Abstract
The presented study analyses the influence of salicylic acid (SA) and the synthetic auxin 1-naphthaleneacetic acid (NAA) on total tanshinone level and on dihydrotanshinone (DHT), cryptotanshinone (CT), tanshinine I (TI) and tanshinone IIA (TIIA) level in Salvia miltiorrhiza callus cultures growing on solid Murashige and Skoog (MS) medium. The influence of SA and NAA was evaluated at 10-day intervals throughout a 80-day treatment period. SA was applied at 0.1, 0.2 and 0.4 mM, and NAA at 2.69, 13.43, 26.85 and 40.28 μM. DHT, CT, TI and TIIA concentrations were measured using HPLC. NAA did not increase the concentration of any tanshinone. SA increased content in a concentration- and time-dependent manner; however, the yields were relatively low, possibly due to the metabolic specificity of S. miltiorrhiza cultivars in Poland. Total tanshinone concentration reached 226.38 ± 37.33 μg g−1 DW after 50 days of 0.4 mM SA elicitation. After 50 days of SA elicitation, the following maximum tanshinone concentrations were observed for 0.4 mM SA: DHT (71.58 ± 12.72 μg g−1 DW), CT (108.54 ± 18.29 μg g−1 DW), TI (29.50 ± 4.13 μg g−1 DW) and TIIA (16.75 ± 2.74 μg g−1 DW). To account for these observed differences in tanshinone biosynthesis, the distribution of SA and auxin responsive cis-active motif in the proximal promoters of the mevalonic acid, methylerythritol-4-phosphate and tanshinone-precursor biosynthesis pathway genes was evaluated in A. thaliana and S. miltiorrhiza. Our findings indicate that the SA-responsive cis-active elements have a much broader distribution than those recognized by auxin-responsive transcription factors.
Key message
Tanshinone concentration in S. miltiorrhiza callus growing on solid, modified MS medium may by increased by salicylic acid elicitation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Salvia miltiorrhiza, also known as Danshen/Tanshen, is a medicinal plant that has been used in Chinese medicine since ancient times (Su et al. 2015). It is applied to treat ischemic cardiovascular disease, coronary heart disease, chronic heart failure, hypertension and hyperlipidemia (Su et al. 2015; Zhou et al. 2005). The medicinal properties of Danshen are mediated mainly by two groups of active components: the tanshinones, a group of lipophilic diterpenoid derivatives with an intense orange-brown color, e.g. tanshinone I (TI), tanshinone IIA (TIIA), cryptotanshinone (CT) and dihydrotanshinone (DHT), and the hydrophilic phenolic acids, including caffeic acid, ferulic acid, rosmarinic acid, salvianolic acid A-G, I-L and N (Wang et al. 2017).
The diterpene tanshinones are synthesized from two five-carbon precursors, isopentenyl diphosphate (IDP) and dimethylallyl diphosphate (DMADP), mainly by the plastidial methylerythritol-4-phosphate (MEP) pathway. The IDP and DMADP moiety is used for the stepwise biosynthesis of a 20-carbon geranylgeranyl diphosphate (GGPP), which then serves as a substrate for copalyl diphosphate synthase 1 and 2, producing (+)-copalyl diphosphate. The (+)-copalyl diphosphate is transformed into miltiradiene, the first committed tanshinone biosynthesis pathway intermediate, by the kaurene-like synthase 1 (Bai et al 2018; Guo and Ma 2019; Szymczyk et al. 2016).
The typical source of S. miltiorrhiza active ingredients is the root, which demonstrates the highest tanshinone concentration (Li et al. 2010). However, the most efficient alternative sources of S. miltiorrhiza plant material are hairy roots and cell cultures (Krajewska-Patan et al. 2007). Although the concentration of tanshinones in these sources is generally lower or at best comparable to roots of field plants, these cultures provide a continuous, season-independent supply of plant material (Wang and Wu 2010). The concentration of tanshinones can also be increased by application of chemical or physical components known as elicitors (Cheng et al. 2013; Hao et al. 2015). Although previous results suggest that salicylic acid positively influences tanshinone biosynthesis in hairy roots or cell cultures, little is known of the content in S. miltiorrhiza calluses growing on solid medium and exclusively elicited by SA; particularly for long culture durations (80 days), allowing for the accumulation of secondary metabolites (Wu et al. 2003). Previous studies related to callus culture elicitation used SA in relatively short tests that lasted for up to 18 days (Li et al. 2003; Chen and Chen 1999). A prompt metabolic response was observed after SA treatment (Hao et al., 2015) suggesting that the total tanshinone concentration in S. miltiorrhiza hairy root culture starts to increase 36 h after 0.1 mM SA treatment. Yu et al. (2016) report that 0.2 mM SA elicitation of suspension cell culture resulted in DHT and CT elevation after 3 days, TIIA augmentation after 4 days, and a reduction in TI concentration. However, Li et al. (2003) report that SA 0.2 mM did not increase the tanshinone concentration in Ti-transformed S. miltiorrhiza cell culture for up to 18 days.
Chen and Chen (1999) report that 50–500 μM SA applied alone is insufficient to induce CT biosynthesis in S. miltiorrhiza suspension cell cultures. SA concentrations higher than 0.2 mM were found to have no effect on CT biosynthesis rate, although it was tested for only a very short period of 5 days (Chen and Chen 1999). Therefore, little is known of the influence of exclusive SA elicitation on callus cultures for longer periods, i.e. up to 80 days, and relatively high concentrations, i.e. up to 0.4 mM. In addition, the influence of auxin phytohormone on callus cultures has not been studied. Only few studies have examined the effect of auxin phytohormones, such as indole-3-acetic acid (IAA) and NAA, on tanshinone concentration in S. miltiorrhiza; however, existing data suggests they do not stimulate tanshinone biosynthesis in hairy root cultures (Gupta et al. 2011). No such study has been performed on callus cultures of S. miltiorrhiza.
Callus culture has significant commercial potential, particularly with regard to the production of plant secondary metabolites with medicinal activity, as well as therapeutic antibodies and other recombinant proteins, in addition to preparing ornamental or agricultural plants by regeneration from calli. Active ingredients may be purified directly from callus culture to relieve the pressure on plants in their natural habitats (Park and Wi 2016; Top et al. 2019; Efferth 2019). Also, callus cultures may be easily converted into single cell cultures growing as suspension in bioreactors. Such culture may be subjected to scale-up and process optimization, similarly to large-scale animal cell cultures (Georgiev et al., 2009, 2018). Some commercially-available drugs are produced by large-scale callus cultures, such as Paclitaxel from yew (Taxus. sp.) or taliglucerase alpha (Elelyso) from carrot Daucus carota (Tabata 2004; Grabowski et al. 2014).
