Abstract
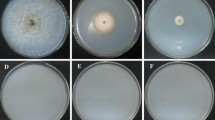
Monascus ruber, a red mold species, has been widely used in the fields of food and medicine. In this research, we transformed Monascus ruber spores using Agrobacterium tumefaciens as a tool for random insertional mutagenesis with the hygromycin phosphotransferase gene as the selected marker. Three types of mutants including citrinin-producing mutants, mutants with abnormal aerial hyphae and pigment change mutants were screened for molecular analysis. Southern blot analysis showed that more than 83.3% of transformants contained single T-DNA insertions. The genomic DNA segments of the transformants flanking the T-DNA could be amplified from their left borders with TAIL-PCR. Homologous comparison using the Blast tool showed that none of the isolated DNA sequences had any similarity to each other, suggesting that the T-DNA was randomly integrated into the fungal genome, which provided the hypothetical reason for the variant phenotypes of the transformants. The successful creation of transformants with a single T-DNA tag insertion may help us to clone functional genes related to the metabolism and differentiation of Monascus spp., which will greatly facilitate the molecular analysis of this important fungus and the improvement of strains at the genetic level.




Similar content being viewed by others
Abbreviations
- ATMT:
-
Agrobacterium tumefaciens-mediated T-DNA transformation
- TAIL-PCR:
-
Thermal asymmetric interlaced PCR
- GABA:
-
γ-Aminobutyric acid
- SON-PCR:
-
Single oligonucleotide nested PCR
References
Antal Z, Rascle C, Fèvre M et al (2004) Single oligonucleotide nested PCR: a rapid method for the isolation of genes and their flanking regions from expressed sequence tags. Curr Genet 46:240–246. doi:10.1007/s00294-004-0524-6
Blanc PJ, Laussac JP, Le Bars J et al (1995) Characterization of monascidin A from Monascus as citrinin. Int J Food Microbiol 27:201–213. doi:10.1016/0168-1605(94)00167-5
Bumpus JA, Trax M, Reisdorph A et al (2008) An in silico analysis of cytochrome c from Phanerochaete chrysosporium: its amino acid sequence and characterization of gene structural elements. In Silico Biol 8(1):1–13
Chen F, Hu X (2005) Study on red fermented rice with high concentration of Monacolin K and low concentration of citrinin. Int J Food Microbiol 103:331–337. doi:10.1016/j.ijfoodmicro.2005.03.002
Detlef W, Jane G (2002) Arabidopsis: a laboratory manual. Cold Spring Harbor Laboratory Press, New York
Ding Y, Shao Y, Xu Y et al (2006) Screening Citrinin mutants from the transformants library of Monascus ruber M-7 by Agrobacterium-mediated DNA transfer. Microbiol Chin 33(4):52–57
Fu JQ (1997) Production and application of red yeast rice (Chinese). Light Industry Press, Beijing
Fu G, Xu Y, Li Y et al (2007) Construction of a replacement vector to disrupt pksCT gene for the mycotoxin citrinin Monascus aurantiacus and maintain food red pigment production. Asia Pac J Clin Nutr 16(suppl 1):137–142
Gheith O, Sheashaa H, Abdelsalam M et al (2008) Efficacy and safety of Monascus purpureus Went rice in subjects with secondary hyperlipidemia. Clin Exp Nephrol 12:189–194. doi:10.1007/s10157-008-0033-x
Hajjaj H, Klaébé A, Loret MO et al (1999) Biosynthetic pathway of citrinin in the filamentous fungus Monascus ruber as revealed by 13C nuclear magnetic resonance. Appl Environ Microbiol 65(1):311–314
Karunakaran M, Nair V, Rho HS et al (2008) Agrobacterium tumefaciens-mediated transformation in Colletotrichum falcatum and C. acutatum. J Microbiol Biotechnol 18(2):234–241
Kim JG, Choi YD, Chang YJ et al (2003) Genetic transformation of Monascus purpureus DSM1379. Biotechnol Lett 25:1509–1514. doi:10.1023/A:1025438701383
Lakrod K, Chaisrisook C, Daniel ZS (2003a) Transformation of Monascus purpureus to hyromycing B resistance with cosmid pMOcosX reduces fertility. Electron J Biotechnol 6(2):143–147
Lakrod K, Chaisrisook C, Skinner DZ (2003b) Expression of pigmentation genes following electroporation of albino Monascus purpureus. J Ind Microbiol Biotechnol 30:369–374. doi:10.1007/s10295-003-0058-9
Lamarre C, Ibrahim-Granet O, Du C et al (2007) Characterization of the SKN7 ortholog of Aspergillus fumigatus. Fungal Genet Biol 44:682–690. doi:10.1016/j.fgb.2007.01.009
Li MX, Gong XY, Zheng J et al (2005) Transformation of Coniothyrium minitans, a parasite of Sclerotinia sclerotiorum, with Agrobacterium tumefaciens. FEMS Microbiol Lett 243:323–329. doi:10.1016/j.femsle.2004.12.033
Li YP, Tan WH, Xu Y (2007) Transformation of protoplast Monascus aurantiacus AS3.4384. Food Sci (Chinese) 28(10):317–321
