Abstract
Dipsacus asperoides contains multiple pharmacologically active compounds. The principal are saponins. The plant can be cultivated, but it contains lower levels of bioactive compounds than the plant in the wild. It may be the reason to exploit the endophytic fungi that colonize the plant roots in order to produce bioactive compounds. However, the endophytic fungi of D. asperoides have not been analyzed in detail. In this study, we isolated and identified 46 endophytic fungal strains from the taproots, lateral roots and leaves, and we used morphological and molecular biological methods to assign them into 15 genera: Fusarium sp., Ceratobasidium sp., Chaetomium sp., Penicillium sp., Aspergillus sp., Talaromyces sp., Cladosporium sp., Bionectria sp., Mucor sp., Trichoderma sp., Myrothecium sp., Clonostachys sp., Ijuhya sp., Leptosphaeria sp. and Phoma sp. Taproots contained abundant endophytic fungi, the numbers of which correlated positively with level of dipsacus saponin VI. Primary fermentation of several endophytic fungal strains from taproots showed that Fusarium, Leptosphaeria, Ceratobasidium sp. and Phoma sp. can produce the triterpenoid saponin. These results may guide efforts to sustainably produce bioactive compounds from D. asperoides.
Graphical abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Dipsacus asperoides is a well-known medicinal plant used to curing occlusion diseases, punch injury, and rheumatism (Niu et al. 2015; Wong et al. 2007). Saponins, the major bioactive compound in D. asperoides, are isolated primarily from the taproots and widely used to treat fractures (Zhang et al. 2003; Jung et al. 2012). D. asperoides in the wild has diminished as a result of exploitation (Zhang et al. 1997; Chen et al. 2014; Wang et al. 2016), and the cultivated plant contains lower levels of dipsacus saponin VI than the plant in the wild. Therefore, rapid, efficient and environmentally sustainable methods are needed to obtain this and other saponins from D. asperoides (Cira et al. 2008; Jiao et al. 2015).
It may be possible to obtain saponins from the endophytic fungi that colonize D. asperoides (Jiao et al. 2015). Such fungi colonize the flowers, seeds, taproots, stems and leaves of many plant species, without causing visible disease symptoms (Aly et al. 2011). Endophytes establish a long-term symbiotic relationship with their plant hosts (Zuccaro et al. 2011). Some endophytic fungi and their metabolites increase resistance to plant pathogens and tolerance to drought (Redman et al. 2002; Waller et al. 2005; Herre et al. 2007; Rodriguez and Redman 2008). Endophytic fungi can combine with other endogenous microorganisms and antibacterial compounds to form a defense system that produces alkaloids to strengthen immunity and maintain growth under stress (Qin et al. 2011; Clay and Holah 1999). Some endophytic fungi produce active proteases helped maintain plant activities, such as pectinase and esterase, which degrade cell walls (Zhao et al. 2016). Pathology can result when programmed senescence in the plant or environmental change perturb the fungal population in the plant (Stamford et al. 2001).
In addition to supporting the growth and productivity of medicinal plants, endophytic fungi can produce bioactive metabolites similar to plant hosts, making them a potential source of medicinal compounds (Chandra 2012). For example, the endophytic fungus isolated from Taxus chinensis can be produced the anti-cancer compound paclitaxel (Li et al. 2009b). Other endophytic fungi produce the drug compounds camptothecin, podophyllotoxin (Eyberger et al. 2006; Puri et al. 2006), hypericin and emodin (Kusari et al. 2009). Endophytic fungi can produce bioactive compounds through industrial fermentation (Winter et al. 2011; Walsh and Fischbach 2010) (Kusari and Spiteller 2011). The research of endophytic fungi may provide new ideas and methods for developing bioactive compounds in medicinal plants in ways that sustain the development of traditional Chinese medicine resources (Zuccaro et al. 2011).
Endophytic fungi have been analyzed in at least 145 species of medicinal plants, but no such analysis has been reported for D. asperoides, to the best of our knowledge. Here we characterized the taxonomic diversity of D. asperoides taproots, lateral roots and leaves, and analyzed the correlation between the number of fungi and the level of saponins. Primary fermentation was performed with several endophytic fungal strains to examine the possibility of large-scale development of natural products.
Materials and methods
Sample collection
Two-year-old D. asperoides from Meihuashan in Weining County, Guizhou Province (N26°23′10.46′′) was planted at the Guiyang University of Chinese Medicine (E106°37′41.64′′). Plant material was washed and soaked in 0.1% SDS for 10 min, then rinsed with double-distilled water. The material was divided into taproots, lateral roots and leaves, which were stored at 4 °C.
Isolation of endophytic fungi
The surface of plant material was sterilized by soaking in 0.1% mercuric chloride for 5 min, then in 75% ethanol for 3 min. The disinfected material was rinsed three times (1 min each time) with sterile water. The material was cut into pieces measuring 0.5 × 0.5 cm, and incubated at 28 °C on petri dishes containing potato dextrose agar (PDA), tryptone soy agar (TSA), beef extract tryptone agar (NA) and Luria–Bertani (LB) medium. Five biological replicates were prepared for each tissue, and growth was monitored every day. Endophytic fungal strains were inoculated on PDA slant culture-medium. After fungal cultures were fully grown, slant culture tubes were closed with tampons wrapped with oilpaper and stored at 4 °C.
Taxonomic identification of endophytic fungi
Identification of endophytic fungi was accomplished following the methods described by Cannon et al. (Cannon and Simmons 2002). In this study, we perform its molecular reidentification, based on the analysis of internal transcript spacer (ITS) regions of endophytic fungi (Ding et al. 2018; Koljalg et al. 2005). Colonial morphology of endophytic fungi was identified using the point planting method as described (Chen et al. 2012). In brief, fungal spores were inoculated onto the center of solid PDA and incubated at 28 °C. Fungal characteristics were recorded every day, including colony shape, height and color of aerial hyphae, base color, growth rate, margin, surface texture, and depth of growth into the agar. At least three cultures were characterized on each petri dish, and on the attempts equated colony morphologies from different plates of the same plant. Endophytic fungi were preliminarily assigned to genera based on spore and culture characteristics.
