Abstract
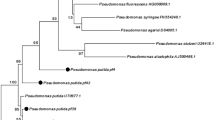
In this study, Pseudomonas species were isolated from the rhizospheres of two plant hosts: rice (Oryza sativa cultivar Pathum Thani 1) and maize (Zea mays cultivar DK888). The genotypic diversity of isolates was determined on basis of amplified rDNA restriction analysis (ARDRA). This analysis showed that both plant varieties selected for two distinct populations of Pseudomonas. The actual biocontrol and plant promotion abilities of these strains was confirmed by bioassays on fungal (Verticillum sp., Rhizoctonia solani and Fusarium sp.) and bacterial (Ralstonia solanacearum and Bacillus subtilis) plant pathogens, as well as indole-3-acetic acid (IAA) production and carbon source utilization. There was a significant difference between isolates from rice and maize rhizosphere in terms of biological control against R. solanacearum and B. subtilis. Interestingly, none of the pseudomonads isolated from maize rhizosphere showed antagonistic activity against R. solanacearum. This study indicated that the percentage of pseudomonad isolates obtained from rice rhizosphere which showed the ability to produce fluorescent pigments was almost threefold higher than pseudomonad isolates obtained from maize rhizosphere. Furthermore, the biocontrol assay results indicated that pseudomonad isolated from rice showed a higher ability to control bacterial and fungal root pathogens than pseudomonad isolates obtained from maize. This work clearly identified a number of isolates with potential for use as plant growth-promoting and biocontrol agents on rice and maize.



Similar content being viewed by others
References
Abbas HK, Cartwright RD, Shier WT, Abouzied MM, Bird CB, Rice LG, Ross PF, Sciumbato GL, Meredith FI (1998) Natural occurrence of fumonisins in rice with Fusarium sheath rot disease. Plant Dis 82:22–25
Bakker PA, Glandorf DC, Viebahn M, Ouwens TW, Smit E, Leeflang P, Wernars K, Thomashow LS, Thomas-Oates JE, van Loon LC (2002) Effects of Pseudomonas putida modified to produce phenazine-1-carboxylic acid and 2,4-diacetylphloroglucinol on the microflora of field grown wheat. Antonie van Leeuwenhoek 81:617–624
Berg G, Roskot N, Steidle A, Eberl L, Zock A, Smalla K (2002) Plant-dependent genotypic and phenotypic diversity of antagonistic rhizobacteria isolated from different Verticillium host plants. Appl Environ Microbiol 68:3328–3338
Cirvilleri G, Spina S, Scuderi G, Gentile A, Catara A (2005) Characterization of antagonistic root-associated fluorescent pseudomonads of transgenic and non-transgenic citrange troyer plants. J Plant Pathol 87:179–186
Desnoues N, Lin M, Guo X, Ma L, Carreño-Lopez R, Elmerich C (2003) Nitrogen fixation genetics and regulation in a Pseudomonas stutzeri strain associated with rice. Microbiology 149:2251–2262
Fenton AM, Stephens PM, Crowley J, O’Callaghan M, O’Gara F (1992) Exploitation of gene(s) involved in 2,4-diacetylphloroglucinol biosynthesis to confer a new biocontrol capability to a Pseudomonas strain. Appl Environ Microbiol 58:3873–3878
Franks A, Ryan RP, Abbas A, Mark GL, O’Gara F (2006) Molecular tools for studying plant growth-promoting rhizobacteria (PGPR). In Molecular techniques for soil and rhizosphere microorganisms. CABI Publishing, Wallingford, Oxfordshire, UK
Haas D, Défago G (2005) Biological control of soil-borne pathogens by fluorescent pseudomonads. Nat Rev Microbiol 3:307–319
Inagaki K (1998) Dispersal of rice sheath blight fungus, Rhizoctonia solani AG-1(IA), and subsequent disease development in paddy fields, from survey of vegetative compatibility groups. Mycoscience 39:391–397
Keel C, Weller DM, Natsch A, Défago G, Cook RJ, Thomashow LS (1996) Conservation of the 2,4-diacetylphloroglucinol biosynthesis locus among fluorescent Pseudomonas strains from diverse geographic locations. Appl Environ Microbiol 62:552–563
Kiely DP, Haynes JM, Higgins CH, Franks A, Mark GL, Morrissey JP, O’Gara F (2006) Exploiting new systems-based strategies to elucidate plant-bacterial interactions in the rhizosphere. Microb Ecol 51:257–266