Lin YL, Wang TH, Lee MH, Su NW (2008a) Biologically active components and nutraceuticals in the Monascus-fermented rice: a review. Appl Microbiol Biotechnol 77:965–973. doi:10.1007/s00253-007-1256-6
Lin CP, Chen YH, Chen JW et al (2008b) Cholestin (Monascus purpureus rice) inhibits homocysteine- induced reactive oxygen species generation, nuclear factor-κB activation, and vascular cell adhesion moleculer expression in human aortic endothelial cells. J Biomed Sci 15:183–196. doi:10.1007/s11373-007-9212-0
Mandt M (1998) Legal opinion on the use of red yeast rice(Angkak) in food. Paper collection for a symposia of Monascus cultures and applications, held from July 8th to 10th, 1998 in Toulouse, France (Monascus purpureus)
Michielse CB, Hooykaas PJ, van den Hondel CA et al (2005) Agrobacterium-mediated transformation as a tool for functional genomics in fungi. Curr Genet 48:1–17. doi:10.1007/s00294-005-0578-0
Ruiz-Herrera J, Gonzalez-Prieto JM, Ruiz-Medrano R (2002) Evolution and phylogenetic relationships of chitin synthases from yeasts and fungi. FEM Yeast Res 1(4):247–256. doi:10.1111/j.1567-1364.2002.tb00042.x
Sambrook J, Russell DW (2001) Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory Press, NewYork
Shao YC, Wang RY, Ding YD, Chen FS, Xie BJ (2006) Construction of T-DNA insertional library of Monascus mediated by Agrobacterium tumefaciens and characteristic analysis of the color mutants. Mycosystema (Chinese) 25(2):247–255
Shao YC, Ding YD, Chen FS et al (2007) Isolation of DNA sequence flanking T-DNA by thermal asymmetric interlaced PCR from Monascus pigment producing mutants. Microbiology (Chinese) 34(2):323–326
Shimizu T, Kinoshita H, Ishihara S et al (2005) Polyketide synthase gene responsible for citrinin biosynthesis in Monascus purpureus. Appl Environ Microbiol 7:3453–3457. doi:10.1128/AEM.71.7.3453-3457.2005
Shimizu T, Kinoshita H, Nihira T (2007) Identification and in vivo functional analysis by gene disruption of ctnA, an activator gene involved in citrinin biosynthesis in Monascus purpureus. Appl Environ Microbiol 73(16):5097–5103. doi:10.1128/AEM.01979-06
Sonia C, Flor P, Martin JF et al (2003) Stable transformants of the azaphilone pigment-producing Monascus purpureus obtained by protoplast transformation and Agrobacterium-mediated DNA transfer. Curr Genet 43:447–452. doi:10.1007/s00294-003-0417-0
Talhinhas P, Muthumeenakshi S, Martins JN et al (2008) Agrobacterium-mediated transformation and insertional mutagenesis in Colletotrichum acutatum for investigating varied pathogenicity lifestyles. Mol Biotechnol 39:57–67. doi:10.1007/s12033-007-9028-1
Tanaka T, Tateno Y, Gojobori T (2005) Evolution of vitamin B6 (pyridoxine) metabolism by gain and loss of genes. Mol Biol Evol 22(2):243–250. doi:10.1093/molbev/msi011
Wang JY, Li HY (2008) Agrobacterium tumefaciens-mediated genetic transformation of the phytopathogenic fungus Penicillium digitatum. J Zhejiang Univ Sci B 9:823–828. doi:10.1631/jzus.B0860006
Wang JJ, Lee CL, Pan TM (2004) Modified mutation method for screening low citrinin-producing strains of Monascus purpureus on Rice Culture. J Agric Food Chem 52:6977–6982. doi:10.1021/jf049783o
Wieser J, Lee BN, John W, Fondon THIII (1994) Genetic requirements for initiating asexual development in Aspergillus nidulans. Curr Genet 27:62–69. doi:10.1007/BF00326580
Yang YJ, Lee I (2008) Agrobactrium tumefaciens-mediated transformation of Monascus ruber. J Microbiol Biotechnol 18(4):754–758
Yu JH (2006) Heterotrimeric G protein signaling and RGSs in Aspergillus nidulans. J Microbiol 44(2):145–154
Yu JH, Keller N (2005) Regulation of secondary metabolism in filamentous fungi. Annu Rev Phytopathol 43:437–458. doi:10.1146/annurev.phyto.43.040204.140214
Acknowledgments
Thanks are due to Dr. Daohong Jiang for his technical assistance and for kindly providing pTFCM for this experiment. This work was financially supported by the National High Technology Research and Development Program of the People’s Republic of China (863 program:No2006AA10Z1A3) and the Financial Aid Program for New Century Talents by the Ministry of Education of the People’s Republic of China (NoNCET-05-0667)
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Shao, Y., Ding, Y., Zhao, Y. et al. Characteristic analysis of transformants in T-DNA mutation library of Monascus ruber . World J Microbiol Biotechnol 25, 989–995 (2009). https://doi.org/10.1007/s11274-009-9977-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11274-009-9977-6