Abstract
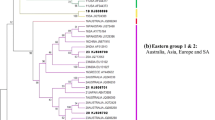
The nucleotide sequences of the complete VP1, VP0, VP3, and partial 3D regions of seven duck hepatitis virus (DHV) serotype 1 (DHV-1) strains isolated in China between 2001 and 2007 and one DHV-1 strain originally obtained from ATCC were determined and compared with previously available DHV sequences in GenBank. Phylogenetic analysis on the basis of VP1 sequences demonstrated three distinct genetic groups. There was an excellent concordance among the genetic groups assigned based on the complete VP0, VP3, and the partial 3D regions. In view of the growing importance of molecular techniques in diagnosis, we propose that the three genetic groups should be termed DHV types A, B, and C. All DHV-1 strains grouped in genotype A, whereas the new serotype strains isolated in Taiwan and the new serotype strains isolated in South Korea clustered into genotypes B and C, respectively, suggesting a potential genetic correlates of serotype. In pairwise comparisons of complete VP1, VP0, and VP3 nucleotide and amino acid sequences and the partial 3D nucleotide sequence, DHVs of the same genotype were clearly distinguished from those of heterologous genotypes. Analysis of the amino acid sequences of the three capsid proteins demonstrated the presence of conserved elements that form the eight-stranded β-barrel structures, as well as intervening domains that vary in sequence between strains of different genotypes as seen in other picorna viruses.


Similar content being viewed by others
References
P.R. Woolcock, in Diseases of Poultry, ed. by Y.M. Saif, H.J. Barnes, J.R. Glisson, A.M. Fadly, L.R. McDougald, D.E. Swayne (Iowa State Press, Ames, 2003). pp. 343–354
P.P. Levine, J. Fabricant, Cornell. Vet. 40, 71 (1950)
F.D. Asplin, Vet. Rec. 77, 1529 (1965)
R.E. Gough, M.S. Collins, E. Borland, I.F. Keymer, Vet. Rec. 114, 279 (1984)
R.E. Gough, E.D. Borland, I.F. Keymer, J.C. Stuart, Avian Pathol. 14, 227 (1985). doi:https://doi.org/10.1080/03079458508436224
T.E. Toth, Avian Dis. 13, 834 (1969). doi:https://doi.org/10.2307/1588590
S.A. Haider, B.W. Calnek, Avian Dis. 23, 715 (1979). doi:https://doi.org/10.2307/1589748
G. Stanway, F. Brown, P. Christian, T. Hovi, T. Hyypiä, A.M.Q. King, N.J. Knowles, S.M. Lemon, P.D. Minor, M.A. Pallansch, A.C. Palmenberg, T. Skern, in Virus Taxonomy ed. by C.M. Fauquet, A. Mayo, J. Maniloff, U. Desselberger, L.A. Ball (Academic Press, San Diego, CA, 2005), pp. 757–778
S.S. Monroe, M.J. Carter, J. Herrmann, D.K. Mitchel, A. Sanchez-Fauquier, in Virus Taxonomy, ed. by C.M. Fauquet, M.A. Mayo, J. Maniloff, U. Desselberger, L.A. Ball (Academic Press, San Diego, CA, 2005), pp. 859–864
C.H. Tseng, H.J. Tsai, Virus Res. 126, 19 (2007). doi:https://doi.org/10.1016/j.virusres.2007.01.012
M.C. Kim, Y.K. Kwon, S.J. Joh, S.J. Kim, C. Tolf, J.H. Kim, H.W. Sung, A.M. Lindberg, J.H. Kwon, Arch. Virol. 152, 2059 (2007). doi:https://doi.org/10.1007/s00705-007-1023-0
C. Ding, D. Zhang, Virology 361, 9 (2007). doi:https://doi.org/10.1016/j.virol.2007.01.007
M.C. Kim, Y.K. Kwon, S.J. Joh, A.M. Lindberg, J.H. Kwon, J.H. Kim, S.J. Kim, J. Gen. Virol. 87, 3307 (2006). doi:https://doi.org/10.1099/vir.0.81804–0
C.H. Tseng, N.J. Knowles, H.J. Tsai, Virus Res. 123, 190 (2007). doi:https://doi.org/10.1016/j.virusres.2006.09.007
C.U.T. Hellen, S. de Breyne, J. Virol. 81, 5850 (2007). doi:https://doi.org/10.1128/JVI.02403–06
J.D. Thompson, D.G. Higgins, T.J. Gibson, Nucl. Acids. Res. 22, 4673 (1994). doi:https://doi.org/10.1093/nar/22.22.4673
K.B. Nicholas, H.B. Jr Nicholas, D.W. Deerfield, EMBNEW. News 4, 14 (1997)
K. Tamura, J. Dudley, M. Nei, S. Kumar, Mol. Biol. Evol. 24, 1596 (2007). doi:https://doi.org/10.1093/molbev/msm092
M.S. Oberste, K. Maher, D.R. Kilpatrick, M.R. Flemister, B.A. Brown, M.A. Pallansch, J. Clin. Microbiol. 37, 1288 (1999)
M.S. Oberste, K. Maher, D.R. Kilpatrick, M.A. Pallansch, J. Virol. 73, 1941 (1999)
E.V. Koonin, J. Gen. Virol. 72, 2197 (1991)
R. Acharya, E. Fry, D. Stuart, G. Fox, D. Rowlands, F. Brown, Nature 337, 709 (1989). doi:https://doi.org/10.1038/337709a0
J.M. Hogle, M. Chow, D.J. Filman, Science 229, 1358 (1985). doi:https://doi.org/10.1126/science.2994218
M. Luo, G. Vriend, G. Kamer, L. Minor, E. Arnold, M.G. Rossmann, U. Boege, D.G. Scraba, G.M. Duke, A.C. Palmenberg, Science 235, 182 (1987). doi:https://doi.org/10.1126/science.3026048
M.G. Rossmann, E. Arnold, J.W. Erickson, E.A. Frankenberger, J.P. Griffith, H.J. Hecht, J.E. Johnson, G. Kamer, M. Luo, A.G. Mosser, Nature 317, 145 (1985). doi:https://doi.org/10.1038/317145a0
L.E. Hanson, H.E. Rhoades, R.L. Schricker, Avian Dis. 8, 196 (1964). doi:https://doi.org/10.2307/1588060
T.E. Toth, Avian Dis. 14, 697 (1970). doi:https://doi.org/10.2307/1588641
M.G. Mateu, Virus Res. 38, 1 (1995). doi:https://doi.org/10.1016/0168–1702(95)00048-U
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wang, L., Pan, M., Fu, Y. et al. Classification of duck hepatitis virus into three genotypes based on molecular evolutionary analysis. Virus Genes 37, 52–59 (2008). https://doi.org/10.1007/s11262-008-0233-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11262-008-0233-1