroocq, S., Harte, E. et al. Genetic diversity and population structure analyses in the Alpine plum (Prunus brigantina Vill.) confirm its affiliation to the Armeniaca section. Tree Genetics & Genomes 17, 2 (2021). https://doi.org/10.1007/s11295-020-01484-6
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11295-020-01484-6