The sequence analysis was also performed to assist in specimen identification. Mycelium was gathered directly from the surface of 4-day-old agar cultures and ground into a powder in liquid nitrogen. The powder was suspended in buffer [200 mM Tris–HCl (pH 8.0), 25 mM EDTA (pH 8.0), 250 mM NaCl and 0.5% SDS (pH 7.5)]. DNA was extracted using phenol and chloroform, and precipitated in ethanol. DNA integrity was analyzed by agarose gel electrophoresis, and purity was assessed using a Micronuclear Quantifier (Nanodrop 2000, Thermo Scientific, USA).
Internal transcript spacer (ITS) regions of endophytic fungi were amplified using polymerase chain reaction (PCR) and the universal ITS primers, V9D (5′-TTAAGTCCCTGCCCTTTGTA-3′) and LS266 (5′-GCATTCCCAAACAACTCGACTC-3′). Reactions (25 µL) contained 100 ng of genomic DNA, 10 µM of each primer, 12 µL of Premix Taq™ (Ex Taq™ 2.0 plus dye) and sterile double-distilled water. Thermal cycling parameters for PCR were as follows: pre-denaturation at 94 °C for 5 min; 30 cycles of denaturation at 94 °C for 30 s, annealing at 53 °C for 30 s and extension at 72 °C for 2 min; and a final extension step at 72 °C for 10 min. PCR products were detected on 1.2% (w/v) agarose gel prepared in 1× TAE buffer and electrophoresed at 100 V for 45 min.
Fragments were eluted and sent to be sequenced by Kingsley Biotech (Nanjing, China). Further information to guide the taxonomic identification of fungal strains came from the Flora of Chinese Mycology. BLAST searches of fungal sequences were conducted to analyze homology with identified sequences in ITS. Moreover, the comparison analysis of UNITE database to complement GenBank results. Tree topologies were evaluated using bootstrap analyses in MEGA6 (1000 bootstrap replicates). Phylogenetic trees were inferred using the neighbor-joining method.
Analysis of endophytic fungal diversity
Menhinick’s index (Dmn) was used to quantify species richness among the isolated endophytic fungi. Dmn was calculated as \(Dmn=S/\sqrt N\), where S refers to the number of different endophytic fungal species, and N refers to the total number of isolated endophytic fungi. The Shannon diversity index (H′) was calculated using \(H'= - \sum\nolimits_{i}^{k} {Pi(LnPi)}\), where Pi = Ni/N, Ni is the number of individuals of the species, and k is the number of different endophytic species in a sample. The isolation rate (IR) was calculated by dividing the total number of isolates in a trial by the total number of samples in the trial. IR was used to measure the richness of endophytic fungi in plant tissues. The Sorensen similarity index (Cs) was calculated as Cs = 2j/(a + b), where j is the number of endophytic fungi common to the two tissues being compared, and a and b are the numbers of endophytic fungi in each tissue. Cs was used to quantify species similarity between different tissues.
Quantification of dipsacus saponin VI
Samples of D. asperoides taproots, lateral roots and leaves were dried and ground into powder. Sample powder were soaked in methanol solution, ultrasonicated for 30 min (power, 100 W; frequency, 40 kHz), allowed to cool, weighed, and membrane-filtered. The filtered sample was analyzed for dipsacus saponin VI on a C18 symmetry column (4.6 × 250 mm, 5 µm) on a Waters HPLC system, with the following chromatography parameters: mobile phase, acetonitrile–water (30:70); flow rate, 1.0 mL/min; injection volume, 20 µL; detection wavelength, 212 nm; and theoretical plate number, ≥ 3000. HPLC run time was 25 min (Pharmacopoeia of the People’s Republic of China, 2015).
HPLC was also conducted with standard dipsacus saponin VI (purity, 91.3%; JY8R—BINA2), which was obtained from the China Food and Drug Certification Research Institute (Beijing, China). The standard was dissolved in methanol to a concentration of 0.15 mg/mL. Retention time of the standard was 18.254 min under our conditions.
Fermentation of endophytic fungi
Taproot mycelium were transferred to an Erlenmeyer flask containing 100 mL liquid medium and cultured at 28 °C for 5 days with shaking at 160 rev min−1. Fungal characteristics were recorded every day, including color, viscosity, odor and other properties of the fermentation broth. Samples of mycelium (100 mg) were harvested by filtering and ground into powder in liquid nitrogen. DNA was extracted and PCR-amplified as described above (Cannon and Simmons 2002).
Statistical analyses
All results were expressed as mean ± SEM. Graphs were prepared using GraphPad Prism 7.0. Differences between mean values were assessed for significance using one-way analysis of variance (ANOVA), followed by the least significant difference (LSD) test for post hoc comparisons (equal variances were assumed). Significance was indicated as follows: *P < 0.05, **P < 0.01, and ***P < 0.005.
Results
Identification of endophytic fungi from D. asperoides roots and leaves
Different tissues of D. asperoides were cultured in PDA, LB, TSA and NA culture media. A total of 46 strains were isolated and preliminarily assigned based on colony and hyphal characteristics (Fig. 1). The largest number of endophytic fungal isolates (40) were found in taproots, followed by leaves (4) and lateral roots (2) (Fig. 1A). The isolates in four media showed that the greatest number was obtained in PDA (37), followed by NA (4), TSA (3) and finally LB medium (2) (Fig. 1B). The IR in taproots (0.40) was significantly higher than that in lateral roots (0.02) or leaves (0.04). Taproots also showed that H′ and Dmn were higher than leaves and lateral roots (Table 1; Fig. 2). These results suggest that the taproots may provide the best niche or entry point for colonization and penetration by endophytic fungi.