SA is a phytohormone produced by plants during their response to abiotic stress, including drought, heavy metal, heat, osmotic and chilling stress. However, SA seems to play a central role in the response to biotropic pathogens, which colonize between plant cells to absorb host nutrients (Li et al. 2019). SA-responsive TGACG cis-active elements have been found in numerous plant promoters (Fonseca et al. 2010; Sun et al. 2018), and SA is known to induce the expression of the SmWRKY1 trans-factor, which strongly induce genes associated with tanshinone biosynthesis through interaction with the W-box TTGAC(C/T) cis-active element (Cao et al. 2018).
Contrary to SA, the influence of auxin on gene expression is not clearly activatory. Among 25 Auxin Response Factors (ARFs) identified in S. miltiorrhiza only seven (SmARF 2, 10, 19, 20, 22, 24, 25) act as transcription activators while the other 18 SmARFs may function as transcription repressors (Xu et al. 2016). Moreover, the binding affinity of the ARFs to DNA and strength of transcription interference depends not only on the presence of cis-active elements recognized by ARFs but also on the spatial organization of these elements allowing ARFs dimerization (Berendzen et al. 2012). Therefore, efficient transcription activation or inhibition of target gene expression by ARFs requires the presence of TGTCGG inverted repeats and TGTCTC or TGTCGG direct repeats (Freire-Rios et al. 2020). Also, the presence of consensus sequence TGTSTSBC (B=C or G or T; S=C or G) in the promoter region is positively associated with auxin-dependent gene expression regulation (Mironova et al. 2014).
The present paper compares the effects of elicitation by the synthetic auxin NAA and SA on the concentrations of four tanshinones CT, DHT, TI and TIIA, as well as total tanshinone content, in S. miltiorrhiza callus cultures grown for up to 80 days on solid MS medium. To explain the metabolic effects indicated by both phytohormones on S. miltiorrhiza callus, the study examined the distribution frequency of SA and auxin-responsive cis-active elements in promoters of selected genes associated with tanshinone precursor biosynthesis.
Materials and methods
Plant material
Salvia miltiorrhiza plants were cultivated from seeds provided by the Medicinal Garden of the Department of Pharmacognosy of the Medical University of Lodz (Poland). The plants were grown in pots of 0.5 L (12 cm in diameter) containing composite soil, at 22 ± 2 °C under natural light. Young plants at the age of 3 months were used for experiments.
Callus induction, salicylic acid and NAA treatment
The induction and proliferation of S. miltiorrhiza callus was performed according to Wu et al. (2003) with modifications: callus induction was performed with 10 × 10 mm leaf pieces from the S. miltiorrhiza plants, compared to 5 × 5 mm described in the original method, and 100 mm glass Petri dishes were used instead of the original 22 × 160 mm glass tubes. The explant surface was disinfected with 70% ethanol for 30 s, followed by treatment with 0.5% sodium hypochlorite with Tween 20 (two drops of Tween 20 per 100 mL) for 10 min; following this, the surface was rinsed five times with sterile distilled water. The explants were placed in 100 mm diameter glass Petri dishes containing 20 mL of solid medium to induce growth of primary callus. The medium consisted of Murashige and Skoog (1962) basal medium known as MS, supplemented with 3% sucrose, 1% Difco Bacto agar (Difco Laboratories, Detroit, Michigan, US) and 1 mg L−1 2,4-dichlorophenoxyacetic acid (2,4-D). The Petri dishes with explants were sealed with two layers of Parafilm (Pechiney Plastic Packaging, Chicago, USA). All procedures were performed under a laminar hood to avoid microbiological contamination. The cultures were incubated at 26 ± 2 °C in darkness for a period of 1 month. To avoid light, the glass Petri dishes were covered with two layers of aluminum foil. The induced primary callus was allowed to proliferate on a solid MS basal medium supplemented with 3% sucrose, 1% Difco Bacto agar (Difco Laboratories, Detroit, Michigan, US), 1 mg L−1 2,4-D and 0.5 mg L−1 N6-benzyladenine (BA). The proliferating callus was cultured three times at 20-day intervals in darkness in 100 mm diameter glass Petri dishes containing 20 mL of solid medium. Then, the proliferated callus was used for experiments with SA and NAA elicitation.
The solid medium consisted of MS basal medium supplemented with 3% sucrose, 1% Difco Bacto agar (Difco Laboratories, Detroit, Michigan, US) and 0.1, 0.2 and 0.4 mM SA sodium salt or 2.69, 13.43, 26.85 and 40.28 μM NAA. The callus cultures were maintained for up to 80 days. Fresh solid medium was provided every 20 days. Callus samples for HPLC experiments were harvested each 10 days, up to 80 days; however samples for growth index calculation were collected after 20, 40, 60 and 80 days. Callus samples harvested after 10 days were grown on the same medium they started growth. Callus samples grown longer than 20 days were transferred to new, fresh medium. For example, callus expected to grow for 30 days was nourished by the initial medium for the first 20 days and a fresh, exchanged portion for the next 10 days.
All the experiments were repeated three times, and growth measurements were performed with three replicates per harvesting.
Before the phytohormones were added, the pH of each medium was adjusted to 5.7 ± 0.1 with 1 N NaOH or HCl, before autoclaving at 121 °C, 105 kPa, 20 min. Stock solutions of phytohormones 2,4-D (0.4 mg/mL), NAA (0.2 mg/mL) and SA sodium salt (100 mM) in 70% ethanol were sterilized with a syringe filter (0.4 μm pore size). They were then added to the warm (50 °C) freshly-autoclaved MS medium under a laminar hood. The control group was the NAA and SA untreated callus.
The callus mass was evaluated on electronic scales localized under a laminar hood to avoid microbiological contamination of the callus. The electronic scales and working surface of the laminar hood were initially sterilized with UV irradiation (30 min) and then with 70% ethanol. The glass material and steel forceps used to manipulate the calluses were sterilized at 200 °C, 30 min before use in the laminar hood. Plastic tips were autoclaved as liquid media.
