Abstract
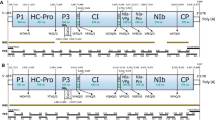
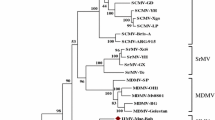
The complete nucleotide sequence of the genome of wheat Eqlid mosaic virus (WEqMV) (excluding the poly A tail) comprised 9636 nucleotides including 5′ and 3′ noncoding regions of 137 and 172 nt, respectively. It contained a single ORF coding for a polyprotein of 3,109 amino acid residues and had a deduced genome organization typical of members of the family Potyviridae and with proteinase cleavage sites very similar to those of the members of the genus Tritimovirus. Pairwise and multiple alignments and phylogenetic analysis showed that WEqMV is a distinct species in the genus Tritimovirus. WEqMV and Wheat streak mosaic virus (WSMV) shared the greatest nucleotide sequence identity in the NIb and HC-Pro cistrons (63.2% and 60.8%, respectively) and the lowest sequence identity in the P1 and CP cistrons (51.2% and 51.1%, respectively). Sequence identity for the complete genome of WEqMV and WSMV was 56.8% at the nucleotide level and 50.7% at the amino acid level. WEqMV had 57.2% nucleotide identity and 50.6% amino acid identity with Oat necrotic mottle virus and 52.5% nucleotide identity and 45.5% amino acid identity with Brome streak mosaic virus. The relationship of WEqMV with other members of the family Potyviridae was more distant. Structural analysis of WEqMV protein showed presence of potential transmembrane helices in 6k1, 6k2, and P3 proteins.





Similar content being viewed by others
References
F. Rabenstein, J. Schubert, Genus Rymovirus. in Virus and Virus Diseases of Poaceae, (INRA Publication, Paris, 2004), pp. 395–401
S.N. Salm, M.E.C. Rey, E.P. Rybicki, Arch. Virol. 141, 2237–2242 (1996). doi:https://doi.org/10.1007/BF01718229
D.C. Stenger, J.S. Hall, L.R. Choi, R. French, Phytopathology 88, 782–787 (1998). doi:https://doi.org/10.1094/PHYTO.1998.88.8.782
D.C. Stenger, R. French, Arch. Virol. 149, 633–640 (2004). doi:https://doi.org/10.1007/s00705-003-0237-z
K. Izadpanah, R. Kamran, in Proc. 12th Iran Plant Protec. Cong., 1995, p. 29
M. Masumi, M. Rastegar, A. Zare, D.-E. Lesemann, F. Ebrahim-Nesbat, K. Izadpanah, Parasitica 61(1), 101–104 (2005)
M. Masumi, K. Izadpanah, Iran. J. Plant Pathol. 38, 35–37 (2002)
P. Froussard, Nucleic Acids Res. 20, 2900 (1992). doi:https://doi.org/10.1093/nar/20.11.2900
R. Kalender, Fast PCR: A program for fast design of PCR primer, DNA and protein manipulation (Version 2.5.59). Institute of Biotechnology, University of Helsinki (2003)
J.D. Thompson, T.J. Gibson, F. Plewniak, F. Jeanmougin, D.G. Higgins, Nucleic Acids Res. 24, 4876–4882 (1997). doi:https://doi.org/10.1093/nar/25.24.4876
N. Saitou, M. Nei. Mol. Biol. Evol. 4, 406–425 (1987)
J. Felsenstein, G.A. Churchill, Mol. Biol. Evol. 13, 93–104 (1996)
R.D.M. Page, Comput. Appl. Biosci. 12, 357–358 (1996)
B. Roast, R. Casadio, P. Fariselli, C. Sander, Protein Sci. 4, 521–533 (1995)
E.L. Sonnhammer, G. von Heijne, A. Krogh, Sixth international conference on intelligent systems for molecular biology (ISMB 98), 1998, pp. 175–182
A. Lupas, Methods Enzymol. 266, 513–525 (1996)
M.J. Adams, J.F. Antoniw, Arch. Virol. 149, 113–135 (2004)
M.J. Adams, J.F. Antoniw, F. Beaudoin, Mol. Plant Pathol. 6, 471–487 (2005). doi:https://doi.org/10.1111/j.1364-3703.2005.00296.x
J. Badge, D.J. Robinson, A.A. Brunt, G.D. Foster, J. Gen. Virol. 78, 253–257 (1997)
M. Masumi, K. Izadpanah, D.-E. Lesemann, in Proc. 14th Iran Plant Protec. Cong., vol. II, 2000a, p. 20
A. Lupas, Curr. Opin. Struct. Biol. 7, 388–393 (1997). doi:https://doi.org/10.1016/S0959-440X(97)80056-5
W.Y. Chung, W.A. Miller, J.F. Atkins, A.E. Firth, Proc. Natl. Acad. Sci. USA 105, 5897–5902 (2008). doi:https://doi.org/10.1073/pnas.0800468105
M. Masumi, K. Izadpanah, F. Ebrahim-Nesbat, in Proc. 14th Iran Plant Protec. Cong., vol. II, 2000b, p. 231
M. Hema, J. Joseph, K. Gopinath, P. Sreenivasulu, H.S. Savithri, Arch. Virol. 144, 479–490 (1999). doi:https://doi.org/10.1007/s007050050519
C. Fauquet, A. Mayo, Arch. Virol. 144, 1249–1273 (1999). doi:https://doi.org/10.1007/s007050050584
E. Johansen, O.F. Rasmussen, M. Heide, B. Borkhardt, J. Gen. Virol. 72, 2625–2632 (1991). doi:https://doi.org/10.1099/0022-1317-72-11-2625
Y. Takimoto, T. Hibi, Virology 332, 199–205 (2005). doi:https://doi.org/10.1016/j.virol.2004.11.026
S. Gazzarrini, M. Kang, S. Epimashko, J.L. Van Etten, J. Dainty, G. Teil et al., Proc. Natl. Acad. Sci. USA 103, 5355–5360 (2006). doi:https://doi.org/10.1073/pnas.0600848103
Acknowledgment
This research was partially supported by funds from Center of Excellence in Plant Virology and the Iranian Chapter of TWAS (ISMO).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Rastegar, M., Izadpanah, K., Masumi, M. et al. Analyses of the complete sequence of the genome of wheat Eqlid mosaic virus, a novel species in the genus Tritimovirus . Virus Genes 37, 212–217 (2008). https://doi.org/10.1007/s11262-008-0252-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11262-008-0252-y