Isolation of endophytic fungi from Dipsacus asperoides. Tissues of Dipsacus asperoides were cultured in the culture medium of PDA, LB, TSA and NA. 46 isolates were identified in 100 taproot segments, 100 lateral root segments and 100 leaf segments based on their morphological characteristics. A Distribution of endophytes in different tissues of D. asperoides: 40 strains (86.96%) were isolated from the taproots, 2 strains (4.35%) were isolated from lateral roots and 4 strains (8.70%) were isolated from the leaves. B Culture of different endophytes from different D. asperoides tissues on different media: PDA supported growth of 37 strains (80.43%); LB medium, 2 strains (4.35%); TSA, 3 strains (6.52%); and NA, 4 strains (8.70%)
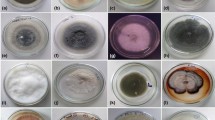
Microscopic analysis of the 46 endophytic fungi allowed them to be assigned preliminarily to Fusarium sp. (samples daef 1–14) on the basis of their irregular, round shape and hyphae uplift, fast growth, yellow pigment production, and presence of conidia or spores; to Ceratobasidium sp. (daef 15–18) on their basis of their loose white hyphae and lack of conidia; to Chaetomiaceae sp. (daef 19–20) on the basis of white colonies and soft hair or cotton with yellow pigment on the back of hyphae; to Penicillium sp. (daef 21–22) on the basis of scattered hyphae, pigment production, and broom-like stem with a string of conidia; to Aspergillus sp. (daef 23) on the basis of white, pilose, cotton-like hyphae with erect hyphae, conidiophores and a hemispherical capsule; to Talaromyces sp. (daef 24) on the basis of green cotton-like hyphae and a small, broom-like stem with conidiophores; to Cladosporium sp. (daef 25) on the basis of green villi-like hyphae with small water-like substances, elliptical and round conidia; and to Bionectria sp. (daef 26–27) on the basis of white cotton-like appearance with some water-like substances, broom-like branches with long spindle-shaped spore stalks on the branchlets, and small curved elliptical and ovate spores on the spore stalks (Figs. 3 and S1).
Morphological characteristics of endophyte fungi. Photographs showing typical morphology of endophyte fungi from taproots, lateral roots and leaves of D. asperoides. A Characteristics of endophytic fungi isolated from the taproots, showing “surface”, “back” and microstructure. These characteristics were observed for the following isolates: daef 11, 15, 22, 23, 25, 40, 41 and 44. Scale bar, 20 µm. B Characteristics of endophytic fungi isolated from the lateral roots, showing “surface”, “back” and microstructure. These characteristics were observed for daef 5 and 21. Scale bar, 20 µm. C Characteristics of endophytic fungi isolated from the leaves, showing “surface”, “back” and microstructure. These characteristics were observed for daef 18 and 37. Scale bar, 20 µm
The remaining endophytic fungal isolates were assigned to genera based on comparison with known fungi: Clonostachys sp. (daef 28–29), Mucor sp. (daef 30–34), Trichoderma sp. (daef 35–36), Myrothecium sp. (daef 37), Ijuhya sp. (deaf 38–39), Leptosphaeria sp. (daef 40–42), Phoma sp. (daef 43–45) and Heliogales sp. (daef 46).
Comparison of ITS sequences from the 46 isolates with fungal sequences in GenBank (Table 2) lead to the identification of 15 genera: Fusarium sp., Ceratobasidium sp., Chaetomium sp., Penicillium sp., Aspergillus sp., Talaromyces sp., Cladosporium sp., Bionectria sp., Mucor sp., Trichoderma sp., Myrothecium sp., Clonostachys sp., Ijuhya sp., Leptosphaeria sp. and Phoma sp. Two strains that could not be assigned to a genus were identified as Chaetomiaceae sp. and Helotiales sp. based on GenBank analysis. Taxonomic identification based on ITS sequencing was consistent with that based on morphological observation. In addition, the results of blastn analysis by UNITE database were consistent with NCBI analysis (Table S1).
The two dominant genera were Fusarium sp. to which 29.09% of isolates, and Ceratobasidium sp. to which 10.91% of isolates. Myrothecium sp. was isolated only from leaves. Cs analysis showed that the tissue pair with greatest similarity was lateral roots and leaves (Cs 2.00), followed by taproots and lateral roots (1.88) and finally taproots and leaves (1.79). These results suggest the heterogeneity of the endophyte assemblage.
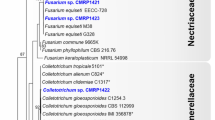
A phylogenetic tree based on ITS sequences (Fig. 4) assigned Fusarium sp. isolates to six clusters, three of which were closely related and clustered with Fusarium globosum (LT746280.1), two of which clustered with Fusarium tricinctum (MG274296.1) and Fusarium sp. (JF740911.1), and one of which was related to Fusarium solani (KY484984.2). Clades comprised daef 6, 7, 13 and 14; daef 4 and 8; and daef 11 and 12. Fusarium sp. was the most frequently isolated fungal genus. The four isolates daef 30 and 32–34 were grouped into a branch with the reference taxon Mucor sp. The daef 15, 16 and 18 and Ceratobasidium sp. were grouped into a branch with 100% bootstrap support, with daef 15 clustering with Ceratobasidium sp. (KC782938.1).
Phylogenetic identification of endophytic fungi from D. asperoides. Phylogenetic tree based on neighbor-joining analysis of ITS sequences from the 46 strains of endophytic fungi isolated from taproots, lateral roots and leaves. ITS sequences obtained were submitted to the NCBI database, and BLAST searches were performed to select species showing 95–100% homology with the isolated species. Closely related species are labeled with taxonomic names, followed by the accession number. Significant bootstrap values are indicated at the branching points
Leptosphaeria isolates formed a cluster with reference taxa Leptosphaeria sp. (KJ934197.1 and AJ317958.1). Phoma sp. isolates formed a cluster with reference taxa Phoma exigua var. (EU343130.1 and EU343168.1). Penicillium sp. isolates were grouped into two clusters: daef 21 clustered with Penicillium janthinellum (MG938669.1), and daef 22 clustered with Penicillium skrjabinii (EU427287.1). Trichoderma isolates formed a cluster with Trichoderma hamatum (KM491888.1), Trichoderma asperellum (KF723005.1) and Trichoderma koningiopsis (GQ229070.1). Clonostachys sp. isolates formed a cluster with Clonostachys rosea f. (HM751081.1), Clonostachys sp. (KC806284.1) and Clonostachys pseudochroleuca (KC806259.1). A Bionectria sp. isolate formed a cluster with Bionectria sp. (KF367470.1) with 100% bootstrap support. Myrothecium sp. isolates were grouped into two clusters: two closely related isolates formed a clade and one isolate formed a clade with Myrothecium roridum (FJ914699.1) and Myrothecium sp. (KY086248.1) with 99% bootstrap support. Chaetomium sp. isolates formed a cluster with Chaetomium sp. (KC007192.1), Chaetomium pseudocochliodes (JN209925.1) and Chaetomium megallocarpum (KC109744.1). The Talaromyces sp. isolate formed a cluster with Talaromyces apiculatus (JN899375.1) with 100% bootstrap support. The Aspergillus sp. isolate formed a cluster with Aspergillus lentulus (KX903293.1) and Aspergillus viridinutans (EF661280.1). The Cladosporium sp. isolate formed a cluster with Cladosporium pseudocladosporioides (KP701943.1), Cladosporium delicatulum (KP701939.1) and Cladosporium cladosporioides (KP701868.1) with 100% bootstrap support. The daef 17 and 31 could not be represented in the phylogenetic tree because of low sequence quality. The daef 46 clustered with Helotiales sp. (FN548161.1 and MG066445.1) with 99% and 100% bootstrap support.