Growth index (GI) was calculated according to the formula described by Godoy-Hernández and Vázquez-Flota (2006). The GI of fresh calls weight is defined as follows: (GIF) = final callus fresh weight (FW) − initial callus fresh weight (FW)/initial callus fresh weight (FW). The same definition was used to calculate the growth index of dry callus—GID.
Preparation of callus extract
The extraction of callus was performed according to Wan et al. (2009). Methanol or 80% methanol solution is now commonly used to extract tanshinones from S. miltiorrhiza plant material (Wei et al. 2019; Zhao et al. 2015; Liu et al. 2017). By using methanol, it is possible to avoids low pressure evaporation of additional solvents, such as chloroform before starting the HPLC procedure (Liu et al. 2006). Removing the evaporation stage is particularly important for experiments based on a small amount of dried callus measured in tens of mg (Zhao et al. 2015). Such very small dried samples produce a tiny amount of extracted tanshinones in the glass vessel that may be not completely solubilized in the added methanol, resulting in relatively high error rates. The callus was harvested carefully and freeze-dried for a 3 days in a lyophylizer Alpha 1-2 LD (Martin Christ, Osterlode, Germany) connected to a GVD1.5 vacuum pump (Atlas Copco, Stockholm, Sweden). The pressure was maintained at 0.1 mbar by a vacuum pump.
The 30 mg dry callus was finely ground with a mortar and a pestle and extracted with 1.5 mL 100% methanol under 90 min sonication (UM1 disintegrator, Unimal, Olsztyn, Poland) at room temperature. The liquid/solid ratio was 50:1 as in the original method (Wan et al. 2009). The samples were then centrifuged at 12,000× rpm for 10 min at room temperature, filtered through 0.45 μm Chromafil membrane (Machery-Nagel, Duren, Germany) and used for HPLC analysis. The samples was stored in darkness at – 25 °C to avoid tanshinone decomposition.
HPLC analysis
Standard HPLC-grade substances such as TI, TIIA, CT and DHT were provided by Sigma Aldrich Poland (Poznan, Poland). Methanol, acetonitrile and water (J.T. Baker HPLC Analyzed) were received from Avantor Performance Materials (Gliwice, Poland). All analyses were performed on the Agilent HPLC System 1200 (Agilent, Palo Alto, US), equipped with a quaternary-pump delivery system, an on-line degasser, an auto sampler, a column temperature controller and a UV–VIS DAD detector. The system was connected with the Agilent ChemStation 2001–2010 software.
Chromatographic separation was performed on an Agilent Zorbax Extend C18 reversed phase column (5 μm, 250 × 4.6 mm) with an Agilent Zorbax Extend C18 guard column (5 μm, 10 × 4.6 mm). The detection was performed at 270 nm, the time of analysis was 20 min, the flow rate was 1.2 mL min−1 and the column temperature was maintained at 20 °C; the sample volume was 20 μL. The mobile phase consisted of A (water for HPLC) and B (acetonitrile). The following gradient program was applied: initially 45% B at 0 min, linearly increasing to 60% B at 2 min, maintaining 60% B from 2 to 9 min, linearly increasing B to 80% at 10 min, linearly increasing B to 82% at 13 min and finally linearly decreasing B to 45% at 20 min. After each analysis, 45% B was pumped and held for 10 min to re-equilibrate the system for baseline stability. The procedure is a modified method of Liu et al. (2006).
Preparation of calibration curves and standard solutions
The starting methanol stock solutions of 0.1 mg mL−1 TI, TIIA, CT and DHT were prepared and then stored in dark, Parafilm-sealed HPLC vials at – 25 °C: storage at 5 °C may cause decomposition. To prepare standard curves, the initial standard solution was appropriately diluted. The retention time for DHT was 7.1 min. The calibration curve is described by the following equation y = 2.9630x + 1.1634, the correlation coefficient is 0.9997, LOD 0.9624 and LOQ 2.9165. The retention time for CT was 10.7 min. The calibration curve for CT is y = 4.3133x + 1.5136, the correlation coefficient 0.9995, the LOD 1.3153 and LOQ 3.986. The retention time for TI was 11.5 min, the calibration curve y = 2.4617x + 2.2342, the correlation coefficient 0.9995, LOD 1.3325 and LOQ 4.077. For TIIA the retention time was 13.7 min, the standard curve y = 5.3145x + 1.3120, the correlation coefficient 0.9997, LOD 1.0337 and LOQ 3.1325. The linearity range of all calibration curves appeared to be 1.25–50 ng.
Promoter analysis
The findings regarding the A. thaliana and S. miltiorrhiza genes used in the study are presented below. They were checked against the PlantPAN3.0, Arabidopsis org-TAIR, NCBI (Nucleotide) and Uniprot databases (Chow et al. 2019; Rhee et al. 2003; NCBI Resource Coordinators 2016; The UniProt Consortium 2021; Majewska et al. 2018). The TSSP software was used to characterize transcription start sites in S. miltiorrhiza promoters (Shahmuradov et al. 2005). The precise description of genes used in the study is presented in Supplement text file 1.
Although the promoter region in A. thaliana is approximately 0.5–1.0 kb long, the in silico searches were concentrated on proximal promoters within 300 bp of the transcription start site (Kristiansson et al. 2009). The decision to decrease the length of studied promoter is based on the higher biological relevance of cis-active motifs found within proximal promoters (Mironova et al. 2014; Keilwagen et al. 2011; Yu et al. 2016).
The gene proximal promoters were searched for the following cis-active elements: SA-responsive TGACG and W-boxes TTGAC(C/T), auxin-responsive TGTCGG, TGTCTC and TGTSTSBC (B=C or G or T, S=C or G) (Fonseca et al. 2010; Sun et al. 2018; Cao et al. 2018; Freire-Rios et al. 2020; Mironova et al. 2014).
Statistical analysis
The results of the HPLC analysis were evaluated by the Kruskal–Wallis test with STATISTICA (StatSoft Inc. 2013, version 13.1). Data concerning wet or dry callus samples mass were compared using the Student’s t-test. Values of p < 0.05 were considered statistically significant.
Results
Changes in tanshinone concentration after SA treatment
Generally, SA elicitation showed time- and concentration-dependent effects. The four tanshinones (TI, TIIA, CT and DHT) were not induced simultaneously, and their elicitation occurred stepwise.
