Abstract
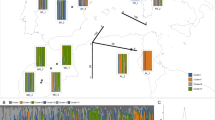
The conservation of genetic resources is a prerequisite for the maintenance of long-lived forest species. Araucaria araucana (Mol.) K. Koch is one of the oldest conifers in South America and a representative symbol of Chilean forest due to its endemicity and longevity. The aim of this study was to evaluate the genetic structure of the current A. araucana populations in Chile, to verify the possible genetic divergence between Coastal and Andean populations and to assess whether bottleneck events have influenced habitat fragmentation and threaten the genetic resources and evolutionary potential of the species. Twelve natural populations, nine from the Andes Cordillera and three from the Coast Cordillera were analysed by means of eight genomic microsatellite markers developed in A. araucana. Results of analysis of molecular variance (AMOVA) highlighted significant differentiation between Coastal and Andean populations (16 %; P = 0.004), detecting one significant barrier that separated populations from both Cordilleras as maximally differentiated areas. At local scale, both ranges revealed significant inter-population differentiation, with higher values for Coastal populations compared with Andean populations. These results suggested the presence of four gene pools (three in the Andes and one in the Coast Cordilleras) and one population (VIL) in the Coast Cordillera that differed to the rest. The differentiation between the Andean and Coastal populations may provide important baseline data that should allow further studies of landscape genetics in the species and that can contribute to develop conservation strategies for its genetic resources.





Similar content being viewed by others
References
Aagesen DL (1998) On the northern fringe of the South American temperate forest. The history and conservation of the monkey-puzzle tree. Environ Hist 3:64–85
Arana MV, Gallo LA, Vendramin GG, Pastorino MJ, Sebastiani F, Marchelli P (2010) High genetic variation in marginal fragmented populations at extreme climatic conditions of the Patagonian Cypress Austrocedrus chilensis. Mol Phylogenet Evol 55:805–815
Azpilicueta MM, Marchelli P, Gallo LA (2009) The effects of Quaternary glaciations in Patagonia as evidenced by chloroplast DNA phylogeography of Southern beech Nothofagus oblique. Tree Genet Genomes 5:561–571
Bagnoli F, Vendramin GG, Buonamici A, Doulis G, Gonzalez-Martinez SC, La Porta N, Magri D, Raddi P, Sebastiani F, Fineschi S (2009) Is Cupressus sempervirens native in Italy? An answer from genetic and paleobotanical data. Mol Ecol 18:2276–2286
Bekessy SA, Allnut TR, Premoli AC, Lara A, Ennos RA, Burgman MA, Cortes M, Newton AC (2002) Genetic variation in the vulnerable and endemic Monkey Puzzle tree, detected using RAPDs. Heredity 88:243–249
Comps B, Gomory D, Letouzey J, Thiebaut B, Petit RJ (2001) Diverging trends between heterozygosity and allelic richness during postglacial colonization in the European beech. Genetics 157:389–397
Cornuet JM, Luikart G (1996) Description and power analysis of two tests for detecting recent population bottlenecks from allele frequency data. Genetics 144:2001–2014
Delmastro R, Donoso C (1980) Review of the distribution, variation and utilization of gene resources of Araucaria araucana (Mol.) K. Koch in Chile. In: Proccedings of the IUFRO Symposio em melboramiento genetico e productividad de especias forestais de rapido crecimiento
Dinerstein E, McCracken GF (1990) Endangered greater one-horned rhinoceros carry high levels of genetic variation. Conserv Biol 4:417–422
Donoso C (1981) Tipos forestales de los bosques nativos de Chile. Documento de trabajo No. 38. Investigación y Desarrollo Forestal FAO/DP/CHI/76/003. Santiago, Chile
Drake F, Martín MA, Herrera MA, Molina JR, Drake-Martin F, Martín LM (2009) Networking sampling of Araucaria araucana (Mol.) K. Koch in Chile and the bordering zone of Argentina: implications for the genetic resources and the sustainable management. iForest 2:207–212
Dupanloup I, Schneider S, Excoffier L (2002) A simulated annealing approach to define the genetic structure of populations. Mol Ecol 11:2571–2581