Positive correlation between dipsacus saponin VI level and number of endophytic fungi in roots
Dipsacus saponin VI was quantified in taproots, lateral roots and leaves using HPLC (Fig. 5A–D). Levels differed significantly in different tissues (P < 0.05). Levels were highest in taproots (2.98%), lower in lateral roots (0.87%) and below the detection limit in leaves (Fig. 5E). Level of dipsacus saponin VI positively correlated with the total number of endophytic fungi (Fig. 5F) and with the number of Fusarium sp. (Fig. 5G), but independent with the number of Mucor sp. in taproots (Fig. 5H).
Dipsacus saponin VI level positively correlated with endophytic fungi in roots of D. asperoides. Dipsacus saponin VI was quantified in taproots, lateral roots and leaves using HPLC. A Chromatogram of the standard dipsacus saponin VI. The y-axis indicates the absorbance of dipsacus saponin VI, and the x-axis indicates the measurement time (min). B Chromatogram of dipsacus saponin VI in taproots. C Chromatogram of dipsacus saponin VI in lateral roots. D Chromatogram of dipsacus saponin VI in leaves. E Quantification of dipsacus saponin VI content in different tissues. Data are mean ± SEM (n = 4). ***P < 0.005 (one-way ANOVA and least significant difference test post hoc). F Correlation analysis between dipsacus saponin VI level and the number of endophytic fungi in taproots and lateral roots. Each isolate is represented by a spot (n = 4, R2 = 0.9035, P = 0.0001). G Correlation analysis between dipsacus saponin VI level and the number of Fusarium sp. in taproots and lateral roots. Each isolate was represented as a spot (n = 4, R2 = 0.9122, P = 0.0001). H Correlation analysis between the dipsacus saponin VI level and the number of Mucor sp. in taproots and lateral roots. Each isolate was represented as a spot (n = 4, R2 = 0.0875, P = 0.0896)
Primary fermentation of endophytic fungi
Selected endophytic fungal isolates were subjected to primary fermentation tests to identify which strains may produce dipsacus saponin VI. Several endophytic fungi enriched in taproots and from different genera were tested: daef 11 (Fusarium sp.), 40 and 41 (Leptosphaeria sp.), 15 (Ceratobasidium sp.) and 44 (Phoma sp.). Within 10 min at 60 °C, all these strains produced foam and showed no fading (Fig. 6A). These strains may produce saponins. In addition, daef 15 produced red pigment, while daef 40, 41 and 44 produced green or deep green pigments.
Primary fermentation of endophytic fungi from Dipsacus asperoides. Five endophytic fungi that were enriched in taproots and came from different genera were subjected to primary fermentation: daef 11 (Fusarium sp.), 40 and 41 (Leptosphaeria sp.), 15 (Ceratobasidium sp.) and 44 (Phoma sp.). A Photographs of the fermentation of five strains of endophytic fungi and their microscopic morphology. Scale bar, 20 µm. B Quantification of synergistic alignment between ITS sequences of the foaming fungus (ITS’) and the ITS sequences of the original fungal isolate
To verify that these fermented fungi were identical to the strains originally isolated and were not contaminated by other microorganisms, we confirmed that the microstructure of the fermented fungi was consistent with that of the original strains. We also amplified ITS regions from mycelium of the fermented strains and confirmed that the sequences were 100% homologous to the regions sequenced from the original strains (Fig. 6B and S2).
Discussion
This study begins the process of correlating production of perhaps the most relevant bioactive compound from this plant, saponins, with the number and diversity of endophytic fungi in different tissues. Our results help clarify the biodiversity and phylogenetic relationships of endophytic fungi in D. asperoides, which can begin to shed light on how endophytic fungi can affect the quality of traditional Chinese medicinal plants.
The 46 endophytic fungi were isolated from different tissues of D. asperoides. This number is slightly lower than what has been reported with other plants, which may mean that some strains stopped colonizing D. asperoides over time, such as due to inhibition by other rapidly growing strains (Gonzaga et al. 2015).
Nearly all the fungal isolates in our study were colonized in taproots, while only two strains were isolated from lateral roots and four strains were isolated from leaves. This suggests that in this medicinal plant, the taproots are most likely to be colonized. The much greater abundance of fungi in taproots reflects that fungi can penetrate host plants via the roots, where they gain access to nutrients in xylem and phloem (Martin et al. 2012; Pfurtscheller and Klimesch 1990; Sheng-Liang et al. 2014). The diversity of endophytic fungi in taproots was more higher than that in other tissues. We guess that these fungi colonized D. asperoides as spores that moved from the soil to the roots; the lateral roots acted simply as transport bridges to carry the fungi to taproots for storage (Courty et al. 2018). In contrast to our results, diversity of endophytic fungi was greatest in the leaves of Gossypium hirsutum (Li et al. 2014) and Miscanthus × giganteus (Schmidt et al. 2018).