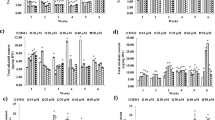
After 50 days of SA elicitation, all tanshinones demonstrated induction for all three SA concentrations, reaching the highest values for 0.4 mM SA DHT (71.58 ± 12.72 μg g−1 DW), CT (108.54 ± 18.29 μg g−1 DW), TI (29.50 ± 4.13 μg g−1 DW) and TIIA (16.75 ± 2.74 μg g−1 DW) (Fig. 1). Lower values were observed than 0.1 mM SA elicitation after 50 days: DHT (19.96 ± 1.16 μg g−1 DW), CT (29.50 ± 1.97 μg g−1 DW), TI (13.00 ± 0.97 μg g−1 DW) and TIIA (5.75 ± 1.24 μg g−1 DW). Also, 0.2 mM SA elicitation demonstrated lower levels for 50 days of treatment: DHT (28.17 ± 2.06 μg g−1 DW), CT (44.33 ± 2.96 μg g−1 DW), TI (15.21 ± 5.68 μg g−1 DW) and TIIA (9.63 ± 0.72 μg g−1 DW). After this time, the elicitation effects of all three SA concentrations were decreasing, and the concentrations of all tanshinones decreased stepwise (Fig. 1).
Concentration of four tanshinones cryptotanshinone (CT), dihydrotanshinone (DHT), tanshinone I (TI) and tanshinone IIA (TIIA) in S. miltiorrhiza callus presented as a function of salicylic acid (SA) concentration (0.1, 0.2 and 0.4 mM) and elicitation time (10–80 days). Control values were not presented as tanshinone concentration was not detectable
Comparing the 40-day and 50-day elicitation by 0.4 mM SA, the concentrations of all four tanshinones could be organized in the following manner CT > DHT > TI > THIIA (Fig. 1).
The presented changes are associated with fluctuations of total tanshinone concentration as compared to control samples (Fig. 2). The maximum total tanshinone concentration observed after day 50 (Fig. 2). All SA concentrations reached maximum elicitation after this period, viz. 0.1 mM (68.21 ± 4.80 μg g−1 DW), 0.2 mM (97.33 ± 5.67 μg g−1 DW) and 0.4 mM SA (226.38 ± 37.30 μg g−1 DW) (Fig. 2). Longer SA elicitation resulted in a significant decrease in the total tanshinone concentration.
Tanshinones have a well-known and strong orange-red coloration (Wu et al. 2003). Appropriate pigmentation changes in callus colors are induced by SA treatment. These changes are well visible after 50 days of elicitation. Danshen calluses treated with 0.1 or 0.2 mM SA demonstrated a yellow to pale-orange pigmentation as compared to control samples (Fig S1D). In addition, S. miltiorrhiza calluses treated with 0.4 mM SA appeared to be darker, which may be related to higher total tanshinone concentration.
Changes of callus growth index induced by SA treatment
The growth index (GI) of the final biomass is calculated by dividing harvested fresh weight or dry weight by the inoculum fresh weight or dry weight. The GI for both fresh (GIF) and dry (GID) callus weight was analysed for 80 days in a 20-day intervals. The GIF for control callus cultures was 3.47–71.15 for the 20 to 80-day interval (Fig. 4), with the GIF for callus cultures being 2.00–27.03 (0.1 mM SA) and 2.12–27.23 (0.2 mM SA) and only 1.59–5.12 at 0.4 mM SA (Fig. 4). The GID values were generally similar to GIF, these being 3.77–80.79 for controls for the 20 to 80-day interval, but 2.42–24.63 (0.1 mM), 2.19–26.42 (0.2 mM SA) and 1.64–5.58 (0.4 mM SA) (Fig. 3).
Changes of tanshinone productivity mediated by SA elicitation
As the highest total tanshinone concentration (226.38 ± 37.30 μg g−1 DW) was observed after day 50 of the 0.4 mM SA elicitation it could be expected that this system should be the best source of tanshinones. In addition, the total tanshinone level observed for 0.2 mM SA, was 2.33 times lower than for 0.4 mM SA.
Treatment with 0.2 mM SA yielded 4.65 ± 0.78 GIF for day 40 and 11.21 ± 0.93 GIF for day 60, encompassing the time point (50 days) of the highest total tanshinone concentration. These values were approximately 2.03–2.99 times higher for cultures induced with 0.4 mM SA (GIF 2.29 ± 0.20 at day 40 and 3.75 ± 0.52 for day 60). Therefore, on day 50, the lower concentration of total tanshinones observed for 0.2 mM SA than 0.4 mM SA may be compensated by the higher GIF of the callus growing on 0.2 mM SA, i.e. approximately 2.03–2.99 times higher.
Distribution of SA responsive cis-active elements within proximal promoters of A. thaliana and S. miltiorrhiza
The SA-responsive cis-elements were found only in one MEP pathway gene, 4-(cytidine 5′-diphospho)-2-C-methyl-d-erythritol kinase (CMK), and one MVA route gene: diphosphomevalonate decarboxylase 1 (PMD1) (Tables S1–2). Furthermore, eight of the 18 tested A. thaliana genes encoding enzymes participating in GGPP biosynthesis indicate an SA-responsive cis-active motif (Table S3). Therefore, the A. thaliana and S. miltiorrhiza genes associated with isoprenoid biosynthesis demonstrate much better representation of SA-responsive motifs than auxin-responsive cis-elements. Among 35 tested A. thaliana genes, as much as ten (28.57%) indicated SA-responsive elements (Fig. 4).
Enzymatic reactions participating in tanshinone biosynthesis (Guo and Ma 2019)
In addition, three among seven (42.86%) S. miltiorrhiza genes present these cis-active motifs, which may explain the increased tanshinone concentration in response to SA treatment observed in the present study (Table S4) (Fig. 4). Among these three, the SmCPS and SmHMGR2 genes were verified experimentally by RT-PCR studies in transgenic plants (Szymczyk et al. 2016, 2018). Similarly, the S. miltiorrhiza genes SmIDI, SmGGPPS, SmHMGS, SmHDR and SmDXR have also been experimentally-confirmed to respond to SA treatment (Hua et al. 2012; Hao et al. 2013, 2015; Jiang et al. 2019; Yan et al. 2009; Zhang et al. 2010, 2016). SmDXS2 gene responded positively to overexpression of SmWRKY1, that is stimulated by SA treatment (Cao et al. 2018).
In addition, closely-spaced SA-responsive elements were observed within two A. thaliana genes: isopentenyl-diphosphate Delta-isomerase and geranylgeranyl diphosphate synthase 9 (Table S5). Such closely-spaced repetitions of cis-active motifs allow the formation of hetero- or homodimers of trans-factors to precisely regulate gene expression.