Earl DA, vonHoldt BM (2012) Structure Harvester: a website and program for visualizing structure output and implementing the Evanno method. Conserv Genet Res 4:359–361
Eckert CG, Samis KE, Lougheed SC (2008) Genetic variation across species’s geographical ranges: the central-marginal hypothesis and beyond. Mol Ecol 17:1170–1188
El Mousadik A, Petit RJ (1996) High level of genetic differentiation for allelic richness among populations of the argan tree (Argania spinosa (L.) Skeels) endemic to Morocco. Theor Appl Genet 92:832
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of cluster of individuals using the software Structure: a simulation study. Mol Ecol 14:2611–2620
Excoffier L, Lischer L (2010) Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Res 10:564–567
Falush D, Stephens M, Pritchard JK (2007) Inference of population structure using multilocus genotype data: dominant markers and null alleles. Mol Ecol Notes 7:574–578
Fussi B, Lexer C, Heinze B (2010) Phylogeography of Populus alba (L.) and Populus tremula (L.) in Central Europe: secondary contact and hybridisation during recolonisation from disconnected refugia. Tree Genet Genomes 6:439–450
Garza JC, Williamson EG (2001) Detection of reduction in population size using data from microsatellite loci. Mol Ecol 10:305–318
González-Martínez SC, Dubreuil M, Riba M, Vendramin GG, Sebastiani F, Mayol M (2010) Spatial genetic structure of Taxus baccata L. in the western Mediterranean Basin: past and present limits to gene movement over a broad geographic scale. Mol Phylogenet Evol 55:805–815
Goudet J (2002) FSTAT, a program to estimate and test gene diversities and fixation indices (version 2.9.3.2). http://www.unil.ch/izea/softwares/fstat.html
Hailer F, Helander B, Folkestad AO, Ganusevich SA, Garstad S, Hauff P, Koren C, Nygård, Volke V, Vilà C, Ellegren H (2006) Bottlenecked but long-lived: high genetic diversity retained in white-tailed eagles upon recovery from population decline. Biol Lett 22:316–319
Herrmann TM (2006) Indigenous knowledge and management of Araucaria araucana forest in the Chilean Andes: implications for native forest conservation. Biodivers Conserv 15:647–662
Heusser CJ, Rabassa J, Brandant A, Stuckenrath R (1988) Late-Holocene vegetation of the Andean Araucaria region, Province Neuquen, Argentina. Mt Res Dev 8:53–63
Hewitt GM (1996) Some genetic consequences of ice ages, and their role in divergence and speciation. J Linn Soc 58:247–276
Hoban SM, Borkowski DS, Brosi SB, McCleary TS, Thompson LM, McLachlan JS, Pereira MA, Schlarbaum S, Romero-Severson JR (2010) Range-wide distribution of genetic diversity in the North American tree Juglans cinerea: a product of range shifts, not ecological marginality or recent population decline. Mol Ecol 19:4876–4891
Hoffmann A, Sierra M, Prosser C, Valle M (2001) Enciclopedia de los Bosques Chilenos. Gráfica Andes, Santiago de Chile
Hollin JT, Schilling DH (1981) Late Wisconsin-Weichselian mountain glaciers and small ice caps. In: Denton G, Hughes TJ (eds) The last ice sheets. Wiley, New York, pp 179–206
Hubisz MJ, Falush D, Stephen M, Pritchard JK (2009) Inferring weak population structure with the assistance of sample group information. Mol Ecol Res 9:1322–1332
IUCN (1996) World list of threatened trees. International Union for the Conservation of Nature, Gland, Switzerland. http://www.iucn.org/themes/ssc/redlist/redlist.htm
Jakobsson M, Rosenberg NA (2007) CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodal in analysis of population structure. Bioinformatics 14:1801–1806
Kershaw AP, McGlone MS (1995) The Quaternary history of the southern conifers. In: Enright N, Hill R (eds) Ecology of the Southern Conifers. Melbourne University Press, Melbourne, pp 30–63
Knotters M, Brus D, Oude J (1995) A comparison of kriging, co-kriging and kriging combined with regression for spatial interpolation of horizon depth with censored observations. Geoderma 67:227–246
Lu G, Wong D (2008) An adaptive inverse-distance weighting spatial interpolation technique. Comput Geosci 34:1044–1055