Endophytic fungi in plants are primarily Ascomycetes and their anamorphs, although they can also be Basidiomycetes, Zygomycetes, and Oomycetes (Soca-Chafre et al. 2011). In D. asperoides, endophytic fungi included many rare species, mainly belong to Deuteromycota, that accounted for 48.94% of isolates; Ascomycetes accounted for only 34.04% of isolates. Some Basidiomycetes and Zygomycetes were observed.
Fusarium sp. is the predominant microflora in D. asperoides. This genus occurs as an endophyte in various cash crops, including Solanum lycopersicum (Aime et al. 2013), Drepanocarpus lunatus (Liu et al. 2016), and Dioscorea zingiberensis (Zhang et al. 2009). Ceratobasidium sp. can cause sheath blight and act as a saprotroph in rice (Mosquera-Espinosa et al. 2013), persimmon (Ceresini et al. 2012) and soybean (Salehi et al. 2005). Aspergillus sp. acts as an endophyte of Opuntia dillenii and several other plants (Li et al. 2009a). Myrothecium sp. acts as an endophyte of Calophyllum apetalum and Garcinia Morella (Ruma et al. 2015).
Several of the endophytic fungi that were identified in D. asperoides can produce bioactive compounds of medicinal interest. Trichoderma sp., Talaromyces sp., Mucor sp. and Penicillium sp. can produce proteases that degrade cellulose (Zhao et al. 2016; Thongekkaew et al. 2013), dairy products, and polysaccharides (Inoue et al. 2015). Fusarium sp. can produce triterpenoid saponins (Cira et al. 2008; Jiao et al. 2015), which are the main secondary metabolites of D. asperoides and used to treat osteoporosis, reduce lipids and protect against oxidation (Wang et al. 2016). Our results suggest that the main location of saponin production in D. asperoides is roots. Levels of dipsacus saponin VI were higher in taproot than in other tissues, and the taproot have been also the greatest number of endophytic fungi. In contrast, dipsacus saponin VI levels in leaves were below the limit of detection, perhaps due to scarcity of interactions between endophytic fungi and host, reflected in the relatively low Dmn and IR. Endophytes can prefer different plant tissues, where they form specific symbiotic relationships; as a result, different tissues contain different profiles of secondary metabolites (Jasinska et al. 2018; Jarvis et al. 1985; Liu et al. 2006; Wang et al. 2016).
Changing the environmental conditions of endophytes can lead them to produce different secondary metabolites (Eaton et al. 2010; Wang et al. 2017), increasing their usefulness as bioactive molecule factories. Many active pharmaceutical compounds have been isolated from filtrates of Fusarium sp. cultures (Suzuki et al. 2013). The leptosins I and J have been isolated from Leptosphaeria mycelium (Takahashi et al. 1994). A cyclic lipodepsipeptide has been isolated from Phoma sp. (Herath et al. 2009). When we subjected Fusarium sp., Leptosphaeria sp., Ceratobasidium sp. and Phoma sp. to primary fermentation, we found that all them could produce triterpenoid saponin. In addition, our isolates (Cladosporium sp., Phoma sp., Fusarium sp., and Penicillium sp.) were produced pigments that may be useful in the food, cosmetic and pharmaceutical industries. These results with primary fermentation may facilitate the development of strategies to produce natural products from D. asperoides (Bick and Rhee 1966; Zheng et al. 2017; Shah et al. 2015).
Our results highlight the diversity of endophytic fungi in medicinal plants and their ability to synthesize bioactive secondary metabolites (Gupta et al. 2018). They may also guide new approaches to synthesize dipsacus saponin VI from D. asperoides, permitting sustainable development of this important traditional Chinese medicine resource.
References
Aime S, Alabouvette C, Steinberg C, Olivain C (2013) The endophytic strain Fusarium oxysporum Fo47: a good candidate for priming the defense responses in tomato roots. Mol Plant-Microbe Interact 26(8):918–926. https://doi.org/10.1094/MPMI-12-12-0290-R
Aly AH, Debbab A, Proksch P (2011) Fungal endophytes: unique plant inhabitants with great promises. Appl Microbiol Biotechnol 90(6):1829–1845. https://doi.org/10.1007/s00253-011-3270-y
Bick IR, Rhee C (1966) Anthraquinone pigments from Phoma foveata Foister. Biochem J 98(1):112–116
Cannon PF, Simmons CM (2002) Diversity and host preference of leaf endophytic fungi in the Iwokrama Forest Reserve. Guyana Mycologia 94(2):210–220
Ceresini PC, Costa–Souza E, Zala M, Furtado EL, Souza NL (2012) Evidence that the Ceratobasidium-like white-thread blight and black rot fungal pathogens from persimmon and tea crops in the Brazilian Atlantic Forest agroecosystem are two distinct phylospecies. Genet Mol Biol 35(2):480–497. https://doi.org/10.1590/S1415-47572012005000032
Chandra S (2012) Endophytic fungi: novel sources of anticancer lead molecules. Appl Microbiol Biotechnol 95(1):47–59. https://doi.org/10.1007/s00253-012-4128-7
Chen J, Wang H, Guo SX (2012) Isolation and identification of endophytic and mycorrhizal fungi from seeds and roots of Dendrobium (Orchidaceae). Mycorrhiza 22(4):297–307. https://doi.org/10.1007/s00572-011-0404-0
Chen DX, Li LY, Zhang X, Wang Y, Zhang Z (2014) Genetic diversity and population structure of wild Dipsacus asperoides in China as indicated by ISSR markers. Genet Mol Res 13(3):6340–6349. https://doi.org/10.4238/2014.February.14.12