Callus response to NAA treatment
The elicitation of S. miltiorrhiza callus cultures by 2.69, 13.43, 26.85 and 40.28 μM NAA did not increase the concentration of four tanshinones TI, TIIA, CT and DHT over the 80 days. The values of TI, TIIA, CT and DHT were so low in the treated samples and the untreated controls that they could not be detected by HPLC.
Changes of callus growth index after NAA treatment
The GIF for control callus cultures was 3.42–70.63 for the 20–80 days interval (Fig. 5). Similar GIF values were obtained for callus cultures growing on 2.69 μM NAA (3.19–67.35), 13.43 μM NAA (3.66–75.34) and 26.85 μM NAA (3.62–83.03). The 40.28 μM NAA treatment inhibited callus growth rate (GIF 2.32–21.69). The values of GID for control and tested samples were similar to GIF (Fig. 5). The color of callus growing on medium containing NAA within the range 2.69–40.28 μM was related to control samples (Fig S1D).
Distribution of auxin responsive cis-active elements within proximal promoters of A. thaliana and S. mitiorrhiza
Analysis of A. thaliana MVA pathway genes showed no auxin-responsive elements in the proximal promoter region localized up to 300 bp upstream from TSS (Table S2). The auxin-responsive elements were found in two genes of the A. thaliana MEP pathway: 2-C-methyl-d-erythritol 4-phosphate cytidylyltransferase and 4-hydroxy-3-methylbut-2-enyl diphosphate reductase (Table S1). Moreover only one among 18 studied A. thaliana genes participating in GGPP biosynthesis, known as geranylgeranyl diphosphate synthase 12 (GGPPS12) displays an auxin-responsive cis-active motif in the proximal promoter (Table S3). The lack of these sequences in the seven tested S. mitiorrhiza genes may be related to the observed scarcity of metabolic response to auxin treatment (Table S4). Previous studies suggest that IAA 2.85 μM (0.5 mg L−1) treatment does not induce of SmHMGR2 gene expression (Szymczyk et al. 2018); this is consistent with the lack of tested auxin-responsive cis-active motifs in SmHMGR2 or its A. thaliana homolog (Table S4) (Fig. 4).
No closely-spaced, tandem or inverted repeats of auxin-responsive elements in proximal promoters were observed, suggesting that the phytohormone has low functional importance in regulating the tested genes (Freire-Rios et al. 2020).
Discussion
Assuming that some plant biosynthetic routes are tissue or organ specific, like the tanshinone or ginsenoside biosynthesis pathway observed in the rhizome, callus cultures offer generally less efficient product biosynthesis (Hao et al. 2015; Szymanska et al. 2013). Genes encoding enzymes of critical importance for late stages of tanshinone biosynthesis, such as SmCPS1, SmKSL1 and SmCYP76AH1 are predominantly expressed in taproot of S. miltiorrhiza (Hao et al. 2020). Therefore the callus cultures could be expected to indicate a lower biosynthesis rate than roots or S. miltiorrhiza plants.
Indeed, previous studies on S. miltiorrhiza calluses show a generaly low total tanshinone concentration of 0.077% (Gryszczynska et al. 2015). Similar values 0.06–0.08% were observed in Ti-transformed suspension cultures of S. miltiorrhiza (Chen et al. 1997). This is considerably lower than in roots regenerated in vitro (0.269–1.137%) (Li et al. 2010) or in native plant roots growing in China (0.260–0.388%) (Cheng et al. 2013). The cultivar dependency is a significant factor, as S. miltiorrhiza plants in Poland are characterized by a lower tanshinone concentration of 0.01–0.26% (Buchwald et al. 2006).
Our results confirm SA-dependent tanshinone elicitation in callus cultures growing on solid medium. Low total tanshinone concentrations of 0.023, 0.010 and 0.007% was observed after 50-day elicitation of 0.4, 0.2 and 0.1 mM SA, respectively, which was consistent with previous observations. Therefore, only the highest SA concentration of 0.4 mM was able to increase the total tanshinone concentration within the lower range observed in native plant roots growing in Poland. However, such high SA concentrations inhibit the callus growth rate and is twice higher than 0.2 mM commonly applied in plant elicitation experiments.
Evaluation of tanshinone types induced by SA indicates that the S. miltiorrhiza callus growing on solid MS medium is dominated by CT with a lower concentration of DHT observed. Similar results suggesting predominant concentration of DHT and CT were provided by other authors in callus or hairy root cultures (Hao et al., 2015; Cheng et al. 2013). Also, Krajewska-Patan et al. (2007) showed that DHT had an advantage (0.27%) over the CT (0.12%) in Danshen callus cultures. Zhao et al. (2010) found that CT was significantly stronger when stimulated by several elicitors as Co2+, Ag+, SA, yeast elicitor, sorbitol and chitosan in S. miltiorrhiza cell cultures. Only results of Cao et al. (2018) show the dominant concentration of TI over CT and DHT in transformed S. miltiorrhiza hairy roots overexpressing SmWRKY1 transcription factor. Similarly, the predominant tanshinone in native S. miltiorrhiza plant roots is TIIA.
These differences may be explained by the tanshinone biosynthesis pathway structure, where the CT is further modified to TIIA, whereas DHT serves as a substrate to produce TI (Guo and Ma 2019). Alternatively, CT plays a role in phytoalexin function, concentrated in cells and directly involved in response to yeast extract elicitation (Chen and Chen 2000).
The time course of tanshinone accumulation in our solid callus culture SA-induced system was slower and not as stable as in previous studies using cytokinin elicitation (Wu et al. 2003). Also S. miltiorrhiza cell suspension or hairy root cultures show a relatively short (24-h) response time to SA treatment (Cheng et al. 2013). In addition, Hao et al. (2015) indicate that the total tanshinone concentration in S. miltiorrhiza hairy root culture starts to increase 36 h after 0.1 mM SA treatment. Our findings show generally lower tanshinone accumulation dynamics; this may be related to the fact they were grown on solid rather than liquid medium, or the generally lower initial tanshinone productivity demonstrated by plants growing in Poland (Buchwald et al. 2006; Cheng et al. 2013).