Magri D, Fineschi S, Bellarosa R, Buonamici A, Sebastani F, Schirone B, Simeone MC, Vendramin GG (2007) The distribution of Quercus suber chloroplast haplotypes matches the paleogeographical history of western Mediterranean. Mol Ecol 16:5259–5266
Marchelli P, Gallo L (2004) The combined role of glaciation and hybridization in shaping the distribution of genetic variation in a Patagonian southern beech. J Biogeogr 31:451–460
Marchelli P, Baier C, Mengel C, Ziegenhagen B, Gallo LA (2010) Biogeographic history of the threatened species Araucaria araucana (Molina) K. Koch and implications for conservation: a case study with organelle DNA markers. Conserv Genet 11:951–963
Marconi G, Martín MA, Cherubini M, Raggi L, Drake F, Villani F, Albertini E, Mattioni C (2011) Microsatellite-AFLP development for Araucaria araucana (Mol.) K. Koch, an endangered conifer of Chilean and Argentinean native forests. Silvae Genet 60:285–288
Markgraf V, McGlone MC, Hope G (1995) Neogene paleoenvironmental and paleoclimatic change in southern temperate ecosystems. A southern perspective. Trends Ecol Evol 10:143–147
Martín MA, Mattioni C, Lusini I, Drake F, Cherubini M, Herrera MA, Villani F, Martín LM (2012) Microsatellite development for the relictual conifer Araucaria araucana (Araucariaceae) using next-generation sequencing. Am J Bot 99:e213–e215
Miller MP (2005) Alleles in the Space (AIS): computer software for the joint analysis of interindividual spatial and genetic information. J Hered 96:722–724
Monmonier M (1973) Maximun-differences barriers: an alternative numerical regionalization method. Geogr Ann B 5:245–261
Moreno R, Zamora R, Molina JR, Vasques A, Herrera MA (2011) Predictive modelling of microhabitats for endemic birds in South Chilean temperate forests using Maximum entropy (Maxent). Ecol Inf 6:364–370
Myers N, Mittermeier RA, Mittermeier CG, Dafonseca GAB, Kent J (2000) Biodiversity hotspots for conservation priorities. Nature 403:853–858
Nei M, Roychoudhury AK (1974) Sampling variance of heterozygosity and genetic distance. Genetics 76:379–390
Nei M, Maruyama T, Chakraborty R (1975) The bottleneck effect and genetic variability in populations. Evolution 29:1–10
Ortego J, Bonal R, Muñoz A (2010) Genetic consequences of habitat fragmentation in long-lived tree species: the case of the Mediterranean holm oak (Quercus ilex, L.). J Hered 101:717–726
Pautasso M (2009) Geographical genetics and conservation of forest tree. Perspect Plant Ecol 11:157–189
Peakall R, Smouse PE (2005) GenAlEx 6: genetic analysis in excell. Population genetic software for teaching and research. Australian National University, Canberra, http://www.anu.edu.au/BoZo/GenAlEx
Peery MZ, Kirby R, Reid BN, Stoelting R, Doucet-Beër E, Robinson S, Vazquez-Carrillo C, Pauli JN, PalsbØll PJ (2012) Reliability of genetic bottleneck tests for detecting recent population declines. Mol Ecol 21:3403–3418
Petit R, Aguinagalde JI, de Beaulieu JL, Bittkau C, Brewer S, Cheddadi R, Ennos R, Fineschi S, Grivet D, Lascoux M, Mohanty A, Müller-Starck GM, Demesure-Musch B, Palme A, Martin JP, Rendell S, Vendramin GG (2003) Glacial refugia: hotspots but not melting pots of genetic diversity. Science 300:1563–1565
Piry S, Luikart G, Cornuet JM (1999) Computer note. Bottleneck: a computer program for detecting recent reductions in effective population size from allele frequency data. J Hered 90:502–503
Pollegioni P, Woeste K, Olimpieri I, Marandola D, Cannata F, Malvolti ME (2011) Long term human impacts on genetic structure of Italian walnut inferred by SSR markers. Tree Genet Genomes 7:707–723
Premoli AC, Souto C, Rovere P, Allnut TR, Newton AC (2002) Patterns of isozyme variation as indicator of biogeographic history in Pilgerodendrom uviferum (D. Don) Florín. Divers Distrib 8:57–66
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Rafii ZA, Dodd RS (1998) Genetic diversity among coastal and Andean natural populations of Araucaria araucana (Mol.) K. Koch. Biochem Syst Ecol 26:441–451
Rondanelli MJ (2000) Vegetational history of the andean forest of Araucaria araucana (Molina) K. Koch, in the river basin of the Bio-Bio river’s High Valley, province of Lonquimay, middle-south Chile during Holocene. Palynological analysis of the “Miraflores 2” profile. Zlb Geo Pal 7(8):1041–1051