Cira LA, Gonzalez GA, Torres JC, Pelayo C, Gutierrez M, Ramirez J (2008) Heterologous expression of Fusarium oxysporum tomatinase in Saccharomyces cerevisiae increases its resistance to saponins and improves ethanol production during the fermentation of Agave tequilana Weber var. azul and Agave salmiana must. Antonie Van Leeuwenhoek 93(3):259–266. https://doi.org/10.1007/s10482-007-9200-4
Clay K, Holah J (1999) Fungal endophyte symbiosis and plant diversity in successional fields. Science 285(5434):1742–1745
Courty PE, Buee M, Tech JJT, Brule D, Colin Y, Leveau JHJ, Uroz S (2018) Impact of soil pedogenesis on the diversity and composition of fungal communities across the California soil chronosequence of Mendocino. Mycorrhiza 28(4):343–356. https://doi.org/10.1007/s00572-018-0829-9
Ding CH, Wang QB, Guo S, Wang ZY (2018) The improvement of bioactive secondary metabolites accumulation in Rumex gmelini Turcz through co-culture with endophytic fungi. Braz J Microbiol 49(2):362–369. https://doi.org/10.1016/j.bjm.2017.04.013
Eaton CJ, Cox MP, Ambrose B, Becker M, Hesse U, Schardl CL, Scott B (2010) Disruption of signaling in a fungal-grass symbiosis leads to pathogenesis. Plant Physiol 153(4):1780–1794. https://doi.org/10.1104/pp.110.158451
Eyberger AL, Dondapati R, Porter JR (2006) Endophyte fungal isolates from Podophyllum peltatum produce podophyllotoxin. J Nat Prod 69(8):1121–1124. https://doi.org/10.1021/np060174f
Gonzaga LL, Costa LE, Santos TT, Araujo EF, Queiroz MV (2015) Endophytic fungi from the genus Colletotrichum are abundant in the Phaseolus vulgaris and have high genetic diversity. J Appl Microbiol 118(2):485–496. https://doi.org/10.1111/jam.12696
Gupta S, Bhatt P, Chaturvedi P (2018) Determination and quantification of asiaticoside in endophytic fungus from Centella asiatica (L.) Urban. World J Microbiol Biotechnol 34(8):111. https://doi.org/10.1007/s11274-018-2493-9
Herath K, Harris G, Jayasuriya H, Zink D, Smith S, Vicente F, Bills G, Collado J, Gonzalez A, Jiang B, Kahn JN, Galuska S, Giacobbe R, Abruzzo G, Hickey E, Liberator P, Xu D, Roemer T, Singh SB (2009) Isolation, structure and biological activity of phomafungin, a cyclic lipodepsipeptide from a widespread tropical Phoma sp. Bioorg Med Chem 17(3):1361–1369. https://doi.org/10.1016/j.bmc.2008.12.009
Herre EA, Mejia LC, Kyllo DA, Rojas E, Maynard Z, Butler A, Van Bael SA (2007) Ecological implications of anti-pathogen effects of tropical fungal endophytes and mycorrhizae. Ecology 88(3):550–558
Inoue H, Yano S, Sawayama S (2015) Effect of beta-Mannanase and beta-mannosidase supplementation on the total hydrolysis of softwood polysaccharides by the Talaromyces cellulolyticus cellulase system. Appl Biochem Biotechnol 176(6):1673–1686. https://doi.org/10.1007/s12010-015-1669-0
Jarvis BB, Lee YW, Yatawara CS, Mazzocchi DB, Flippen-Anderson JL, Gilardi R, George C (1985) 7 Alpha-hydroxytrichodermol, a new trichothecene from Myrothecium roridum. Appl Environ Microbiol 50(5):1225–1228
Jasinska A, Goralczyk A, Sobon A, Dlugonski J (2018) Novel laccase-like multicopper oxidases from the Myrothecium roridum fungus—production enhancement, identification and application in the dye removal process. Acta Biochim Polonica 65(2):287–295. https://doi.org/10.18388/abp.2017_2546
Jiao X, Lu X, Chen AJ, Luo Y, Hao JJ, Gao W (2015) Effects of Fusarium solani and F. oxysporum infection on the metabolism of Ginsenosides in American ginseng roots. Molecules 20(6):10535–10552. https://doi.org/10.3390/molecules200610535
Jung HW, Jung JK, Son KH, Lee DH, Kang TM, Kim YS, Park YK (2012) Inhibitory effects of the root extract of Dipsacus asperoides C.Y. T.M.Ai on collagen-induced arthritis in mice. J Ethnopharmacol 139(1):98–103. https://doi.org/10.1016/j.jep.2011.10.020
Koljalg U, Larsson KH, Abarenkov K, Nilsson RH, Alexander IJ, Eberhardt U, Erland S, Hoiland K, Kjoller R, Larsson E, Pennanen T, Sen R, Taylor AF, Tedersoo L, Vralstad T, Ursing BM (2005) UNITE: a database providing web-based methods for the molecular identification of ectomycorrhizal fungi. New Phytol 166(3):1063–1068. https://doi.org/10.1111/j.1469-8137.2005.01376.x
Kusari S, Spiteller M (2011) Are we ready for industrial production of bioactive plant secondary metabolites utilizing endophytes? Nat Prod Rep 28(7):1203–1207. https://doi.org/10.1039/c1np00030f
Kusari S, Zuhlke S, Kosuth J, Cellarova E, Spiteller M (2009) Light-independent metabolomics of endophytic Thielavia subthermophila provides insight into microbial hypericin biosynthesis. J Nat Prod 72(10):1825–1835. https://doi.org/10.1021/np9002977
Li DL, Li XM, Wang BG (2009a) Natural anthraquinone derivatives from a marine mangrove plant-derived endophytic fungus Eurotium rubrum: structural elucidation and DPPH radical scavenging activity. J Microbiol Biotechnol 19(7):675–680
Li YC, Tao WY, Cheng L (2009b) Paclitaxel production using co-culture of Taxus suspension cells and paclitaxel-producing endophytic fungi in a co-bioreactor. Appl Microbiol Biotechnol 83(2):233–239. https://doi.org/10.1007/s00253-009-1856-4
Li ZF, Wang LF, Feng ZL, Zhao LH, Shi YQ, Zhu HQ (2014) Diversity of endophytic fungi from different Verticillium–wilt-resistant Gossypium hirsutum and evaluation of antifungal activity against Verticillium dahliae in vitro. J Microbiol Biotechnol 24(9):1149–1161
Liu JY, Huang LL, Ye YH, Zou WX, Guo ZJ, Tan RX (2006) Antifungal and new metabolites of Myrothecium sp. Z16, a fungus associated with white croaker Argyrosomus argentatus. J Appl Microbiol 100(1):195–202. https://doi.org/10.1111/j.1365-2672.2005.02760.x