Although 0.4 mM SA induced 2.33-times higher total tanshinone concentration than 0.2 mM SA after 50 days of induction, the decreased tanshinone concentration is compensated by GIF, which was 2.03–2.99 times higher for a lower SA value. Similar values of GI were observed for dry weight (GID). The GI noted for callus cultures on solid medium was generally lower than GI for callus cultures on liquid medium. Li et al. (2003) observed GI values of 11.44 in controls and 9.93 for samples treated with 0.2 mM SA for 18 days. Our data show much slower GI of 3.47 ± 0.15 for the control group and 2.12 ± 0.27 for 0.2 mM SA after 20 days of culture on solid medium.
Elicitation of callus by auxin (NAA) did not show increased tanshinone concentration nor any significant change in GIF or GID for 2.69, 13.43, 26.85 and 40.28 μM NAA. Only the highest NAA concentration 40.28 μM (7.5 mg L−1) NAA slowed the callus growth rate. It is possible that SA and NAA elicitation may influence the tanshinone biosynthesis rate though inhibiting the transcription activity of most ARFs in S. miltiorrhiza (Xu et al. 2016). Also, the expression of certain genes, such as SmHMGR2, in S. miltiorrhiza is not induced by auxin IAA (Szymczyk et al. 2018).
In contrast, SA induces the expression of genes associated with tanshinone biosynthesis (Hua et al. 2012; Hao et al. 2013, 2015; Yan et al. 2009; Zhang et al. 2010, 2016). Most of these genes or their A. thaliana homologs indicate SA-responsive cis-elements in proximal promoters. In contrast, compared to SA-responsive elements, the auxin-responsive cis-active elements are much rarer in the tested proximal promoters of A. thaliana and S. miltiorrhiza genes participating in isoprenoid biosynthesis.
Conclusion
The presented study compares the influence of SA and NAA elicitation on tanshinone concentration in S. miltiorrhiza callus cultures growing for up to 80 days on a solid MS medium. The SA indicated clear but relatively low induction of four tanshinones, viz. CT, DHT, TI and TIIA, as well as total concentration, which may be due to the relatively low tanshinone concentration is S. miltiorrhiza plants growing in Poland. However, NAA elicitation did not increase tanshinone concentration. The influence of either phytohormone on tanshinone concentration seems to be proportional to the distribution frequency of auxin and SA-responsible cis-active elements in proximal promoters of S. miltiorrhiza or A. thaliana homologous genes, encoding for the enzymes of the MEP and MVA pathways or the later stages of tanshinone precursor (miltiradiene) biosynthesis.
Abbreviations
- NAA:
-
1-Naphthaleneacetic acid
- IAA:
-
Indole-3-acetic acid
- BA:
-
N6-benzyladenine
- SA:
-
Salicylic acid
- 2,4-D:
-
2,4-Dichlorophenoxyacetic acid
- MEP:
-
Methylerythritol-4-phosphate
- MVA:
-
Mevalonic acid
References
Bai Z, Li W, Jia Y, Yue Z, Jiao J, Huang W, Xia P, Liang Z (2018) The ethylene response factor SmERF6 co-regulates the transcription of SmCPS1 and SmKSL1 and is involved in tanshinone biosynthesis in Salvia miltiorrhizza hairy roots. Planta 248:243–255
Berendzen KW, Weiste Ch, Wanke D, Kilian J, Harter K, Dröge-Laser W (2012) Bioinformatic cis-analyses performed in Arabidopsis and rice disclose bZIP and MYB-related binding sites as potential AuxRE-coupling elements in auxin-mediated transcription. BMC Plant Biol 12:125. https://doi.org/10.1186/1471-2229-12-125
Buchwald W, Forycka A, Mścisz A, Mielcarek S, Mrozikiewicz PM, Baraniak M (2006) Zmiany zawartości substancji biologicznie czynnych w korzeniach Salvia miltiorrhza Bunge. Herba Pol 52:64
Cao W, Wang Y, Shi M, Hao X, Zhao W, Wang Y, Ren J, Kai G (2018) Transcription factor SmWRKY1 positively promotes the biosynthesis of tanshinones in Salvia miltiorrhiza. Front Plant Sci 9:554. https://doi.org/10.3389/fpls.2018.00554.eCollection2018
Chen H, Chen F (1999) Effect of methyl jasmonate and salicylic acid on cell growth and cryptotanshinone formation in Ti transformed Salvia miltiorrhiza suspension cell cultures. Biotechnol Lett 21:803–807
Chen H, Chen F (2000) Induction of phytoalexin formation in crown gall and hairy root culture of Salvia miltiorrhiza by methyl viologen. Biotechnol Lett 22:715–720
Chen H, Yuan JP, Chen F, Zhang YL, Song JY (1997) Tanshinone production in Ti transformed Salvia miltiorrhiza cell suspension cultures. J Biotechnol 58:147–156
Cheng Q, He Y, Li G, Liu Y, Gao W, Huang L (2013) Effects of combined elicitors on tanshinone metabolic profiling and SmCPS expression in Salvia miltiorrhiza hairy root cultures. Molecules 18:7473–7485. https://doi.org/10.3390/molecules18077473
Chow ChN, Lee TY, Hung YCh, Li GZ, Tseng KCh, Liu YH, Kuo PL, Zheng HQ, Chang WCh (2019) PlantPAN3.0: a new and updated resource for reconstructing regulatory networks from ChIP-seq experiments in plants. Nucleic Acids Res 47(D1):D1155–D1163
Efferth TH (2019) Biotechnology applications of plant callus cultures. Engineering 5:50–59
Fonseca JP, Menossi M, Thibaud-Nissen F, Town CD (2010) Functional analysis of a TGA factor-binding site located in the promoter region controlling salicylic-acid induced NIMIN-1 expression in Arabidopsis. Gen Mol Res 9:167–175
Freire-Rios A, Tanaka K, Crespo I, van der Wijk E, Sizentsova Y, Levitsky V, Lindhoud S, Fontana M, Hohlbein J, Boer DR, Mironova V, Weijers D (2020) Architecture of DNA elements mediating ARF transcription factor binding and auxin-responsive gene expression in Arabidopsis. Proc Natl Acad Sci USA 117:24557–24566