Lipman JG (1904) Soil bacteriological studies. Further contributions to the physiology and morphology of the members of the Azotobacter group. Report of the New Jersey State Agricultural Experiment Station 25:237–289
McSpadden Gardener BB, Mavrodi DV, Thomashow LS, Weller DM (2001) A rapid polymerase chain reaction-based assay characterizing rhizosphere populations of 2,4-diacetylphloroglucinol-producing bacteria. Phytopathology 91:44–54
Moënne-Loccoz Y, Tichy H-V, O’Donnell A, Simon R, O’Gara F (2001) Impact of 2,4-diacetylphloroglucinol producing biocontrol strain Pseudomonas fluorescens F113 on intraspecific diversity of resident culturable fluorescent pseudomonads associated with the roots of field-grown sugar beets seedlings. Appl Environ Microbiol 67:3418–3425
Morrissey JP, Dow JM, Mark GL, O’Gara F (2004) Are microbes at the root of a solution to world food production? EMBO Rep 5:922–926
Nielsen MN, Sorensen J, Fels J, Pedersen HC (1998) Secondary metabolite- and endochitinase-dependent antagonism toward plant-pathogenic microfungi of Pseudomonas fluorescens isolates from sugar beet rhizosphere. Appl Environ Microbiol 64:3563–3569
NSW Department of Environment and Climate Change, Sydney Australia (2007) Botanic gardens trust. http://www.rbgsyd.nsw.gov.au/. Cited 28 May 2007
Nuntagij A, Abe M, Uchimi T, Seki Y, Boonkerd N, Higashi S (1997) Characterization of bradyrhizobium strains isolated from soybean cultivation in Thailand. J Gen Appl Microbiol 43:183–187
O’Sullivan DJ, O’Gara F (1992) Traits of fluorescent Pseudomonas spp. involved in suppression of plant root pathogens. Microbiol Rev 56:662–676
Picard C, Frascaroli E, Bosco M (2004) Frequency and biodiversity of 2,4 diacetylphloroglucinol-producing rhizobacteria are differentially affected by the genotype of two maize inbred lines and their hybrid. FEMS Microbiol Ecol 49:207–215
Raaijmakers JM, Weller DM (2001) Exploiting genotypic diversity of 2,4-diacetylphloroglucinol-producing Pseudomonas spp.: characterization of superior root-colonizing P. fluorescens strain Q8r1-96. Appl Environ Microbiol 67:2545–2554
Rangarajan S, Saleena LM, Nair S (2002) Diversity of Pseudomonas spp. isolated from rice rhizosphere populations grown along a salinity gradient. Microb Ecol 43:280–289
Rodriguez F, Pfender WF (1997) Antibiosis and antagonism of Sclerotinia homoeocarpa and Drechslera poae by Pseudomonas fluorescens PF-5 in vitro and in planta. Phytopathology 87:614–621
Ross IL, Alami Y, Harvey PR, Achouak W, Ryder MH (2000) Genetic diversity and biological control activity of novel species of closely related pseudomonads isolated from wheat field soils in south Australia. Appl Environ Microbiol 66:1609–1616
Sambrook J, Russell DW (2001) Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory, New York
Sharifi-Tehrani A, Zala M, Natsch A, Moënne-Loccoz Y, Défago G (1998) Biocontrol of soil-borne fungal plant diseases by 2,4-diacetylphloroglucinol-producing fluorescent pseudomonads with different restriction profiles of amplified 16S rDNA. Eur J Plant Pathol 104:631–643
Shurtleff MC, Edwards DI, Noel GR, Pederson WL, White DG (1993) Diseases of corn or maize (Zea mays L.). In: Common names of plant diseases. The American Phytopathological Society. Available via DIALOG. http://www.apsnet.org/online/common/names/corn.asp. Cited 27 May 2007
Weisburg WG, Barns SM, Pelletier DA, Lane DJ (1991) 16S ribosomal DNA amplification for phylogenetic study. J Bacteriol 173:697–703
Acknowledgements
This work was supported by a grant from the Royal Golden Jubilee Ph.D. program (Thailand Research Fund) and Suranaree University of Technology. F.O. was supported in part by grants awarded by Science Foundation Ireland (SFI 02/IN.1/B1261 and 04/BR/B0597 to F.O.) and the Health Research Board (RP/2004/145 and RP/2006/271 to F.O.).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Lawongsa, P., Boonkerd, N., Wongkaew, S. et al. Molecular and phenotypic characterization of potential plant growth-promoting Pseudomonas from rice and maize rhizospheres. World J Microbiol Biotechnol 24, 1877–1884 (2008). https://doi.org/10.1007/s11274-008-9685-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11274-008-9685-7