Liu S, Dai H, Orfali RS, Lin W, Liu Z, Proksch P (2016) New fusaric acid derivatives from the endophytic fungus Fusarium oxysporum and their phytotoxicity to barley leaves. J Agric Food Chem 64(16):3127–3132. https://doi.org/10.1021/acs.jafc.6b00219
Martin LL, Ross Friedman CM, Phillips LA (2012) Fungal endophytes of the obligate parasitic dwarf mistletoe Arceuthobium americanum (Santalaceae) act antagonistically in vitro against the native fungal pathogen Cladosporium (Davidiellaceae) of their host. Am J Bot 99(12):2027–2034. https://doi.org/10.3732/ajb.1200189
Mosquera-Espinosa AT, Bayman P, Prado GA, Gomez-Carabali A, Otero JT (2013) The double life of Ceratobasidium: orchid mycorrhizal fungi and their potential for biocontrol of Rhizoctonia solani sheath blight of rice. Mycologia 105(1):141–150. https://doi.org/10.3852/12-079
Niu Y, Li C, Pan Y, Li Y, Kong X, Wang S, Zhai Y, Wu X, Fan W, Mei Q (2015) Treatment of Radix Dipsaci extract prevents long bone loss induced by modeled microgravity in hindlimb unloading rats. Pharm Biol 53(1):110–116. https://doi.org/10.3109/13880209.2014.911920
Pfurtscheller G, Klimesch W (1990) Topographical display and interpretation of event-related desynchronization during a visual-verbal task. Brain Topogr 3(1):85–93
Puri SC, Nazir A, Chawla R, Arora R, Riyaz–Ul–Hasan S, Amna T, Ahmed B, Verma V, Singh S, Sagar R, Sharma A, Kumar R, Sharma RK, Qazi GN (2006) The endophytic fungus Trametes hirsuta as a novel alternative source of podophyllotoxin and related aryl tetralin lignans. J Biotechnol 122(4):494–510. https://doi.org/10.1016/j.jbiotec.2005.10.015
Qin S, Xing K, Jiang JH, Xu LH, Li WJ (2011) Biodiversity, bioactive natural products and biotechnological potential of plant-associated endophytic actinobacteria. Appl Microbiol Biotechnol 89(3):457–473. https://doi.org/10.1007/s00253-010-2923-6
Redman RS, Sheehan KB, Stout RG, Rodriguez RJ, Henson JM (2002) Thermotolerance generated by plant/fungal symbiosis. Science 298(5598):1581. https://doi.org/10.1126/science.1072191
Rodriguez R, Redman R (2008) More than 400 million years of evolution and some plants still can’t make it on their own: plant stress tolerance via fungal symbiosis. J Exp Bot 59(5):1109–1114. https://doi.org/10.1093/jxb/erm342
Ruma K, Sunil K, Kini KR, Prakash HS (2015) Genetic diversity and antimicrobial activity of endophytic Myrothecium spp. isolated from Calophyllum apetalum and Garcinia morella. Mol Biol Rep 42(11):1533–1543. https://doi.org/10.1007/s11033-015-3884-8
Salehi A, Mohammadi M, Okhovvat SM, Omidi M (2005) Chitinase gene transformation through Agrobacteriumand its explanation in soybean in order to induce resistance to root rot caused by Rhizoctonia solani. Commun Agric Appl Biol Sci 70(3):399–406
Schmidt CS, Mrnka L, Frantik T, Lovecka P, Vosatka M (2018) Plant growth promotion of Miscanthus × giganteus by endophytic bacteria and fungi on non-polluted and polluted soils. World J Microbiol Biotechnol 34(3):48. https://doi.org/10.1007/s11274-018-2426-7
Shah SP, Kalia KS, Patel JS (2015) Optimization of cellulase production by Penicillium oxalicum using banana agrowaste as a substrate. J Gen Appl Microbiol 61(2):35–43. https://doi.org/10.2323/jgam.61.35
Sheng-Liang Z, Shu-Zhen Y, Zhen-Ying W, Shuang-Lin C (2014) Endophytic fungi associated with Macrosolen tricolor and its host Camellia oleifera. World J Microbiol Biotechnol 30(6):1775–1784. https://doi.org/10.1007/s11274-014-1600-9
Soca-Chafre G, Rivera-Orduna FN, Hidalgo-Lara ME, Hernandez-Rodriguez C, Marsch R, Flores-Cotera LB (2011) Molecular phylogeny and paclitaxel screening of fungal endophytes from Taxus globosa. Fungal Biol 115(2):143–156. https://doi.org/10.1016/j.funbio.2010.11.004
Stamford TL, Stamford NP, Coelho LC, Araujo JM (2001) Production and characterization of a thermostable alpha-amylase from Nocardiopsis sp. endophyte of yam bean. Bioresour Technol 76(2):137–141
Suzuki M, Nishida N, Ishihara A, Nakajima H (2013) New 3-O-alkyl-4a,10a-dihydrofusarubins produced by Fusarium sp. Mj-2. Biosci Biotechnol Biochem 77(2):271–275. https://doi.org/10.1271/bbb.120670
Takahashi C, Numata A, Matsumura E, Minoura K, Eto H, Shingu T, Ito T, Hasegawa T (1994) Leptosins I and J, cytotoxic substances produced by a Leptosphaeria sp. Physico-chemical properties and structures. J Antibiot 47(11):1242–1249
Thongekkaew J, Ikeda H, Masaki K, Iefuji H (2013) Fusion of cellulose binding domain from Trichoderma reesei CBHI to Cryptococcus sp. S-2 cellulase enhances its binding affinity and its cellulolytic activity to insoluble cellulosic substrates. Enzyme Microb Technol 52(4–5):241–246. https://doi.org/10.1016/j.enzmictec.2013.02.002
Waller F, Achatz B, Baltruschat H, Fodor J, Becker K, Fischer M, Heier T, Huckelhoven R, Neumann C, von Wettstein D, Franken P, Kogel KH (2005) The endophytic fungus Piriformospora indica reprograms barley to salt-stress tolerance, disease resistance, and higher yield. Proc Natl Acad Sci USA 102(38):13386–13391. https://doi.org/10.1073/pnas.0504423102
Walsh CT, Fischbach MA (2010) Natural products version 2.0: connecting genes to molecules. J Am Chem Soc 132(8):2469–2493. https://doi.org/10.1021/ja909118a
Wang JY, Liang YL, Hai MR, Chen JW, Gao ZJ, Hu QQ, Zhang GH, Yang SC (2016) Genome-wide transcriptional excavation of Dipsacus asperoides Unmasked both Cryptic Asperosaponin biosynthetic genes and SSR markers. Front Plant Sci 7:339. https://doi.org/10.3389/fpls.2016.00339