Georgiev MI, Weber J, Maciuk A (2009) Bioprocessing of plant cell cultures for mass production of targeted compounds. Appl Microbiol Biotechnol 83:809–823
Georgiev V, Slavov A, Vasileva I, Pavlov A (2018) Plant cell culture as emerging technology for production of active cosmetic ingredients. Eng Life Sci 18:779–798
Godoy-Hernández G, Vázquez-Flota FA (2006) Growth measurements: estimation of cell division and cell expansion. Methods Mol Biol 318:51–58
Grabowski GA, Golembo M, Shaaltiel Y (2014) Taliglucerase alfa: an enzyme replacement therapy using plant cell expression technology. Mol Genet Metab 112:1–8
Gryszczynska A, Opala B, Lowicki Z, Dreger M, Gorska-Paukszta M, Szulc M, Kaminska E, Litwin E, Struzik P, Dyr W, Wyszogrodzka E, Mikolajczak PL (2015) Bioactive compounds determination in the callus and hydroalcoholic extracts from Salvia miltiorrhiza and Salvia przewalskii—preliminary study on their anti-alcoholic activity effects. Phytochem Lett 11:399–403
Guo J, Ma Y (2019) Biosynthetic pathway of tanshinones in Salvia miltiorhiza. In: Lu S (ed) The Salvia miltiorrhiza genome, 1st edn. Springer, Cham, Switzerland, pp 129–140
Gupta SK, Liu RB, Liaw SY, Chan HS, Tsay HS (2011) Enhanced tanshinone production in hairy roots of ‘Salvia miltiorrhiza Bunge’ under the influence of plant growth regulators in liquid culture. Bot Stud 52:435–443
Hao G, Shi R, Tao R, Fang Q, Jiang X, Ji H, Feng L, Huang L (2013) Cloning, molecular characterization and functional analysis of 1-hydroxy-2-methyl-2-(E)-butenyl-4-diphosphate reductase (HDR) gene for diterpenoid tanshinone biosynthesis in Salvia miltiorrhiza Bge. F. alba. Plant Physiol Biochem 70:21–32
Hao X, Shi M, Cui L, Xu Ch, Zhang Y, Kai G (2015) Effects of methyl jasmonate and salicylic acid on tanshinone production and biosynthetic gene expression in transgenic Salvia miltiorrhiza hairy roots. Biotechnol Appl Biochem 62:24–31
Hao X, Pu Z, Cao G, You D, Zhou Y, Deng Ch, Shi M, Nile SH, Wang Y, Zhou W, Kai G (2020) Tanshinone and salvianolic acid biosynthesis are regulated by SmMYB98 in Salvia miltiorrhiza hairy roots. J Adv Res 23:1–12
Hua W, Song J, Li C, Wang Z (2012) Molecular cloning and characterization of the promoter of SmGGPPs and its expression pattern of Salvia miltiorrhiza. Mol Biol Rep 39:5775–5783
Jiang Y, Wang L, Lu S, Xue Y, Wei X, Lu J, Zhang Y (2019) Transcriptome sequencing of Salvia miltiorrhiza after infection by its endophytic fungi and identification of genes related to tanshinone biosynthesis. Pharm Biol 57:760–769
Keilwagen J, Grau J, Paponov IA, Posch S, Strickert M, Grosse I (2011) De-novo discovery of differentially abundant transcription factor binding sites including their positional preference. PLoS Comput Biol 7:e1001070. https://doi.org/10.1371/journal.pcbi.1001070
Krajewska-Patan A, Dreger M, Górska-Paukszta M, Mścisz A, Mielcarek S, Baraniak M, Buchwald W, Marecik R, Grajek W, Mrozikiewicz PM (2007) Salvia miltiorrhiza Bunge in vitro cultivation in callus cultures. Herba Pol 53:88–96
Kristiansson E, Thorsen M, Tamás MJ, Nerman O (2009) Evolutionary forces acts on promoter length: identification of enriched cis-regulatory elements. Mol Biol Evol 26:1299–1307
Li GJ, Wang SC, Xia K, Zhou X (2003) Effect of yeast elicitor and salicylic acid on the fluctuation of phytohormone contents in Ti-transformed Salvia miltiorrhiza cell cultures. Plant Growth Regul 39:27–32
Li MH, Peng Y, Xiao PG (2010) Distribution of tanshinones in the genus Salvia (family Lamiaceae) from China and its systematic significance. J Syst Evol 48:118–122
Li N, Han X, Feng D, Yuan D, Huang LJ (2019) Signaling crosstalk between salicylic acid and ethylene/jasmonate in plant defense: do we understand what they are whispering? Int J Mol Sci 20:671. https://doi.org/10.3390/ijms20030671
Liu AH, Lin YH, Yang M, Sun JH, Guo H, Guo DA (2006) High-performance liquid chromatographic determination of tanshinones in the roots of Salvia miltiorrhiza and related traditional Chinese medical preparations. J Pharm Pharm Sci 9:1–9
Liu Y, Sun G, Zhong Z, Ji L, Zhang Y, Zhou J, Zheng X, Deng K (2017) Overexpression of AtEDT1 promotes root elongation and affects medicinal secondary metabolite biosynthesis in roots of transgenic Salvia miltiorrhiza. Protoplasma 254:1617–1625
Majewska M, Wysokińska H, Kuźma Ł, Szymczyk P (2018) Eukaryotic and prokaryotic promoter databases as valuable tools in exploring the regulation of gene transcription: a comprehensive overview. Gene 644:38–48
Mironova VV, Omelyanchuk NA, Wiebe DS, Levitsky VG (2014) Computational analysis of auxin responsive elements in the Arabidopsis thaliana L. genome. BMC Genomics 15:S4
Murashige T, Skoog F (1962) A revised medium for rapid growth and bioassays for tobacco tissue cultures. Physiol Plant 15:473–497
NCBI Resource Coordinators (2016) Database resources of the national center of biotechnology information. Nucleic Acids Res 44(Database issue):D7–D19
Park KY, Wi SJ (2016) Potential of plants to produce recombinant protein products. J Plant Biol 59:559–568
Rhee SY, Beavis W, Berardini TZ, Chen G, Dixon D, Doyle A, Garcia-Hernandez M, Huala E, Lander G, Montoya M, Miller N, Mueller LA, Mundodi S, Reiser L, Tacklind J, Weems DC, Wu Y, Xu I, Yoo D, Yoon J, Zhang P (2003) The Arabidopsis information resource (TAIR): a model organism database providing a centralized, curated, gateway to Arabidopsis biology, research materials and community. Nucleic Acids Res 31:224–228
Shahmuradov IA, Solovyev VV, Gammerman AJ (2005) Plant promoter prediction with confidence estimation. Nucleic Acids Res 33:1069–1076