Rosenberg NA (2004) DISTRUCT: a program for the graphical display of population structure. Mol Ecol Notes 4:137–138
Ruiz E, González F, Torres-Diaz C, Fuentes G, Mardones M, Stuessy T, Samuel R, Becerra J, Silva M (2007) Genetic and differentiation within and among Chilean populations of Araucaria araucana (Araucariaceae) based on allozyme variabilty. Taxon 56:1221–1228
Schneider S, Roessli D, Excoffier L (2000) Arlequin: a software for population genetics data analysis, version 3.1. genetics and Biometry Laboratory, Department of Anthropology, University of Geneva, Switzerland
Scotti-Saintagne C, Dick CW, Caron H, Vendramin GG, Troispoux V, Sire P, Casalis M, Buonamici A, Valencia R, Lemes MR, Gribel R, Scotti I (2013) Amazon diversification and cross-Andean dispersal of the widespread Neotropical tree species Jacaranda copaia (Bignoniaceae). J Biogeogr 40:707–719
Spear SF, Peterson CR, Matocq M, Storfer A (2006) Molecular evidence for historical and recent population size reductions of tiger salamanders (Ambystoma tigrinum) in Yellowstone National Park. Conserv Genet 7:605–611
Van Oosterhout C, Hutchinson WF, Wills DPM, Shipley P (2004) Micro-Checker: software for identifying and correcting genotyping errors in microsatellite data. Mol Ecol Notes 4:535–538
Veblen TT (1982) Regeneration patterns in Araucaria araucana forest in Chile. J Biogeogr 9:11–28
Villagrán C, Le-Quesne C, Aravena JC, Jiménez H, Hinojosa F (1998) El rol del clima cuaternario en la distribución actual de la vegetación de Chile central-sur. Bamb Geograph Schr 15:227–242
Wang XR, Szmidt AE (2001) Molecular markers in population genetics of forest trees. Scand J For Res 16:199–220
Watson DF (1992) Contouring: a guide to the analysis and display of spatial data. Pergamon Press, New York
Weir BS, Cockerham CC (1984) Estimating F-statistics for the analysis of populations structure. Evolution 38:1358–1370
Widmer A, Lexer C (2001) Glacial refugia sanctuaries for allelic richness, but not for gene diversity. Trends Ecol Evol 16:267–269
Williamson-Natesan EG (2005) Comparison of methods for detecting bottlenecks from microsatellite loci. Conserv Genet 6:551–562
Acknowledgments
This research was partially supported by grants (No. 207-141-018-1.0) from the Research Service of Concepción University (Chile), A/023099/09 and A/030789/10, of the Spanish Agency of International Cooperation for the Development from the Spanish Ministry for Foreign Affairs and Cooperation. The first author is grateful to Agrifood Campus of International Excellence (ceiA3) from the Spanish Ministry of Education and the Ministry of Science and Innovation for the financial support.
Data archiving statement
Following the standard Tree Genetics and Genomes policy, we declare that the samples and populations SSR data will be uploaded in the TreeGenes Database (http://dendrome.ucdavis.edu).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by P. Ingvarsson
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
Matrix of correlation between geographical variables and diversity parameters. (DOC 28 kb)
ESM 2
Percentage of membership (admixture proportion-Q) of each population in each cluster inferred by STRUCTURE software. A Q-value greater than 0.75 indicates that the population belongs to the cluster. (DOC 33 kb)
ESM 3
Inference of K, the most probable number of clusters, using Structure software, with locprior. A) Log-likelihood value of data L(K) as a function of K averaged over six replicates; B) Log-likelihood of the data (ΔK) as a function of K, calculated over six replicates. Plots made using Structure Harvester (Earl and von Holdt 2011). (DOC 45 kb)
ESM 4
Genetic diversity indices for the four groups of populations identified by Samova software analysis. (DOC 34 kb)
Rights and permissions
About this article
Cite this article
Martín, M.A., Mattioni, C., Lusini, I. et al. New insights into the genetic structure of Araucaria araucana forests based on molecular and historic evidences. Tree Genetics & Genomes 10, 839–851 (2014). https://doi.org/10.1007/s11295-014-0725-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11295-014-0725-1