Wang X, Li Y, Song X, Meng Q, Zhu J, Zhao Y, Yu W (2017) Influence of host tree species on isolation and communities of mycorrhizal and endophytic fungi from roots of a tropical epiphytic orchid, Dendrobium sinense (Orchidaceae). Mycorrhiza 27(7):709–718. https://doi.org/10.1007/s00572-017-0787-7
Winter JM, Behnken S, Hertweck C (2011) Genomics-inspired discovery of natural products. Curr Opin Chem Biol 15(1):22–31. https://doi.org/10.1016/j.cbpa.2010.10.020
Wong RW, Rabie AB, Hagg EU (2007) The effect of crude extract from Radix Dipsaci on bone in mice. Phytother Res 21(6):596–598. https://doi.org/10.1002/ptr.2126
Zhang Y, Kiyohara H, Matsumoto T, Yamada H (1997) Fractionation and chemical properties of immunomodulating polysaccharides from roots of Dipsacus asperoides. Planta Medica 63(5):393–399. https://doi.org/10.1055/s-2006-957720
Zhang ZJ, Qian YH, Hu HT, Yang J, Yang GD (2003) The herbal medicine Dipsacus asper wall extract reduces the cognitive deficits and overexpression of beta–amyloid protein induced by aluminum exposure. Life Sci 73(19):2443–2454
Zhang R, Li P, Xu L, Chen Y, Sui P, Zhou L, Li J (2009) Enhancement of diosgenin production in Dioscorea zingiberensis cell culture by oligosaccharide elicitor from its endophytic fungus Fusarium oxysporum Dzf17. Nat Prod Commun 4 (11):1459–1462
Zhao XH, Wang W, Tong B, Zhang SP, Wei DZ (2016) A newly isolated Penicillium oxalicum 16 cellulase with high efficient synergism and high tolerance of monosaccharide. Appl Biochem Biotechnol 178(1):173–183. https://doi.org/10.1007/s12010-015-1866-x
Zheng L, Cai Y, Zhou L, Huang P, Ren X, Zuo A, Meng X, Xu M, Liao X (2017) Benzoquinone from Fusarium pigment inhibits the proliferation of estrogen receptor-positive MCF-7 cells through the NF-kappaB pathway via estrogen receptor signaling. Int J Mol Med 39(1):39–46. https://doi.org/10.3892/ijmm.2016.2811
Zuccaro A, Lahrmann U, Guldener U, Langen G, Pfiffi S, Biedenkopf D, Wong P, Samans B, Grimm C, Basiewicz M, Murat C, Martin F, Kogel KH (2011) Endophytic life strategies decoded by genome and transcriptome analyses of the mutualistic root symbiont Piriformospora indica. PLoS Pathog 7(10):e1002290. https://doi.org/10.1371/journal.ppat.1002290
Acknowledgements
This research was financially supported by the Regional Science Fund of the National Natural Science Foundation of China (81160501; 81860675), the China Agriculture Research System (CARS-21), the First-class Discipline Construction Project in Guizhou Province of China [GNYL (2017) 008], and the Chinese Medicine Public Health Program of the National Administration of Traditional Chinese Medicine [Caishe (2014) No. 76]. We thank Dr. Shenghua Wei for supplying the samples, Prof. Tao Zhou for field knowledge about D. asperoides and microbiology, and Jinqiang Zhang for useful comments on the manuscript. We also thank Yongqiang Zhou for guidance during experiments.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflicts of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
11274_2019_2616_MOESM1_ESM.pptx
Fig. S1. Morphological characteristics of endophyte fungi. Photographs showing typical morphology of residual endophyte fungi in Fig. S1. Characteristics of endophytic fungi from D. asperoides taproots are shown at “surface” and “back”; microstructure is also shown. Scale bar, 20 μm. (PPTX 4076 KB)
11274_2019_2616_MOESM2_ESM.pptx
Fig. S2. Homology comparison of endophytic fungi with foaming properties. Original sequence map of homology alignment between ITS sequences from the foaming fungi (ITS’) and the ITS sequences from the original fungal isolates, showing 100% homology for all the isolates (daef 11, 15, 40, 41 and 44). The alignment was produced using BioEdit 7.1.3. (PPTX 2070 KB)
11274_2019_2616_MOESM3_ESM.docx
Table S1. Similarity between the isolates from Dipsacus asperoides and the closest species in UNITE database. The statistical table shows the homology of the rDNA-ITS sequence of isolates from D. asperoides to the closest fungal sequences in UNITE database, based on blastn analysis. The strain ID has the format: latin initials of Dipsacus asperoides, the initial letter of the endophytic fungus and the strain number. The “Accession No. “is the GenBank accession number. The “Sequences producing significant alignments (Accession No.)” indicating the analysis of sequences between the fungi isolated from Dipsacus asperoides and 1 to 3 fungi from 15 best matches (as judged by the e value) fungi obtained from UNITE blastn comparative analysis. Identities (%) is the homology (similarity) obtained by comparing the sequences between the two strains. Score (Bits) is a log-scaled version of a score. It is a rescaled version of the raw alignment score that is independent of the size of the search space. The “E-value” is correction of the p-value for multiple testing. It indicates the possibility that the similarity between other sequences and the target sequence. The lower the score, the better. (DOCX 27 KB)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Gong, A., Zhou, T., Xiao, C. et al. Association between dipsacus saponin VI level and diversity of endophytic fungi in roots of Dipsacus asperoides. World J Microbiol Biotechnol 35, 42 (2019). https://doi.org/10.1007/s11274-019-2616-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-019-2616-y