Su CY, Ming QL, Rahman K, Han T, Qin LP (2015) Salvia miltiorrhiza: traditional medicinal uses, chemistry and pharmacology. Chin J Nat Med 13:163–182
Sun T, Busta L, Zhang Q, Ding P, Jetter R, Zhang Y (2018) TGACG-binding factor 1 (TGA1) and TGA4 regulate salicylic acid and pipecolic acid biosynythesis by modulating the expression SYSTEMIC ACQUIRED RESISTANCE DEFICIENCT 1 (SARD1) and CALMODULIN-BINDING PROTEIN 60g (CBP60g). New Phytol 217:344–354
Szymanska G, Kochan E, Szymczyk P (2013) Field cultivation and in vitro cultures, root-forming callus cultures, and adventitious root cultures of Panax quinquefolium, as a source of ginsenosides. Z Narturforsch C J Biosci 68:482–488
Szymczyk P, Skała E, Grąbkowska R, Jeleń A, Żebrowska M, Balcerczak E (2016) Isolation and characterization of a copalyl diphosphate synthase gene promoter from Salvia miltiorrhiza. Acta Soc Bot Pol 85:3513
Szymczyk P, Grąbkowska R, Skała E, Żebrowska M, Balcerczak E, Jeleń A (2018) Isolation and characterization of a 3-hydroxy-3-methylglutaryl coenzyme A reductase 2 promoter from Salvia miltiorrhiza. J Plant Biochem Biotechnol 27:223–236
Tabata H (2004) Paclitaxel production by plant-cell-culture technology. Adv Biochem Eng Biotechnol 87:1–23
The UniProt Consortium (2021) UniProt: the universal protein knowledgebase in 2021. Nucleic Acids Res 49(D1):D480–D489
Top O, Geisen U, Decker EL, Reski R (2019) Critical evaluation of strategies for the production of blood coagulation factors in plant-based systems. Front Plant Sci 10:261
Wan X, Wang Y, Row KH (2009) Separation of tanshinone I, tanshinone IIA, and cryptotanshinone from Salvia miltiorrhiza Bunge by normal phase HPLC. J Liq Chromatogr Relat Technol 32:544–552
Wang WW, Wu JY (2010) Tanshinone biosynthesis in Salvia miltiorrhiza and production in plant tissue cultures. Appl Microbiol Biotechnol 88:437–449
Wang L, Ma R, Liu C, Liu H, Zhu R, Guo S, Tang M, Li Y, Niu J, Fu M, Gao S (2017) Salvia miltiorrhiza: a potential red light to the development of cardiovascular diseases. Curr Pharm Des 23:1077–1097
Wei T, Gao Y, Deng K, Zhang L, Yang M, Liu X, Qi C, Wang Ch, Song W, Zhang Y, Chen Ch (2019) Enhancement of tanshinone production in Salvia miltiorrhiza hairy root cultures by metabolic engineering. Plant Methods 15:53
Wu ChT, Mulagabal V, Nalawade SM, Chen ChL, Yang TF, Tsay HS (2003) Isolation and quantitative analysis of cryptotanshinone, an active quinoid diterpene formed in callus of Salvia miltiorrhiza Bunge. Biol Pharm Bull 26:845–848
Xu Z, Ji A, Song J, Chen S (2016) Genome-wide analysis of auxin response factor gene family members in medicinal model plant Salvia miltiorrhiza. Biol Open 5:848–857
Yan X, Zhang L, Wang J, Liao P, Zhang Y, Zhang R, Kai G (2009) Molecular characterization and expression of 1-deoxy-D-xylulose 5-phosphate reductoisomerase (DXR) gene from Salvia miltiorrhiza. Acta Physiol Plant 31:1015–1022
Yu ChP, Lin JJ, Li WH (2016) Positional distribution of transcription factor binding sites in Arabidopsis thaliana. Sci Rep 6:25164
Zhang L, Yan X, Wang J, Li S, Liao P, Kai G (2010) Molecular cloning and expression analysis of a new putative gene encoding 3-hydroxy-3-methylglutaryl-CoA synthase from Salvia miltiorrhiza. Acta Physiol Plant 33:953–961
Zhang X, Dong J, Liu H, Wang J, Qi Y, Liang Z (2016) Transcriptome sequencing in response to salicylic acid in Salvia miltiorrhiza. PLoS One 11:e0147849
Zhao JL, Zhou LG, Wu JY (2010) Effects of biotic and abiotic elicitors on cell growth and tanshinone accumulation in Salvia miltiorrhiza cell cultures. Appl Microbiol Biotechnol 87:137–144
Zhao S, Zhang J, Tan R, Yang L, Zheng X (2015) Enhancing diterpenoid concentration in Salvia miltiorrhiza hairy roots through pathway engineering with maize C1 transcription factor. J Exp Bot 66:7211–7226
Zhou L, Zuo Z, Chow MS (2005) Danshen: an overview of its chemistry, pharmacology, and clinical use. J Clin Pharmacol 45:1345–1389
Acknowledgements
The authors gratefully acknowledge the help of Dr. Grażyna Szymańska at preparing the figures presented in the article.
Funding
Research supported by statutory funds of Department of Pharmaceutical Biotechnology (503/3-012-02/503-31-001), Department of Medical Biochemistry (500/6-086-01/500-41), and Department of Biology and Pharmaceutical Botany (503/3-012-01/503-31-001-19-00) at the Medical University of Łódź (Łódź, Poland).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by Maria Margarida Oliveira.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
11240_2021_2141_MOESM1_ESM.tiff
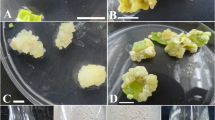
Fig S1 Pictures of S. miltiorrhiza callus growing on solid MS medium containing 1 mg L-1 2,4-dichlorophenoxyacetic acid (2,4-D) obtained after two weeks (A) and four weeks cultivation (B). Callus growing on solid MS medium containing 1 mg L-1 2,4-D and 0.5 mg L-1 N6-benzyladenine for 40 days (C). Callus culture on solid MS medium after 50 days of SA elicitation (D). From left to right following SA concentrations were applied; 0.0, 0.1, 0,2 and 0.4 mM (D). Scale bar indicates 1 cm. (TIFF 6130 kb)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Szymczyk, P., Szymańska, G., Kochan, E. et al. Elicitation of solid callus cultures of Salvia miltiorrhiza Bunge with salicylic acid and a synthetic auxin (1-naphthaleneacetic acid). Plant Cell Tiss Organ Cult 147, 491–502 (2021). https://doi.org/10.1007/s11240-021-02141-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11240-021-02141-z